

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.11 - phase: 0
(309 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
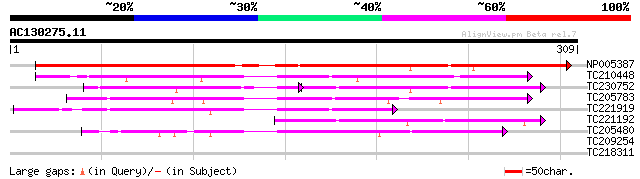
Score E
Sequences producing significant alignments: (bits) Value
NP005387 putative narbonin 237 4e-63
TC210448 similar to UP|CHI1_TULBA (Q9SLP4) Chitinase 1 precursor... 116 1e-26
TC230752 weakly similar to UP|RUAP_SOYBN (P39657) RuBisCO-associ... 62 5e-21
TC205783 UP|RUAP_SOYBN (P39657) RuBisCO-associated protein, comp... 80 1e-15
TC221919 weakly similar to UP|RUAP_SOYBN (P39657) RuBisCO-associ... 80 2e-15
TC221192 similar to UP|Q8WZI7 (Q8WZI7) Ferulic acid esterase A p... 79 2e-15
TC205480 weakly similar to UP|Q08883 (Q08883) Narbonin, partial ... 65 3e-11
TC209254 homologue to UP|Q9FUN3 (Q9FUN3) QM-like protein, partia... 27 9.0
TC218311 similar to UP|Q84MB9 (Q84MB9) At1g48300, partial (19%) 27 9.0
>NP005387 putative narbonin
Length = 873
Score = 237 bits (605), Expect = 4e-63
Identities = 135/297 (45%), Positives = 182/297 (60%), Gaps = 5/297 (1%)
Frame = +1
Query: 15 IFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNF 74
IFREYIGVK + L DFP E+I+ + EFHFILG E Y GK G F +W+ +F
Sbjct: 13 IFREYIGVKPNSETLHDFPHEIIDTENLEFHFILGFATESYYESGKSTGNFEESWDSESF 192
Query: 75 SPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQEYSKDS 134
V KLK+ + VKV+ISIGG G + PF+P E W A++S+K++ +Q+YS +S
Sbjct: 193 GLENVKKLKERYPEVKVVISIGGRGVQTPFHPAEENVWVENAQESLKQI---FQKYSNES 363
Query: 135 SSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSK-SIKLVSIAPTELLK 193
S +IDGIDINYE+ + + + F+ +G LI +LKK +I +VSIAP+E
Sbjct: 364 GS--------MIDGIDINYEHIS-SDEPFARLVGLLITELKKDDDLNINVVSIAPSETNS 516
Query: 194 PHYHKLYWANKDIINWVDYKFYNQ--TVSSADELVNLYNKLLNEYGTDVKLLPGVSTDPD 251
HY KLY A KD INWVDY+F NQ V D V ++ + +Y K+LPG STDPD
Sbjct: 517 SHYQKLYNAKKDYINWVDYQFSNQHKPVHKDDHFVEIFKTVEKDYHLR-KVLPGFSTDPD 693
Query: 252 --SNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYFLEEVLQDLL 306
+ +TRDVFI GC L ++ SLPG+F WNA+DS P + + P+ +E LQ LL
Sbjct: 694 DAKHNKITRDVFIGGCTRLKQTSSLPGVFFWNAHDSVTPRLDGDKPFIVELTLQQLL 864
>TC210448 similar to UP|CHI1_TULBA (Q9SLP4) Chitinase 1 precursor (Tulip
bulb chitinase-1) (TBC-1) , partial (70%)
Length = 1157
Score = 116 bits (291), Expect = 1e-26
Identities = 88/281 (31%), Positives = 138/281 (48%), Gaps = 10/281 (3%)
Frame = +1
Query: 15 IFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGK---GEFYRTWNF 71
+FREYIG D P IN ++ FHFIL + + G G+F W+
Sbjct: 199 LFREYIGALFNGVKFSDVP---INPNVN-FHFILSFAIDYDTSSGSPSPTNGKFNIFWDN 366
Query: 72 NNFSPAKVAKLKKDHKNVKVIISIGGFGAENP---FNPKEIESWSTKAKQSIKKLINEYQ 128
N +P +V+ +K + NVKV +S+GG + F+P ++SW + A S+ +I EY
Sbjct: 367 QNLTPTQVSSIKAKNPNVKVALSLGGDTVSSAFAYFDPTSVDSWVSNAVSSLTNIIKEYN 546
Query: 129 EYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAP 188
+DGIDI+YE+ + + F+ IG+LI+ L KS I SIAP
Sbjct: 547 -----------------LDGIDIDYEHFKADSETFAQSIGKLIQTL-KSRGVITFASIAP 672
Query: 189 --TELLKPHYHKLYWANKDIINWVDYKFYNQTVS-SADELVNLYNKLLNEYGTDVKLLPG 245
+ ++ HY L+ + II++V+++FY S + + ++ +NK + Y L
Sbjct: 673 FDNDEVQSHYLALWKSYGHIIDYVNFQFYAYDRSTTVSQFIDYFNKQSSNYNGGKVL--- 843
Query: 246 VSTDPDSNTNMTRD-VFIKGCKSLLESESLPGIFVWNANDS 285
VS + D + + D F C L + L GIFVW A+DS
Sbjct: 844 VSFNSDGSRGLIPDNGFFTACSMLKTQQKLHGIFVWCADDS 966
>TC230752 weakly similar to UP|RUAP_SOYBN (P39657) RuBisCO-associated
protein, partial (9%)
Length = 1028
Score = 61.6 bits (148), Expect(2) = 5e-21
Identities = 43/136 (31%), Positives = 66/136 (47%), Gaps = 3/136 (2%)
Frame = +1
Query: 160 PDEFSSCIGELIRKLKKSSKSIKLVSIAPTELLKPHYHK-LYWANKDIINWVDYKFYNQ- 217
P +F C+G+LIR LK+ + SI+P+ L Y+ LY A ++WVDY+F ++
Sbjct: 427 PGDFIECVGQLIRNLKEKG-VVSEASISPSFALNEEYYPLLYSAVSFFVDWVDYQFQSEL 603
Query: 218 -TVSSADELVNLYNKLLNEYGTDVKLLPGVSTDPDSNTNMTRDVFIKGCKSLLESESLPG 276
V LV YN+L Y KL G S + + ++ VF G +L+ PG
Sbjct: 604 KPVFDPTTLVKRYNELTKLYPRR-KLFAGYSAENEDWATLSPIVFFLGGMDILKKRKAPG 780
Query: 277 IFVWNANDSAMPSNED 292
+ + N A N+D
Sbjct: 781 VSIHYHNYYADAPNKD 828
Score = 57.0 bits (136), Expect(2) = 5e-21
Identities = 37/123 (30%), Positives = 61/123 (49%), Gaps = 3/123 (2%)
Frame = +3
Query: 41 IEEFHFILGTLREVYSGDGKG-KGEFYRTWNFNNFSPAKVAKLKKDHKNV--KVIISIGG 97
++E+ L T Y +G G F TW+ + +P +A+ K + NV KV ISIG
Sbjct: 102 LKEYQIAL-TFASDYDDEGAPTNGVFRPTWDLSKVTPESIARFKDKNPNVDIKVFISIGN 278
Query: 98 FGAENPFNPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSN 157
G ++PF P ++W A +S+ LI ++Y+ +DGID+ YE+ +
Sbjct: 279 RGTQHPFKPLNNKTWIDNATESLTHLIKN-EDYNLH------------VDGIDVLYEHID 419
Query: 158 CNP 160
+P
Sbjct: 420 ASP 428
>TC205783 UP|RUAP_SOYBN (P39657) RuBisCO-associated protein, complete
Length = 1353
Score = 80.1 bits (196), Expect = 1e-15
Identities = 70/272 (25%), Positives = 122/272 (44%), Gaps = 18/272 (6%)
Frame = +2
Query: 32 FPAEMINDDIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNFSPAKVAKLKKDHK---- 87
F ++I ++I EF L R+ Y G+ G+F W+ +P + K KK ++
Sbjct: 134 FLNQVIPENITEFQVTLSLARD-YDGNNSTNGKFIPYWDTEKVTPEVIKKFKKKYEPTAL 310
Query: 88 NVKVIISIGGFGAENPF---NPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCDD 144
VKV++SIG + PF + E+W ++A S+K +I Y
Sbjct: 311 RVKVLVSIGNKNKQFPFTIGSDSNSEAWVSEATASLKSIIKTYN---------------- 442
Query: 145 IIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAPT-ELLKPHYHKLYWAN 203
+DGID++YE N +F + +G L+R LK+ +K I + S A + + ++ L +A
Sbjct: 443 -LDGIDVSYEDIAANEADFVNSVGGLVRNLKQ-NKLITVASFATSADAANNKFYNLLYAE 616
Query: 204 KD-------IINWVDYKFYNQTVSSADELVNLYNKLL---NEYGTDVKLLPGVSTDPDSN 253
++WV + T S A+ + +L K+L NEY L ST +
Sbjct: 617 YATFFDTVVFLSWVGF-----TPSRANPVASLEEKILAVANEYKAVKAFLVAYSTVAEDW 781
Query: 254 TNMTRDVFIKGCKSLLESESLPGIFVWNANDS 285
N + +F LL + ++ G + +D+
Sbjct: 782 ANFSPPIFFITLHGLLGNSAVKGASIKVISDA 877
>TC221919 weakly similar to UP|RUAP_SOYBN (P39657) RuBisCO-associated
protein, partial (31%)
Length = 688
Score = 79.7 bits (195), Expect = 2e-15
Identities = 66/213 (30%), Positives = 96/213 (44%), Gaps = 4/213 (1%)
Frame = +2
Query: 3 HTHQINAVVKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGK 62
H I+ I+FREY K L D P D+ E L R+ Y G G +
Sbjct: 77 HISHISMAGSKIVFREYTSDDSFTK-LTDIPK-----DVSEIQVALTFARD-YDGVGSTQ 235
Query: 63 GEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKE--IESWSTKAKQSI 120
G+F W+ +P +A K + + KV++SIG PF E + +W A +S+
Sbjct: 236 GKFIPYWDTAIVNPDVIANFKSKYPSAKVLVSIGNNKRHFPFKIDEDRVTAWINAATESL 415
Query: 121 KKLINEYQEYSKDSSSTDECHCDDIIDGIDINYE-YSNCNPDEFSSCIGELIRKLKKSSK 179
++I EY +DGID+ YE NP F CIGELI+ L K +K
Sbjct: 416 TRIIKEYN-----------------LDGIDVYYEDIEVSNPAIFVHCIGELIKTL-KLNK 541
Query: 180 SIKLVSIAPT-ELLKPHYHKLYWANKDIINWVD 211
I SI+P E+ Y L+ +D I++V+
Sbjct: 542 VITRASISPNIEINDNFYRSLHQNFEDHIDYVN 640
>TC221192 similar to UP|Q8WZI7 (Q8WZI7) Ferulic acid esterase A precursor,
partial (6%)
Length = 706
Score = 79.3 bits (194), Expect = 2e-15
Identities = 48/153 (31%), Positives = 82/153 (53%), Gaps = 5/153 (3%)
Frame = +3
Query: 145 IIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAPTELLKPHYHKLYWANK 204
++ GID+ Y++ + +P +F+ C+G+LIR LK++ + SI+P+ Y+ L +++
Sbjct: 54 VVHGIDVLYDHIDASPADFTECVGQLIRNLKENG-VVSQASISPSSYPNQEYYPLLYSSV 230
Query: 205 D-IINWVDYKFY--NQTVSSADELVNLYNKLLNEYGTDVKLLPGVSTDPDSNTNMTRDVF 261
++WVDY+F +Q V LV YN+L Y KL G S + + ++ VF
Sbjct: 231 PFFVDWVDYQFQSEDQPVLEPTTLVKRYNELTKVY-PKTKLFAGYSAENEDWATVSPFVF 407
Query: 262 IKGCKSLLESESLPGIFV--WNANDSAMPSNED 292
G LL+ S PG+ + N S P+N+D
Sbjct: 408 FLGAMDLLKKRSAPGVSIHYHNYYASEAPNNKD 506
>TC205480 weakly similar to UP|Q08883 (Q08883) Narbonin, partial (8%)
Length = 1157
Score = 65.5 bits (158), Expect = 3e-11
Identities = 69/248 (27%), Positives = 108/248 (42%), Gaps = 16/248 (6%)
Frame = +3
Query: 40 DIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNFSPAKVA--KLKKDHKN--VKVIISI 95
D+ E+ L Y+G+ GKG+F W+ +N +P + KL D N VKV +SI
Sbjct: 165 DVTEYQLCLA-----YAGES-GKGKFEPKWDTDNVTPQLIEQFKLNCDASNLPVKVFVSI 326
Query: 96 GGFGAENPFNPKE--IESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCDDIIDGIDINY 153
GG ++PF + W A ++ +INEY IDGID+NY
Sbjct: 327 GG---DDPFKVAAGGADDWVLNATTTLHHMINEYH-----------------IDGIDVNY 446
Query: 154 EYSNCNPDEFSSCIGELIRKLKKSSKSIKLV-SIAPTELL-KPHYHKLY------WANKD 205
CN F +CI LI+ L+++ + V S+AP + Y LY W
Sbjct: 447 TTIQCNQTLFVNCIKGLIKGLREALCTTGFVASLAPDAATNETFYLPLYRSIDNSWIGTV 626
Query: 206 IIN-WVDYKFYNQTVSSADELVNLYNKLLNEYG-TDVKLLPGVSTDPDSNTNMTRDVFIK 263
+ +VD++ + + LV K++ EY D K+L G S + ++ +F
Sbjct: 627 VFQCFVDFEPLDPN-RPVESLVRQVEKIVGEYDYYDSKILLGYSAIKEDWNKVSPPLFFL 803
Query: 264 GCKSLLES 271
L+ S
Sbjct: 804 ALNRLVHS 827
>TC209254 homologue to UP|Q9FUN3 (Q9FUN3) QM-like protein, partial (96%)
Length = 1060
Score = 27.3 bits (59), Expect = 9.0
Identities = 17/69 (24%), Positives = 27/69 (38%)
Frame = +2
Query: 40 DIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFG 99
D H R + G+ K R W F FS ++ KLK +++ V ++ G
Sbjct: 572 DANSHHAQEALRRAKFKFPGRQKIILSRKWGFTKFSRSEYLKLKSENRIVPDGVNAKVLG 751
Query: 100 AENPFNPKE 108
P +E
Sbjct: 752 CHGPLANRE 778
>TC218311 similar to UP|Q84MB9 (Q84MB9) At1g48300, partial (19%)
Length = 1004
Score = 27.3 bits (59), Expect = 9.0
Identities = 12/33 (36%), Positives = 16/33 (48%)
Frame = +3
Query: 115 KAKQSIKKLINEYQEYSKDSSSTDECHCDDIID 147
KAK K+ E S SS +C CD ++D
Sbjct: 33 KAKLKAAKMNCESSSSSSSESSDSDCGCDQVVD 131
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,612,320
Number of Sequences: 63676
Number of extensions: 203568
Number of successful extensions: 692
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 677
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 679
length of query: 309
length of database: 12,639,632
effective HSP length: 97
effective length of query: 212
effective length of database: 6,463,060
effective search space: 1370168720
effective search space used: 1370168720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC130275.11


