

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.4 - phase: 0
(362 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
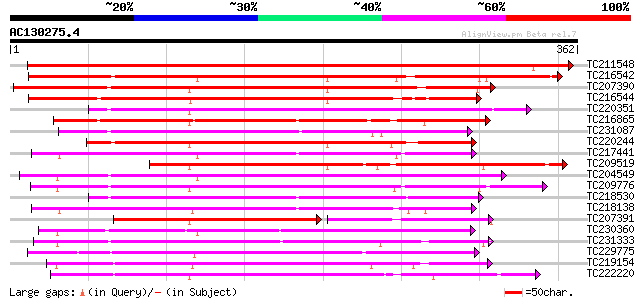
Score E
Sequences producing significant alignments: (bits) Value
TC211548 weakly similar to UP|Q7XIP7 (Q7XIP7) Receptor-like kina... 545 e-155
TC216542 weakly similar to UP|Q9LL55 (Q9LL55) Receptor-like kina... 258 3e-69
TC207390 weakly similar to UP|Q9ATQ5 (Q9ATQ5) LRK33, partial (34%) 243 1e-64
TC216544 weakly similar to UP|Q9LL55 (Q9LL55) Receptor-like kina... 241 5e-64
TC220351 similar to UP|Q961E7 (Q961E7) GH28523p (CG1830-PB) , pa... 221 4e-58
TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 214 5e-56
TC231087 similar to UP|Q39203 (Q39203) Receptor-like protein kin... 209 1e-54
TC220244 weakly similar to UP|Q9LL58 (Q9LL58) Receptor-like kina... 204 7e-53
TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic em... 196 1e-50
TC209519 similar to UP|Q84K89 (Q84K89) Receptor kinase LRK10 (Re... 193 1e-49
TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-l... 191 4e-49
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 189 2e-48
TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partia... 187 8e-48
TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis r... 187 8e-48
TC207391 similar to UP|Q9SWU8 (Q9SWU8) Receptor-like kinase (Fra... 147 2e-47
TC230360 similar to GB|AAP68229.1|31711746|BT008790 At5g63940 {A... 179 2e-45
TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, parti... 179 2e-45
TC229775 weakly similar to UP|KPEL_DROME (Q05652) Probable serin... 178 3e-45
TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 176 1e-44
TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like prot... 174 7e-44
>TC211548 weakly similar to UP|Q7XIP7 (Q7XIP7) Receptor-like kinase
TAK33-like protein, partial (58%)
Length = 1194
Score = 545 bits (1404), Expect = e-155
Identities = 269/353 (76%), Positives = 304/353 (85%), Gaps = 4/353 (1%)
Frame = +2
Query: 12 KPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAE 71
KPIRFT QQLRIATDNYS LLGSGGFG VYKG FSNGT+VAVKVLRGSS+K+IDEQFMAE
Sbjct: 2 KPIRFTDQQLRIATDNYSFLLGSGGFGVVYKGSFSNGTIVAVKVLRGSSDKRIDEQFMAE 181
Query: 72 VGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLHEIAIGT 131
VGTIG++HHFNLVRLYGFCFER+L ALVYEYM NG+L++YLFHE L +EKLHEIA+GT
Sbjct: 182 VGTIGKVHHFNLVRLYGFCFERHLRALVYEYMVNGALEKYLFHENTTLSFEKLHEIAVGT 361
Query: 132 ARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTP 191
ARGIAYLHEECQ RIIHYDIKPGNILLD+NF PKVADFGLAK CNR+NTHI+MTGGRGTP
Sbjct: 362 ARGIAYLHEECQQRIIHYDIKPGNILLDRNFCPKVADFGLAKLCNRDNTHISMTGGRGTP 541
Query: 192 GYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLL 251
GYAAPELW+PFP+THKCDVYSFGMLLFEIIGRRRN I ESQ WFP+WVW++ DA +
Sbjct: 542 GYAAPELWLPFPVTHKCDVYSFGMLLFEIIGRRRNHNINLPESQVWFPMWVWERFDAENV 721
Query: 252 GEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPKTFNPFQH 311
+ + CGIE++N+EIAER++KVAL CVQY+PE RPIMSVVVKMLEGS+E+PK NPFQH
Sbjct: 722 EDLISACGIEDQNREIAERIVKVALSCVQYKPEARPIMSVVVKMLEGSVEVPKPLNPFQH 901
Query: 312 LIDGTEFTTHSVQESNTYT-TSV---SSVMVSDSSIVCATPIMRKYEIELASS 360
LID T T VQ S T T TS+ SS +V+ S V ATP+M KYEIELAS+
Sbjct: 902 LIDWTPPPTDPVQASQTNTDTSICSDSSELVAKSVHVVATPVMTKYEIELASA 1060
>TC216542 weakly similar to UP|Q9LL55 (Q9LL55) Receptor-like kinase, partial
(38%)
Length = 1435
Score = 258 bits (659), Expect = 3e-69
Identities = 152/359 (42%), Positives = 222/359 (61%), Gaps = 18/359 (5%)
Frame = +2
Query: 13 PIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEV 72
PI ++ ++++ + LG GG+G V+KG +G VA+K+L + D F++E+
Sbjct: 368 PIGYSYKEIKKMARGFKEKLGGGGYGIVFKGKLHSGPSVAIKMLHKAKGNGQD--FISEI 541
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV--LGYEKLHEIAIG 130
TIGRIHH N+V+L G+C E + ALVYE+M NGSLD+++F + L Y++++ IAIG
Sbjct: 542 ATIGRIHHQNVVQLIGYCVEGSKRALVYEFMPNGSLDKFIFTKDGNIHLTYDEIYNIAIG 721
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGT 190
ARGIAYLH C+ +I+H+DIKP NILLD+ F PKV+DFGLAK +N+ +TMT RGT
Sbjct: 722 VARGIAYLHHGCEMQILHFDIKPHNILLDETFTPKVSDFGLAKLYPIDNSIVTMTAARGT 901
Query: 191 PGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNL-AIKNTESQEWFPIWVWKK-- 245
GY APEL+ I+HK DVYSFGMLL E+ +R+NL + SQ +FP W++ +
Sbjct: 902 IGYMAPELFYKNIGGISHKADVYSFGMLLMEMASKRKNLNPHADHSSQLYFPFWIYNQLG 1081
Query: 246 KDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEG---SLEI 302
K+ + E G+ E+ +IA++MI V+LWC+Q +P RP M+ VV+MLEG SLEI
Sbjct: 1082KETDIEME-----GVTEEENKIAKKMIIVSLWCIQLKPTDRPSMNKVVEMLEGDIESLEI 1246
Query: 303 P--------KTFNPFQHLIDGTEFTTHSVQESNTYTTSVSSVMVSDSSIVCATPIMRKY 353
P +T Q + +T + SN +++ +V D+ C T +M Y
Sbjct: 1247PPKPSLYPHETMENDQSIYSSQTMSTDFISSSNYSKEILTNPLVEDAI**C-TVVMYLY 1420
>TC207390 weakly similar to UP|Q9ATQ5 (Q9ATQ5) LRK33, partial (34%)
Length = 1342
Score = 243 bits (620), Expect = 1e-64
Identities = 133/316 (42%), Positives = 192/316 (60%), Gaps = 8/316 (2%)
Frame = +3
Query: 3 KFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNK 62
KFL D EKP RFT L+ T + LG G G V++G SN +VAVK+L + +
Sbjct: 174 KFLEDYRAEKPARFTYADLKRITGGFKEKLGEGAHGAVFRGKLSNEILVAVKILNNTEGE 353
Query: 63 KIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF---HETKVL 119
++F+ EVG +G+IHH N+VRL GFC E ALVY NGSL R++ + L
Sbjct: 354 --GKEFINEVGIMGKIHHINVVRLLGFCAEGFHRALVYNLFPNGSLQRFIVPPDDKDHFL 527
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
G+EKL +IA+G A+GI YLHE C IIH+DI P N+LLD NF PK++DFGLAK C++
Sbjct: 528 GWEKLQQIALGIAKGIEYLHEGCNQPIIHFDINPHNVLLDDNFTPKISDFGLAKLCSKNP 707
Query: 180 THITMTGGRGTPGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNLAIKNTES-QE 236
+ ++MT RGT GY APE++ +++K D+YS+GMLL E++G R+N+ + + +
Sbjct: 708 SLVSMTAARGTLGYIAPEVFSKNFGNVSYKSDIYSYGMLLLEMVGGRKNVDMSSAQDFHV 887
Query: 237 WFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
+P W+ D + C I +IA+++ V LWC+Q++P RP + V++ML
Sbjct: 888 LYPDWIHNLIDGDVHIHVEDECDI-----KIAKKLAIVGLWCIQWQPVNRPSIKSVIQML 1052
Query: 297 E--GSLEIPKTFNPFQ 310
E G ++ NPFQ
Sbjct: 1053ETGGESQLNVPPNPFQ 1100
>TC216544 weakly similar to UP|Q9LL55 (Q9LL55) Receptor-like kinase, partial
(36%)
Length = 1229
Score = 241 bits (614), Expect = 5e-64
Identities = 132/294 (44%), Positives = 196/294 (65%), Gaps = 5/294 (1%)
Frame = +2
Query: 13 PIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEV 72
PIR++ ++++ + + LG GG+G+V+KG +G+ VA+K+L +K + F++EV
Sbjct: 299 PIRYSYKEVKKMAGGFKDKLGEGGYGSVFKGKLGSGSCVAIKML--GKSKGNGQDFISEV 472
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFH--ETKVLGYEKLHEIAIG 130
TIGR +H N+V+L GFC + ALVYE+M NGSLD+++F E+ L Y++++ I+I
Sbjct: 473 ATIGRTYHQNIVQLIGFCVHGSKRALVYEFMPNGSLDKFIFSKDESIHLSYDRIYNISIE 652
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGT 190
ARGIAYLH C+ +I+H+DIKP NILLD+NF PKV+DFGLAK +N+ + T RGT
Sbjct: 653 VARGIAYLHYGCEMQILHFDIKPHNILLDENFTPKVSDFGLAKLYPIDNSIVPRTAARGT 832
Query: 191 PGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNL-AIKNTESQEWFPIWVWKKKD 247
GY APEL+ I+HK DVYS+GMLL E+ +R+NL SQ +FP W++
Sbjct: 833 IGYMAPELFYNNIGGISHKADVYSYGMLLMEMASKRKNLNPHAERSSQLFFPFWIYNH-- 1006
Query: 248 AGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLE 301
+G+ + +E+ +E ++MI VALWC+Q +P RP M+ VV+MLEG +E
Sbjct: 1007---IGDEEDI-EMEDVTEE-EKKMIIVALWCIQLKPNDRPSMNKVVEMLEGDIE 1153
>TC220351 similar to UP|Q961E7 (Q961E7) GH28523p (CG1830-PB) , partial (11%)
Length = 1064
Score = 221 bits (563), Expect = 4e-58
Identities = 130/289 (44%), Positives = 172/289 (58%), Gaps = 6/289 (2%)
Frame = +1
Query: 51 VAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDR 110
VAVK L G ++QF EV TI HH NLVRL GFC E LVYE+M NGSLD
Sbjct: 1 VAVKQLEGIEQG--EKQFRMEVATISSTHHLNLVRLIGFCSEGRHRLLVYEFMKNGSLDD 174
Query: 111 YLF----HETKVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKV 166
+LF H K+L +E IA+GTARGI YLHEEC+ I+H DIKP NILLD+N+ KV
Sbjct: 175 FLFLTEQHSGKLLNWEYRFNIALGTARGITYLHEECRDCIVHCDIKPENILLDENYVAKV 354
Query: 167 ADFGLAKNCN-RENTHITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRR 225
+DFGLAK N +++ H T+T RGT GY APE PIT K DVY +GM+L EI+ RR
Sbjct: 355 SDFGLAKLINPKDHRHRTLTSVRGTRGYLAPEWLANLPITSKSDVYGYGMVLLEIVSGRR 534
Query: 226 NLAIKNTESQEWFPIWVWKKKDAG-LLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPE 284
N + +++ F IW +++ + G + G +E + E R I+ + WC+Q +P
Sbjct: 535 NFDVSEETNRKKFSIWAYEEFEKGNISGILDKRLANQEVDMEQVRRAIQASFWCIQEQPS 714
Query: 285 LRPIMSVVVKMLEGSLEIPKTFNPFQHLIDGTEFTTHSVQESNTYTTSV 333
RP MS V++MLEG E + P + +++G T + SN SV
Sbjct: 715 HRPTMSRVLQMLEGVTEPERPPAP-KSVMEGAVSGTSTYLSSNASAFSV 858
>TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (28%)
Length = 1358
Score = 214 bits (545), Expect = 5e-56
Identities = 130/289 (44%), Positives = 178/289 (60%), Gaps = 10/289 (3%)
Frame = +1
Query: 29 SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYG 88
+N +G GGFG VYKG FS+GT++AVK L S +++ + +F+ E+G I + H +LV+LYG
Sbjct: 4 ANKIGEGGFGPVYKGCFSDGTLIAVKQL-SSKSRQGNREFLNEIGMISALQHPHLVKLYG 180
Query: 89 FCFERNLIALVYEYMGNGSLDRYLF----HETKVLGYEKLHEIAIGTARGIAYLHEECQH 144
C E + + LVYEYM N SL R LF H+ K L + ++I +G ARG+AYLHEE +
Sbjct: 181 CCVEGDQLLLVYEYMENNSLARALFGAEEHQIK-LDWTTRYKICVGIARGLAYLHEESRL 357
Query: 145 RIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMPFPI 204
+I+H DIK N+LLD++ PK++DFGLAK +NTHI+ T GT GY APE M +
Sbjct: 358 KIVHRDIKATNVLLDQDLNPKISDFGLAKLDEEDNTHIS-TRIAGTFGYMAPEYAMHGYL 534
Query: 205 THKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLLGEAMIVCGIEEK- 263
T K DVYSFG++ EII R N + + +E F + W A LL E + + ++
Sbjct: 535 TDKADVYSFGIVALEIINGRSNTI--HRQKEESFSVLEW----AHLLREKGDIMDLVDRR 696
Query: 264 -----NKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPKTFN 307
NKE A MIKVAL C LRP MS VV MLEG + + + F+
Sbjct: 697 LGLEFNKEEALVMIKVALLCTNVTAALRPTMSSVVSMLEGKIVVDEEFS 843
>TC231087 similar to UP|Q39203 (Q39203) Receptor-like protein kinase
precursor, partial (30%)
Length = 826
Score = 209 bits (533), Expect = 1e-54
Identities = 128/278 (46%), Positives = 166/278 (59%), Gaps = 14/278 (5%)
Frame = +1
Query: 32 LGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCF 91
+G GGFGTV++G S+ ++VAVK L +++F AEV TIG I H NLVRL GFC
Sbjct: 1 VGHGGFGTVFQGELSDASVVAVKRLERPGGG--EKEFRAEVSTIGNIQHVNLVRLRGFCS 174
Query: 92 ERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDI 151
E + LVYEYM NG+L+ YL E L ++ +A+GTA+GIAYLHEEC+ IIH DI
Sbjct: 175 ENSHRLLVYEYMQNGALNVYLRKEGPCLSWDVRFRVAVGTAKGIAYLHEECRCCIIHCDI 354
Query: 152 KPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMPFPITHKCDVY 211
KP NILLD +F KV+DFGLAK R+ + + +T RGT GY APE IT K DVY
Sbjct: 355 KPENILLDGDFTAKVSDFGLAKLIGRDFSRVLVT-MRGTWGYVAPEWISGVAITTKADVY 531
Query: 212 SFGMLLFEIIGRRRNLAIK--------NTESQE------WFPIWVWKKKDAGLLGEAMIV 257
S+GM L E+IG RRN+ ES + +FP W ++ G + + M
Sbjct: 532 SYGMTLLELIGGRRNVEAPLSAGGGGGGGESGDEMGGKWFFPPWAAQRIIEGNVSDVMDK 711
Query: 258 CGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKM 295
N E A R+ VA+WC+Q +RP M +VVKM
Sbjct: 712 RLGNAYNIEEARRVALVAVWCIQDDEAMRPTMGMVVKM 825
>TC220244 weakly similar to UP|Q9LL58 (Q9LL58) Receptor-like kinase, partial
(32%)
Length = 1112
Score = 204 bits (518), Expect = 7e-53
Identities = 111/258 (43%), Positives = 164/258 (63%), Gaps = 9/258 (3%)
Frame = +1
Query: 50 MVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLD 109
+VAVK+L + D F+ EVGT+G+IHH N+VRL GFC + ALVY++ NGSL
Sbjct: 37 LVAVKILNDTVGDGKD--FINEVGTMGKIHHVNVVRLLGFCADGFHRALVYDFFPNGSLQ 210
Query: 110 RYLF---HETKVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKV 166
R+L ++ LG+EKL +IA+G A+G+ YLH C RI+H+DI P NIL+D +F PK+
Sbjct: 211 RFLAPPDNKDVFLGWEKLQQIALGVAKGVEYLHLGCDQRILHFDINPHNILIDDHFVPKI 390
Query: 167 ADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRR 224
DFGLAK C + + +++T RGT GY APE++ +++K D+YS+GMLL E++G R
Sbjct: 391 TDFGLAKLCPKNQSTVSITAARGTLGYIAPEVFSRNFGNVSYKSDIYSYGMLLLEMVGGR 570
Query: 225 RNLAIKNTES-QEWFPIWV---WKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQ 280
+N + ES Q +P W+ K +D + E +E + IA+++ V LWC++
Sbjct: 571 KNTNMSAEESFQVLYPEWIHNLLKSRDVQVTIE-------DEGDVRIAKKLAIVGLWCIE 729
Query: 281 YRPELRPIMSVVVKMLEG 298
+ P RP M V++MLEG
Sbjct: 730 WNPIDRPSMKTVIQMLEG 783
>TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic embryogenesis
receptor-like kinase 3 {Arabidopsis thaliana;} , partial
(54%)
Length = 1008
Score = 196 bits (498), Expect = 1e-50
Identities = 123/293 (41%), Positives = 172/293 (57%), Gaps = 9/293 (3%)
Frame = +2
Query: 15 RFTGQQLRIATDNYSN--LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEV 72
RF+ ++L++ATDN+SN +LG GGFG VYKG ++G++VAVK L+ + QF EV
Sbjct: 116 RFSLRELQVATDNFSNKHILGRGGFGKVYKGRLADGSLVAVKRLKEERTQGG*LQFQTEV 295
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV---LGYEKLHEIAI 129
I H NL+RL GFC LVY YM NGS+ L + LG+ + IA+
Sbjct: 296 EMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERQESQPPLGWPERKRIAL 475
Query: 130 GTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRG 189
G+ARG+AYLH+ C +IIH D+K NILLD+ F V DFGLAK + ++TH+T T RG
Sbjct: 476 GSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVT-TAVRG 652
Query: 190 TPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKK---K 246
T G+ APE + K DV+ +G++L E+I +R + + + + W K K
Sbjct: 653 TIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDWVKGLLK 832
Query: 247 DAGLLGEAMIVCGIE-EKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEG 298
D L E ++ ++ N E E++I+VAL C Q P RP MS VV+MLEG
Sbjct: 833 DRKL--ETLVDADLQGSYNDEEVEQLIQVALLCTQGSPMERPKMSEVVRMLEG 985
>TC209519 similar to UP|Q84K89 (Q84K89) Receptor kinase LRK10 (Receptor
kinase LRK14), partial (19%)
Length = 866
Score = 193 bits (490), Expect = 1e-49
Identities = 119/279 (42%), Positives = 173/279 (61%), Gaps = 12/279 (4%)
Frame = +2
Query: 90 CFERNLIALVYEYMGNGSLDRYLF--HETKVLGYEKLHEIAIGTARGIAYLHEECQHRII 147
C E ALVYE+M NGSLD+Y+F E+ L Y+K +EI +G ARGIAYLH++C +I+
Sbjct: 2 CAEGEKHALVYEFMPNGSLDKYIFSKEESVSLSYDKTYEICLGIARGIAYLHQDCDVQIL 181
Query: 148 HYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMP--FPIT 205
H+DIKP NILLD NF PKV+DFGLAK ++ I +TG RGT GY APEL+ ++
Sbjct: 182 HFDIKPHNILLDDNFIPKVSDFGLAKLYPIKDKSIILTGLRGTFGYMAPELFYKNIGGVS 361
Query: 206 HKCDVYSFGMLLFEIIGRRRNLAIKNTE--SQEWFPIWVWKKKDAGLLGEAMIVCGIEEK 263
+K DVYSFGMLL E+ RRRN + +TE SQ +FP W++ D + + + + + E+
Sbjct: 362 YKADVYSFGMLLMEMGSRRRN-SNPHTEHSSQHFFPFWIY---DHFMEEKDIHMEEVSEE 529
Query: 264 NKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLE----IPK-TFNPFQHLIDGTEF 318
+K + ++M V+LWC+Q +P RP M VV+MLEG +E PK F P + ID +
Sbjct: 530 DKILVKKMFIVSLWCIQLKPNDRPSMKKVVEMLEGKVENIDMPPKPVFYPHETTIDSDQA 709
Query: 319 T-THSVQESNTYTTSVSSVMVSDSSIVCATPIMRKYEIE 356
+ + S + S+ + ++S T +M K+ ++
Sbjct: 710 SWSDSTSSCKNIDKTKSNFSLENNS--*QTMVMNKFNLQ 820
>TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-like protein,
partial (54%)
Length = 2032
Score = 191 bits (485), Expect = 4e-49
Identities = 124/316 (39%), Positives = 172/316 (54%), Gaps = 5/316 (1%)
Frame = +3
Query: 7 DMEREKPIRFTGQQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKI 64
D+ ++F + AT+ +S N LG GGFG VYKG S+G +VAVK L SS +
Sbjct: 765 DIPTVDSLQFDFSTIEAATNKFSADNKLGEGGFGEVYKGTLSSGQVVAVKRLSKSSGQG- 941
Query: 65 DEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV--LGYE 122
E+F EV + ++ H NLVRL GFC + LVYEY+ N SLD LF K L +
Sbjct: 942 GEEFKNEVVVVAKLQHRNLVRLLGFCLQGEEKILVYEYVPNKSLDYILFDPEKQRELDWG 1121
Query: 123 KLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHI 182
+ ++I G ARGI YLHE+ + RIIH D+K NILLD + PK++DFG+A+ + T
Sbjct: 1122 RRYKIIGGIARGIQYLHEDSRLRIIHRDLKASNILLDGDMNPKISDFGMARIFGVDQTQG 1301
Query: 183 TMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWV 242
+ GT GY APE M + K DVYSFG+LL EI+ ++N + T+ E +
Sbjct: 1302 NTSRIVGTYGYMAPEYAMHGEFSVKSDVYSFGVLLMEILSGKKNSSFYQTDGAEDLLSYA 1481
Query: 243 WKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLE-GSLE 301
W+ G E M E N+ R I + L CVQ P RP M+ +V ML+ ++
Sbjct: 1482 WQLWKDGTPLELMDPILRESYNQNEVIRSIHIGLLCVQEDPADRPTMATIVLMLDSNTVT 1661
Query: 302 IPKTFNPFQHLIDGTE 317
+P P + GT+
Sbjct: 1662 LPTPTQPAFFVHSGTD 1709
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein
, partial (47%)
Length = 1305
Score = 189 bits (479), Expect = 2e-48
Identities = 130/339 (38%), Positives = 182/339 (53%), Gaps = 9/339 (2%)
Frame = +2
Query: 14 IRFTGQQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAE 71
+RF + AT +S N LG GGFG VYKG+ +G VAVK L S + E+F E
Sbjct: 38 LRFDFSTIEAATQKFSEANKLGEGGFGEVYKGLLPSGQEVAVKRLSKISGQG-GEEFKNE 214
Query: 72 VGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF--HETKVLGYEKLHEIAI 129
V + ++ H NLVRL GFC E LVYE++ N SLD LF + K L + + ++I
Sbjct: 215 VEIVAKLQHRNLVRLLGFCLEGEEKILVYEFVANKSLDYILFDPEKQKSLDWTRRYKIVE 394
Query: 130 GTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRG 189
G ARGI YLHE+ + +IIH D+K N+LLD + PK++DFG+A+ + T G
Sbjct: 395 GIARGIQYLHEDSRLKIIHRDLKASNVLLDGDMNPKISDFGMARIFGVDQTQANTNRIVG 574
Query: 190 TPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWK--KKD 247
T GY +PE M + K DVYSFG+L+ EII +RN + T+ E + WK K +
Sbjct: 575 TYGYMSPEYAMHGEYSAKSDVYSFGVLILEIISGKRNSSFYETDVAEDLLSYAWKLWKDE 754
Query: 248 AGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEG---SLEIPK 304
A L E M E + R I + L CVQ P RP M+ VV ML+ +L++P
Sbjct: 755 APL--ELMDQSLRESYTRNEVIRCIHIGLLCVQEDPIDRPTMASVVLMLDSYSVTLQVPN 928
Query: 305 TFNPFQHLIDGTEFTTHSVQESNTYTTSVSSVMVSDSSI 343
P ++ TE + + TT+ +S V+D S+
Sbjct: 929 --QPAFYINSRTEPNMPKGLKIDQSTTNSTSKSVNDMSV 1039
>TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partial (25%)
Length = 1190
Score = 187 bits (474), Expect = 8e-48
Identities = 104/252 (41%), Positives = 150/252 (59%)
Frame = +3
Query: 51 VAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDR 110
+AVK L S++ QF+ E+ TI + H NLV+LYG C E + LVYEY+ N SLD+
Sbjct: 6 IAVKQLSVGSHQG-KSQFITEIATISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQ 182
Query: 111 YLFHETKVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFG 170
LF + L + ++I +G ARG+ YLHEE + RI+H D+K NILLD PK++DFG
Sbjct: 183 ALFGKCLTLNWSTRYDICLGVARGLTYLHEESRLRIVHRDVKASNILLDYELIPKISDFG 362
Query: 171 LAKNCNRENTHITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIK 230
LAK + + THI+ TG GT GY APE M +T K DV+SFG++ E++ R N
Sbjct: 363 LAKLYDDKKTHIS-TGVAGTIGYLAPEYAMRGHLTEKADVFSFGVVALELVSGRPNSDSS 539
Query: 231 NTESQEWFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMS 290
+ + W W+ + + + ++ + E N+E +R++ +AL C Q P LRP MS
Sbjct: 540 LEGEKVYLLEWAWQLHEKNCIID-LVDDRLSEFNEEEVKRVVGIALLCTQTSPTLRPSMS 716
Query: 291 VVVKMLEGSLEI 302
VV ML G +E+
Sbjct: 717 RVVAMLSGDIEV 752
>TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis receptor
kinase 1, partial (60%)
Length = 1444
Score = 187 bits (474), Expect = 8e-48
Identities = 120/294 (40%), Positives = 170/294 (57%), Gaps = 10/294 (3%)
Frame = +1
Query: 15 RFTGQQLRIATDNYSN--LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEV 72
RF+ ++L++ATD++SN +LG GGFG VYKG ++G++VAVK L+ + QF EV
Sbjct: 118 RFSLRELQVATDSFSNKNILGRGGFGKVYKGRLADGSLVAVKRLKEERTPGGELQFQTEV 297
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE---TKVLGYEKLHEIAI 129
I H NL+RL GFC LVY YM NGS+ L + L + +A+
Sbjct: 298 EMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPPYQEPLDWPTRKRVAL 477
Query: 130 GTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRG 189
G+ARG++YLH+ C +IIH D+K NILLD+ F V DFGLAK + ++TH+T T RG
Sbjct: 478 GSARGLSYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVT-TAVRG 654
Query: 190 TPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAG 249
T G+ APE + K DV+ +G++L E+I +R + + + + W K G
Sbjct: 655 TIGHIAPEYLSTGKSSEKTDVFGYGIMLLELITGQRAFDLARLANDDDVMLLDWVK---G 825
Query: 250 LLGE---AMIVCGIEEKN--KEIAERMIKVALWCVQYRPELRPIMSVVVKMLEG 298
LL E M+V + N + E++I+VAL C Q P RP MS VV+MLEG
Sbjct: 826 LLKEKKLEMLVDPDLQTNYIETEVEQLIQVALLCTQGSPMDRPKMSEVVRMLEG 987
>TC207391 similar to UP|Q9SWU8 (Q9SWU8) Receptor-like kinase (Fragment),
partial (29%)
Length = 963
Score = 147 bits (370), Expect(2) = 2e-47
Identities = 70/136 (51%), Positives = 95/136 (69%), Gaps = 3/136 (2%)
Frame = +1
Query: 67 QFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF---HETKVLGYEK 123
+F+ EV +G+IHH N+VRL G+C E ALVY + NGSL ++F + LG+EK
Sbjct: 16 RFINEVEIMGKIHHINVVRLLGYCAEGIHRALVYNFFPNGSLQSFIFPPDDKQNFLGWEK 195
Query: 124 LHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHIT 183
L IA+G A+GI YLH+ C H IIH+DI P N+LLD NF PK++DFGLAK C++ + ++
Sbjct: 196 LQNIALGIAKGIGYLHQGCNHPIIHFDINPHNVLLDDNFTPKISDFGLAKLCSKNPSLVS 375
Query: 184 MTGGRGTPGYAAPELW 199
MT RGT GY APE++
Sbjct: 376 MTAARGTLGYIAPEVF 423
Score = 60.1 bits (144), Expect(2) = 2e-47
Identities = 36/109 (33%), Positives = 62/109 (56%), Gaps = 3/109 (2%)
Frame = +2
Query: 204 ITHKCDVYSFGMLLFEIIGRRRNLAIKNTES-QEWFPIWVWKKKDAGLLGEAMIVCGIEE 262
+++K D+YS+GMLL E++G R+N+ + E +P W+ L+ + + +E
Sbjct: 443 VSYKSDIYSYGMLLLEMVGGRKNVDTSSPEDFHVLYPDWMHD-----LVHGDVHIHVEDE 607
Query: 263 KNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPKTF--NPF 309
+ +IA ++ V LWC+Q++P RP + V++MLE E T NPF
Sbjct: 608 GDVKIARKLAIVGLWCIQWQPLNRPSIKSVIQMLESKEEDLLTVPPNPF 754
>TC230360 similar to GB|AAP68229.1|31711746|BT008790 At5g63940 {Arabidopsis
thaliana;} , partial (42%)
Length = 1533
Score = 179 bits (454), Expect = 2e-45
Identities = 102/285 (35%), Positives = 165/285 (57%), Gaps = 6/285 (2%)
Frame = +2
Query: 19 QQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIG 76
Q+L AT N++ NL+G GG VY+G +G +AVK+L+ S N + ++F+ E+ I
Sbjct: 431 QELLSATSNFASDNLIGRGGCSYVYRGCLPDGKELAVKILKPSEN--VIKEFVQEIEIIT 604
Query: 77 RIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV----LGYEKLHEIAIGTA 132
+ H N++ + GFC E N + LVY+++ GSL+ L H KV G+++ +++A+G A
Sbjct: 605 TLRHKNIISISGFCLEGNHLLLVYDFLSRGSLEENL-HGNKVDCSAFGWQERYKVAVGVA 781
Query: 133 RGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPG 192
+ YLH C +IH D+K NILL +F P+++DFGLA + ++HIT T GT G
Sbjct: 782 EALDYLHNGCAQAVIHRDVKSSNILLADDFEPQLSDFGLA-SWGSSSSHITCTDVAGTFG 958
Query: 193 YAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLLG 252
Y APE +M +T K DVY+FG++L E++ R+ + ++ + QE +W + G
Sbjct: 959 YLAPEYFMHGRVTDKIDVYAFGVVLLELLSNRKPINNESPKGQESLVMWATPILEGGKFS 1138
Query: 253 EAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
+ + E N +RMI A C++ P LRP +++ + LE
Sbjct: 1139QLLDPSLGSEYNTCQIKRMILAATLCIRRIPRLRPQINLALLDLE 1273
>TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partial (38%)
Length = 1009
Score = 179 bits (454), Expect = 2e-45
Identities = 116/304 (38%), Positives = 175/304 (57%), Gaps = 10/304 (3%)
Frame = +3
Query: 16 FTGQQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVG 73
FT +L++AT +S N L GGFG+V++G+ +G ++AVK + +S + D++F +EV
Sbjct: 6 FTFSELQLATGGFSQANFLAEGGFGSVHRGVLPDGQVIAVKQYKLASTQG-DKEFCSEVE 182
Query: 74 TIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF-HETKVLGYEKLHEIAIGTA 132
+ H N+V L GFC + LVYEY+ NGSLD +++ + VL + +IA+G A
Sbjct: 183 VLSCAQHRNVVMLIGFCVDDGRRLLVYEYICNGSLDSHIYRRKQNVLEWSARQKIAVGAA 362
Query: 133 RGIAYLHEECQ-HRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTP 191
RG+ YLHEEC+ I+H D++P NILL +F V DFGLA+ + T GT
Sbjct: 363 RGLRYLHEECRVGCIVHRDMRPNNILLTHDFEALVGDFGLAR-WQPDGDMGVETRVIGTF 539
Query: 192 GYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQ----EWFPIWVWKKKD 247
GY APE IT K DVYSFG++L E++ R+ + I + Q EW + K+
Sbjct: 540 GYLAPEYAQSGQITEKADVYSFGIVLLELVTGRKAVDINRPKGQQCLSEWARPLLEKQAT 719
Query: 248 AGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLE--IPKT 305
L+ ++ C ++++ RM+K + C+ P LRP MS V++MLEG + +P T
Sbjct: 720 YKLIDPSLRNCYVDQE----VYRMLKCSSLCIGRDPHLRPRMSQVLRMLEGDILM*LPIT 887
Query: 306 FNPF 309
F PF
Sbjct: 888 F-PF 896
>TC229775 weakly similar to UP|KPEL_DROME (Q05652) Probable
serine/threonine-protein kinase pelle , partial (8%)
Length = 1250
Score = 178 bits (452), Expect = 3e-45
Identities = 112/292 (38%), Positives = 163/292 (55%), Gaps = 3/292 (1%)
Frame = +1
Query: 12 KPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAE 71
K ++T +L T+N++ +LG GGFG VY G F + T VAVK+L S+ + EQF+AE
Sbjct: 37 KQRQYTFNELVKITNNFTRILGRGGFGKVYHG-FIDDTQVAVKMLSPSAVRGY-EQFLAE 210
Query: 72 VGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET---KVLGYEKLHEIA 128
V + R+HH NL L G+C E N I L+YEYM NG+LD + ++ K L +E +IA
Sbjct: 211 VKLLMRVHHRNLTSLVGYCNEENNIGLIYEYMANGNLDEIVSGKSSRAKFLTWEDRLQIA 390
Query: 129 IGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGR 188
+ A+G+ YLH C+ IIH D+K NILL++NF K+ADFGL+K+ + T
Sbjct: 391 LDAAQGLEYLHNGCKPPIIHRDVKCANILLNENFQAKLADFGLSKSFPTDGGSYMSTVVA 570
Query: 189 GTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDA 248
GTPGY PE + +T K DVYSFG++L E++ + AI T + WV
Sbjct: 571 GTPGYLDPEYSISSRLTEKSDVYSFGVVLLEMVTGQP--AIAKTPDKTHISQWVKSMLSN 744
Query: 249 GLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSL 300
G + E+ + R++++ + V P RP MS +V L+ L
Sbjct: 745 GDIKNIADSRFKEDFDTSSVWRIVEIGMASVSISPFKRPSMSYIVNELKECL 900
>TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (31%)
Length = 1363
Score = 176 bits (447), Expect = 1e-44
Identities = 113/294 (38%), Positives = 168/294 (56%), Gaps = 9/294 (3%)
Frame = +1
Query: 24 ATDNY--SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHF 81
ATDN+ +N +G GGFG VYKG G +AVK L S + I E F+ EV I ++ H
Sbjct: 337 ATDNFLLNNKIGEGGFGPVYKGKLVGGQEIAVKRLSSLSGQGITE-FITEVKLIAKLQHR 513
Query: 82 NLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE--TKVLGYEKLHEIAIGTARGIAYLH 139
NLV+L G C + LVYEY+ NGSL+ ++F + +K+L + + I +G ARG+ YLH
Sbjct: 514 NLVKLLGCCIKGQEKLLVYEYVVNGSLNSFIFDQIKSKLLDWPRRFNIILGIARGLLYLH 693
Query: 140 EECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELW 199
++ + RIIH D+K N+LLD+ PK++DFG+A+ + T GT GY APE
Sbjct: 694 QDSRLRIIHRDLKASNVLLDEKLNPKISDFGMARAFGGDQTEGNTNRVVGTYGYMAPEYA 873
Query: 200 MPFPITHKCDVYSFGMLLFEIIGRRRNLAI---KNTESQEWFPIWVWKKKDAGLLGEAMI 256
+ K DV+SFG+LL EI+ +N ++ T + + +WK+++A L ++ I
Sbjct: 874 FDGNFSIKSDVFSFGILLLEIVCGIKNKSLCHENQTLNLVGYAWALWKEQNALQLIDSGI 1053
Query: 257 --VCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPKTFNP 308
C I E R I V+L CVQ PE RP M+ V++ML +++ + P
Sbjct: 1054KDSCVIPE-----VLRCIHVSLLCVQQYPEDRPTMTSVIQMLGSEMDMVEPKEP 1200
>TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like protein kinase,
RLK3 precursor, partial (35%)
Length = 1304
Score = 174 bits (440), Expect = 7e-44
Identities = 109/321 (33%), Positives = 174/321 (53%), Gaps = 8/321 (2%)
Frame = +3
Query: 27 NYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRL 86
+Y +G GGFG VYKG+ +G +AVK L SS + +E F E+ I ++ H NLV L
Sbjct: 228 SYEKRIGEGGFGVVYKGVLPDGREIAVKKLSKSSGQGANE-FKNEILLIAKLQHRNLVTL 404
Query: 87 YGFCFERNLIALVYEYMGNGSLDRYLF--HETKVLGYEKLHEIAIGTARGIAYLHEECQH 144
GFC E + L+YE++ N SLD +LF H +K L + + ++I G A+GI+YLHE +
Sbjct: 405 LGFCLEEHEKMLIYEFVSNKSLDYFLFDSHRSKQLNWSERYKIIEGIAQGISYLHEHSRL 584
Query: 145 RIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMPFPI 204
++IH D+KP N+LLD N PK++DFG+A+ + GT GY +PE M
Sbjct: 585 KVIHRDLKPSNVLLDSNMNPKISDFGMARIVAIDQLQGKTNRIVGTYGYMSPEYAMHGQF 764
Query: 205 THKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLLGEAMIVCGIEEKN 264
+ K DV+SFG+++ EII +RN ++ + + W++ + EA + +
Sbjct: 765 SEKSDVFSFGVIVLEIISAKRNTRSVFSDHDDLLS-YTWEQ----WMDEAPLNIFDQSIE 929
Query: 265 KEIAE-----RMIKVALWCVQYRPELRPIMSVVVKMLEGSL-EIPKTFNPFQHLIDGTEF 318
E + + I++ L CVQ +P+ RP ++ V+ L S+ E+P P + G
Sbjct: 930 AEFCDHSEVVKCIQIGLLCVQEKPDDRPKITQVISYLNSSITELPLPKKPIRQ--SGIVQ 1103
Query: 319 TTHSVQESNTYTTSVSSVMVS 339
+ S+ T S++ + VS
Sbjct: 1104KIAVGESSSGSTPSINEMSVS 1166
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,703,072
Number of Sequences: 63676
Number of extensions: 225367
Number of successful extensions: 2805
Number of sequences better than 10.0: 942
Number of HSP's better than 10.0 without gapping: 2065
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2096
length of query: 362
length of database: 12,639,632
effective HSP length: 98
effective length of query: 264
effective length of database: 6,399,384
effective search space: 1689437376
effective search space used: 1689437376
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC130275.4


