

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130200.9 - phase: 0
(393 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
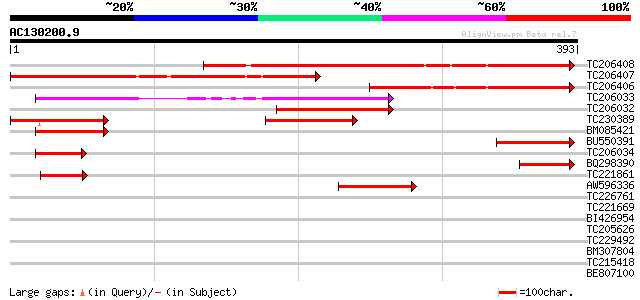
TC206408 similar to UP|Q9LRY5 (Q9LRY5) Gb|AAF16548.1 (AT3g24740/... 399 e-112
TC206407 similar to UP|Q9LRY5 (Q9LRY5) Gb|AAF16548.1 (AT3g24740/... 363 e-101
TC206406 weakly similar to UP|Q9LRY5 (Q9LRY5) Gb|AAF16548.1 (AT3... 211 5e-55
TC206033 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/... 192 2e-49
TC206032 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/... 97 2e-20
TC230389 95 6e-20
BM085421 similar to GP|11034559|dbj P0410E01.4 {Oryza sativa (ja... 81 7e-16
BU550391 homologue to GP|19715579|gb| AT3g24740/K7P8_3 {Arabidop... 80 2e-15
TC206034 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/... 67 2e-11
BQ298390 60 2e-09
TC221861 similar to GB|AAM19869.1|20453261|AY097353 AT4g31410/F8... 60 2e-09
AW596336 similar to GP|18650602|gb| AT4g31410/F8F16_230 {Arabido... 59 3e-09
TC226761 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding p... 38 0.007
TC221669 32 0.38
BI426954 similar to GP|5734709|gb|A Contains PF|00069 Eukaryotic... 32 0.50
TC205626 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g7539... 30 1.9
TC229492 similar to UP|Q9FT86 (Q9FT86) Peroxin Pex14, partial (13%) 30 1.9
BM307804 similar to GP|18766642|gb| NIMA-related protein kinase ... 29 3.2
TC215418 similar to GB|AAB90084.1|2649426|AE001023 A. fulgidus p... 29 3.2
BE807100 weakly similar to SP|O60879|DIA2 Diaphanous protein hom... 29 3.2
>TC206408 similar to UP|Q9LRY5 (Q9LRY5) Gb|AAF16548.1 (AT3g24740/K7P8_3),
partial (44%)
Length = 903
Score = 399 bits (1026), Expect = e-112
Identities = 203/257 (78%), Positives = 222/257 (85%)
Frame = +3
Query: 135 QDPSRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEV 194
QDPSRHLDQHDEGILETADSE LQDRAVLE D+D + +SK +L CPLCRG+VL WEV
Sbjct: 3 QDPSRHLDQHDEGILETADSETLQDRAVLE---DLDADASESKLNLKCPLCRGSVLNWEV 173
Query: 195 VEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTREQAWQQFERQR 254
VEEARNYLN KKRSCSRDSCSF GDYLELRRHARRVHPTSRPS+VDPTRE+AW+ FERQR
Sbjct: 174 VEEARNYLNMKKRSCSRDSCSFVGDYLELRRHARRVHPTSRPSNVDPTRERAWRHFERQR 353
Query: 255 EYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNGNVPWLTTTTILFQM 314
EYGDIVSAIQSA+PGAV+VGDY LENGDGIGRL EG+ N N PWL TTILFQM
Sbjct: 354 EYGDIVSAIQSAMPGAVLVGDYALENGDGIGRL-QDERVEGNIDNANRPWL-ATTILFQM 527
Query: 315 MDNTIEIVREPRARSSNGWSRHRRSSDRRRYLWGENLLGLQDNEVEEDLRIFNELVEDAS 374
MD+TIEIVREPRA SS W+RHRRSS+RRRYLWGE+LLGL DN++E+DLRIF + EDAS
Sbjct: 528 MDSTIEIVREPRAHSS-AWTRHRRSSERRRYLWGESLLGLHDNDIEDDLRIFRDAGEDAS 704
Query: 375 HVPRRRRRLNRTRSNED 391
VPRRRRRL RTRSNED
Sbjct: 705 PVPRRRRRLTRTRSNED 755
>TC206407 similar to UP|Q9LRY5 (Q9LRY5) Gb|AAF16548.1 (AT3g24740/K7P8_3),
partial (31%)
Length = 894
Score = 363 bits (932), Expect = e-101
Identities = 180/215 (83%), Positives = 196/215 (90%)
Frame = +2
Query: 1 MAGFKRRLCNDSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS 60
MAG KRRLC+DSD+HALH+ELDEVSCPICMDHPHNAVLLLCSSH+KGCRSYICDTSYRHS
Sbjct: 263 MAGVKRRLCSDSDIHALHKELDEVSCPICMDHPHNAVLLLCSSHEKGCRSYICDTSYRHS 442
Query: 61 NCLDRFKKLRDNSKENPNLQSSLINTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTL 120
NCLDRFKK+RDN KEN NL SSL+NTNN SGNSFDIN T+QSDMHDV+E L+QNEIN L
Sbjct: 443 NCLDRFKKMRDNFKENQNLPSSLVNTNN-SGNSFDINLTVQSDMHDVNE--LHQNEINAL 613
Query: 121 LSVGIPQGSRQGDAQDPSRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSL 180
LSVG+ QGSRQGDAQDP+R LDQHDEGILETADSENLQDRAV+ E+L+ DNSSE SK +L
Sbjct: 614 LSVGLAQGSRQGDAQDPNRLLDQHDEGILETADSENLQDRAVI-EDLNADNSSE-SKLNL 787
Query: 181 HCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCS 215
CPLCRG VL W+VVEEARNYLN KKRSCSRDSCS
Sbjct: 788 KCPLCRGAVLNWKVVEEARNYLNMKKRSCSRDSCS 892
>TC206406 weakly similar to UP|Q9LRY5 (Q9LRY5) Gb|AAF16548.1
(AT3g24740/K7P8_3), partial (14%)
Length = 881
Score = 211 bits (537), Expect = 5e-55
Identities = 109/142 (76%), Positives = 120/142 (83%)
Frame = +3
Query: 250 FERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNGNVPWLTTTT 309
FE QREYG VSAIQSA+PGAV+VGDYVLENGDGIGRLP EG+ GN N PWLTTT
Sbjct: 6 FEDQREYGGWVSAIQSAVPGAVLVGDYVLENGDGIGRLPDERA-EGNIGNANGPWLTTT- 179
Query: 310 ILFQMMDNTIEIVREPRARSSNGWSRHRRSSDRRRYLWGENLLGLQDNEVEEDLRIFNEL 369
ILFQMMD+T+EIVREPRA SS W+RHRRS +RRRYLWGENLLGL DN++E+DLRIF +
Sbjct: 180 ILFQMMDSTVEIVREPRAHSS-AWTRHRRSDERRRYLWGENLLGLHDNDIEDDLRIFRDA 356
Query: 370 VEDASHVPRRRRRLNRTRSNED 391
EDAS VPRRRRRL RTRSNED
Sbjct: 357 GEDASPVPRRRRRLTRTRSNED 422
>TC206033 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/MPE11_6),
partial (40%)
Length = 1468
Score = 192 bits (489), Expect = 2e-49
Identities = 100/249 (40%), Positives = 146/249 (58%), Gaps = 1/249 (0%)
Frame = +2
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRF-KKLRDNSKENP 77
+E +E CP+CM+HPHNAVLL+CSSH+KGCR Y+C+TSYRHSNCLD+F K D S P
Sbjct: 440 KEWEEARCPVCMEHPHNAVLLICSSHEKGCRPYMCNTSYRHSNCLDQFCKSFADTSSTIP 619
Query: 78 NLQSSLINTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTLLSVGIPQGSRQGDAQDP 137
++ + +T S + +P+ +G Q
Sbjct: 620 QVEGQISDTFESR--------------------------------IDVPEDRNEGTVQS- 700
Query: 138 SRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEVVEE 197
+D ++ SE+L V + L +N D+KS L CPLCRG + W+VVE
Sbjct: 701 --RIDVPEDR------SEDL----VTMQTLSCEN---DAKSKLVCPLCRGQIKEWKVVEA 835
Query: 198 ARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREYG 257
AR+++N K RSCS ++C+F+G Y +LR+HAR HP RPS VDP R+++W++ ERQR+ G
Sbjct: 836 ARHFMNEKSRSCSCETCNFSGTYPDLRKHARLEHPLERPSAVDPERQRSWRRLERQRDLG 1015
Query: 258 DIVSAIQSA 266
D++S +Q++
Sbjct: 1016DLLSTLQTS 1042
>TC206032 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/MPE11_6),
partial (23%)
Length = 422
Score = 96.7 bits (239), Expect = 2e-20
Identities = 39/81 (48%), Positives = 62/81 (76%)
Frame = +3
Query: 186 RGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTREQ 245
+G + W+VVE AR+++N K R+CS ++C+F+G Y +LR HAR HP RPS VDP R++
Sbjct: 3 QGKIKEWKVVEAARHFMNEKSRNCSCETCNFSGTYPDLRNHARLEHPLERPSAVDPERQR 182
Query: 246 AWQQFERQREYGDIVSAIQSA 266
+W++ ERQR+ GD++S +Q++
Sbjct: 183 SWRRLERQRDLGDLLSTLQTS 245
>TC230389
Length = 785
Score = 94.7 bits (234), Expect = 6e-20
Identities = 41/73 (56%), Positives = 54/73 (73%), Gaps = 5/73 (6%)
Frame = +2
Query: 1 MAGFKRRLCNDSDMHALHR-----ELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDT 55
+A KR +C D + EL++V+C +CM++PHNAVLLLCSSHDKGCR Y+C T
Sbjct: 272 LASCKRDICEDMCQKKCSKALEKKELEDVTCSVCMEYPHNAVLLLCSSHDKGCRPYMCGT 451
Query: 56 SYRHSNCLDRFKK 68
S+RHSNCLD++KK
Sbjct: 452 SFRHSNCLDQYKK 490
Score = 85.1 bits (209), Expect = 5e-17
Identities = 36/64 (56%), Positives = 44/64 (68%)
Frame = +2
Query: 178 SSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPS 237
+ L CPLCRG V GW VVE R+YLN KKR C +D C + G Y EL++H R HP++RP
Sbjct: 587 TELACPLCRGQVKGWTVVEPVRDYLNAKKRGCMQDDCLYVGSYKELKKHVRAEHPSARPR 766
Query: 238 DVDP 241
VDP
Sbjct: 767 MVDP 778
>BM085421 similar to GP|11034559|dbj P0410E01.4 {Oryza sativa (japonica
cultivar-group)}, partial (16%)
Length = 428
Score = 81.3 bits (199), Expect = 7e-16
Identities = 33/50 (66%), Positives = 41/50 (82%)
Frame = +2
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKK 68
+E +E CPICM+ PHN VLL CSS++KGC Y+C+TSYRHSNCLD+F K
Sbjct: 254 KEWEEARCPICMEPPHNGVLLKCSSYEKGCLPYMCNTSYRHSNCLDQFCK 403
>BU550391 homologue to GP|19715579|gb| AT3g24740/K7P8_3 {Arabidopsis
thaliana}, partial (3%)
Length = 425
Score = 79.7 bits (195), Expect = 2e-15
Identities = 37/54 (68%), Positives = 42/54 (77%)
Frame = -2
Query: 338 RSSDRRRYLWGENLLGLQDNEVEEDLRIFNELVEDASHVPRRRRRLNRTRSNED 391
R + RYLWGENLLGL DN++E+ RIF + EDAS VPRRRRRL RTRSNED
Sbjct: 424 RXDEXXRYLWGENLLGLHDNDIEDXXRIFRDAGEDASPVPRRRRRLTRTRSNED 263
>TC206034 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/MPE11_6),
partial (12%)
Length = 484
Score = 66.6 bits (161), Expect = 2e-11
Identities = 24/35 (68%), Positives = 31/35 (88%)
Frame = +1
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYIC 53
+E +E CP+CM+HPHNAVLL+CSSH+KGCR Y+C
Sbjct: 379 KEWEEARCPVCMEHPHNAVLLICSSHEKGCRPYMC 483
>BQ298390
Length = 424
Score = 60.1 bits (144), Expect = 2e-09
Identities = 28/38 (73%), Positives = 32/38 (83%)
Frame = +1
Query: 354 LQDNEVEEDLRIFNELVEDASHVPRRRRRLNRTRSNED 391
L DN++E+DLRIF + EDAS VPRRRRRL RTRSNED
Sbjct: 1 LHDNDIEDDLRIFRDAGEDASPVPRRRRRLTRTRSNED 114
>TC221861 similar to GB|AAM19869.1|20453261|AY097353 AT4g31410/F8F16_230
{Arabidopsis thaliana;} , partial (12%)
Length = 440
Score = 60.1 bits (144), Expect = 2e-09
Identities = 22/33 (66%), Positives = 28/33 (84%)
Frame = +3
Query: 22 DEVSCPICMDHPHNAVLLLCSSHDKGCRSYICD 54
D CPIC+D PHN+VLL CSS+DKGCR+++CD
Sbjct: 342 DNAVCPICLDFPHNSVLLQCSSYDKGCRAFVCD 440
>AW596336 similar to GP|18650602|gb| AT4g31410/F8F16_230 {Arabidopsis
thaliana}, partial (30%)
Length = 339
Score = 59.3 bits (142), Expect = 3e-09
Identities = 26/54 (48%), Positives = 36/54 (66%)
Frame = +3
Query: 229 RVHPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGD 282
R HP + PS +DP R+ W+ F++ E D++S I S IP VV+GDYV+E GD
Sbjct: 6 REHPHACPSKIDPVRQLDWKNFQQSSEIIDVLSTIHSEIPRGVVLGDYVIEYGD 167
>TC226761 similar to UP|Q9FYE4 (Q9FYE4) EF-hand calcium binding protein-like
(At5g04170), partial (63%)
Length = 1169
Score = 38.1 bits (87), Expect = 0.007
Identities = 31/88 (35%), Positives = 42/88 (47%), Gaps = 2/88 (2%)
Frame = -2
Query: 213 SCSFAGDYLELRRHARRVHPTSRPSDVDPTREQAW-QQFERQR-EYGDIVSAIQSAIPGA 270
S S +LE R H R V P RE+ W Q+ ER R + +A SA+
Sbjct: 457 SASVLVRHLEARHHVR----------VCPWRERRWHQRRERARVRRSRVAAAASSAVGVG 308
Query: 271 VVVGDYVLENGDGIGRLPPGGGREGSNG 298
+V+ +V+ G G GRL GGG G +G
Sbjct: 307 LVLRGFVVGGGVGGGRLGVGGGGVGLSG 224
>TC221669
Length = 784
Score = 32.3 bits (72), Expect = 0.38
Identities = 14/23 (60%), Positives = 18/23 (77%)
Frame = +3
Query: 260 VSAIQSAIPGAVVVGDYVLENGD 282
+S I S+ PGA+V+GDYVLE D
Sbjct: 3 ISTILSSTPGAMVLGDYVLEPND 71
>BI426954 similar to GP|5734709|gb|A Contains PF|00069 Eukaryotic protein
kinase domain. {Arabidopsis thaliana}, partial (4%)
Length = 423
Score = 32.0 bits (71), Expect = 0.50
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = -1
Query: 326 RARSSNGWSRHRRSSDRRRYLWGENLL 352
R R S GW R RR S R+R WG +L
Sbjct: 189 RRRGSGGWRRWRRCSARKRRSWGRAVL 109
>TC205626 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g75390), partial
(34%)
Length = 1184
Score = 30.0 bits (66), Expect = 1.9
Identities = 17/58 (29%), Positives = 26/58 (44%)
Frame = -3
Query: 286 RLPPGGGREGSNGNGNVPWLTTTTILFQMMDNTIEIVREPRARSSNGWSRHRRSSDRR 343
R G R S G+G V W ++ F + + R PR++ +G RH+R R
Sbjct: 273 RTSRGRRRRRSGGDGFVAWGCESS*KFFLKHLIVN*ARNPRSKRGSGL*RHKREKGNR 100
>TC229492 similar to UP|Q9FT86 (Q9FT86) Peroxin Pex14, partial (13%)
Length = 830
Score = 30.0 bits (66), Expect = 1.9
Identities = 14/38 (36%), Positives = 19/38 (49%)
Frame = +3
Query: 292 GREGSNGNGNVPWLTTTTILFQMMDNTIEIVREPRARS 329
G+ NG+ VPW T + + +DN IE P A S
Sbjct: 186 GKIQDNGDNTVPWWQTKNVRIKEIDNEIEYNGAPAASS 299
>BM307804 similar to GP|18766642|gb| NIMA-related protein kinase {Populus x
canescens}, partial (13%)
Length = 422
Score = 29.3 bits (64), Expect = 3.2
Identities = 21/57 (36%), Positives = 26/57 (44%), Gaps = 8/57 (14%)
Frame = +3
Query: 48 CRSYICDTSYRHSNCLD--------RFKKLRDNSKENPNLQSSLINTNNSSGNSFDI 96
C +CDTSY C D RFKK D K + Q++ T SS NS D+
Sbjct: 18 CTIKVCDTSYVRPGCTDAWQGIKRSRFKK-NDEDKSGSSDQNA--TTGASSHNSSDL 179
>TC215418 similar to GB|AAB90084.1|2649426|AE001023 A. fulgidus predicted
coding region AF1174 {Archaeoglobus fulgidus DSM 4304;} ,
partial (8%)
Length = 1700
Score = 29.3 bits (64), Expect = 3.2
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = -3
Query: 31 DHPHNAVLLLCSSHDKGCRSYICDTSYRHSNC 62
+HP + LLC S+D+ YI SYR NC
Sbjct: 1563 EHP*QILYLLCFSYDRDQILYILCFSYR*GNC 1468
>BE807100 weakly similar to SP|O60879|DIA2 Diaphanous protein homolog 2
(Diaphanous-related formin 2) (DRF2). [Human] {Homo
sapiens}, partial (3%)
Length = 385
Score = 29.3 bits (64), Expect = 3.2
Identities = 17/64 (26%), Positives = 23/64 (35%)
Frame = -2
Query: 279 ENGDGIGRLPPGGGREGSNGNGNVPWLTTTTILFQMMDNTIEIVREPRARSSNGWSRHRR 338
+ G G G PP GG +G G+G P + + PR G R +R
Sbjct: 177 KGGKGGGPTPPQGGEKGGKGDGGGP-----------------LFKNPRGEKGGGAKRKKR 49
Query: 339 SSDR 342
R
Sbjct: 48 GEPR 37
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.133 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,823,005
Number of Sequences: 63676
Number of extensions: 297343
Number of successful extensions: 2103
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 2071
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2092
length of query: 393
length of database: 12,639,632
effective HSP length: 99
effective length of query: 294
effective length of database: 6,335,708
effective search space: 1862698152
effective search space used: 1862698152
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC130200.9


