

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130200.5 - phase: 0
(312 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
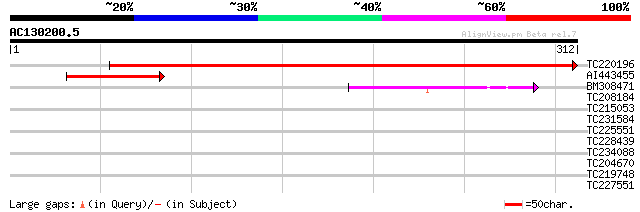
TC220196 similar to GB|AAO63361.1|28950875|BT005297 At5g48850 {A... 404 e-113
AI443455 similar to PIR|T45759|T457 MS5-like protein - Arabidops... 84 6e-17
BM308471 similar to PIR|T51865|T51 MS5-like protein [imported] -... 55 4e-08
TC208184 UP|Q39834 (Q39834) Clathrin heavy chain, complete 34 0.097
TC215053 homologue to UP|Q39834 (Q39834) Clathrin heavy chain, p... 31 0.63
TC231584 30 1.1
TC225551 similar to UP|Q9AYT8 (Q9AYT8) Mangrin, partial (68%) 30 1.1
TC228439 similar to UP|Q93VK5 (Q93VK5) At1g31800/68069_m00159, p... 29 2.4
TC234088 homologue to UP|Q94C23 (Q94C23) AT4g13510/T6G15_60, par... 28 5.3
TC204670 similar to UP|Q43499 (Q43499) Peroxidase precursor , pa... 27 9.1
TC219748 similar to GB|AAQ62420.1|34146824|BT010419 At3g12170 {A... 27 9.1
TC227551 homologue to UP|O65879 (O65879) Translation initiation ... 27 9.1
>TC220196 similar to GB|AAO63361.1|28950875|BT005297 At5g48850 {Arabidopsis
thaliana;} , partial (65%)
Length = 1185
Score = 404 bits (1039), Expect = e-113
Identities = 196/257 (76%), Positives = 229/257 (88%)
Frame = +3
Query: 56 QLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQ 115
+LV+KDPE AIV FWKAIN+GDKVDSALKDMAVVMKQLDR++EAIEAI+SFR LC+K SQ
Sbjct: 219 KLVEKDPEAAIVLFWKAINSGDKVDSALKDMAVVMKQLDRSDEAIEAIRSFRSLCSKQSQ 398
Query: 116 ESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEAFNGRTTKTARSHGKKFQVSIKQET 175
ESLDNVL+DLYKKCG+++EQIE+LKRKL+LIYQGEAFNG+ TKTARSHGKKFQVSIKQET
Sbjct: 399 ESLDNVLIDLYKKCGKIDEQIEMLKRKLKLIYQGEAFNGKLTKTARSHGKKFQVSIKQET 578
Query: 176 ARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANKALNLALCLMRQSRYEEAYLILEQV 235
+RLLGNLGWAYMQK NYMMAEVV++KAQ+ID D NKA NL LCL+RQ+RYEEA L+LE V
Sbjct: 579 SRLLGNLGWAYMQKMNYMMAEVVYRKAQIIDPDCNKACNLGLCLIRQARYEEAQLVLEDV 758
Query: 236 LQGKLPGSDEIKSRNRAEELLVELSANLPQPKFMDDLGLDDDLLKGIDGLLNVWSPTRSR 295
L+G LPGSD+ K+R RA++L EL + LP P F D LGLDD+ +KG++ L+N W P RS+
Sbjct: 759 LKGNLPGSDDSKARKRAQDLRTELRSMLPPPHFSDLLGLDDEFIKGLEQLMNEWGPIRSK 938
Query: 296 RLPIFEEISSFRDQLAC 312
RLPIFEEISSFRDQLAC
Sbjct: 939 RLPIFEEISSFRDQLAC 989
>AI443455 similar to PIR|T45759|T457 MS5-like protein - Arabidopsis thaliana,
partial (16%)
Length = 344
Score = 84.3 bits (207), Expect = 6e-17
Identities = 38/54 (70%), Positives = 45/54 (82%)
Frame = +3
Query: 32 KEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKD 85
+ + +HV HKVP GDTPYV+AK+ QLV+KDPE AI FW AINAGD+VDSALKD
Sbjct: 183 RSETFHVAHKVPIGDTPYVRAKNVQLVNKDPERAIPLFWAAINAGDRVDSALKD 344
>BM308471 similar to PIR|T51865|T51 MS5-like protein [imported] - Arabidopsis
thaliana (fragment), partial (36%)
Length = 418
Score = 55.1 bits (131), Expect = 4e-08
Identities = 34/109 (31%), Positives = 56/109 (51%), Gaps = 4/109 (3%)
Frame = +3
Query: 187 MQKTNYMMAEVVFKKAQMIDADANKALNLALCLMRQSRYEEA----YLILEQVLQGKLPG 242
MQ+ NY+ AE +++A I D NK NL +CLM+Q R EA Y + V+ G
Sbjct: 6 MQQNNYIEAEDAYRRALSIAPDNNKMCNLGICLMKQGRIGEAKETLYRVKPAVMDGPRGS 185
Query: 243 SDEIKSRNRAEELLVELSANLPQPKFMDDLGLDDDLLKGIDGLLNVWSP 291
+K+ RA+++L +L + + K +D + L + G ++W P
Sbjct: 186 DSHLKAYERAQQMLKDLESEM-MNKGVDRIE-QSRLFEAFLGSSSIWQP 326
>TC208184 UP|Q39834 (Q39834) Clathrin heavy chain, complete
Length = 5624
Score = 33.9 bits (76), Expect = 0.097
Identities = 32/97 (32%), Positives = 46/97 (46%), Gaps = 5/97 (5%)
Frame = +3
Query: 51 KAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFR--G 108
KA H +LV K VA+ N V+ AL ++ V + DR E+I+ +F G
Sbjct: 4443 KAGHLRLV-KPYMVAV-----QSNNVSAVNEALNEIYVEEEDYDRLRESIDLHDNFDQIG 4604
Query: 109 LCNK---HSQESLDNVLLDLYKKCGRVEEQIELLKRK 142
L K H + V +YKK GR ++ IEL +K
Sbjct: 4605 LAQKIETHELLEMRRVAAYIYKKAGRWKQSIELCHKK 4715
>TC215053 homologue to UP|Q39834 (Q39834) Clathrin heavy chain, partial (22%)
Length = 1446
Score = 31.2 bits (69), Expect = 0.63
Identities = 28/96 (29%), Positives = 44/96 (45%), Gaps = 5/96 (5%)
Frame = +2
Query: 51 KAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGL- 109
KA H +LV K +A+ N V+ AL ++ V + DR E+I+ +F +
Sbjct: 395 KAGHLRLV-KPYMIAV-----QSNNVSAVNEALNEIYVEEEDYDRLRESIDLHDNFDQIG 556
Query: 110 ----CNKHSQESLDNVLLDLYKKCGRVEEQIELLKR 141
KH + V +YKK GR ++ I L K+
Sbjct: 557 LAQKIEKHELLEMRRVAAYIYKKAGRWKQSIALSKK 664
>TC231584
Length = 780
Score = 30.4 bits (67), Expect = 1.1
Identities = 20/90 (22%), Positives = 40/90 (44%)
Frame = +2
Query: 66 IVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDL 125
I+ +WK + +KVDS+ D A + D ++ E++ + + LD+VL +
Sbjct: 485 IIRWWKWLQNSNKVDSSNTDSAAAHRNQDNPKKTEESVGNCPVCRKPFLAKDLDHVLALV 664
Query: 126 YKKCGRVEEQIELLKRKLRLIYQGEAFNGR 155
++ E L + +L+ + N R
Sbjct: 665 GSYSSQLSSDNEELNEEEKLLQSEDEINRR 754
>TC225551 similar to UP|Q9AYT8 (Q9AYT8) Mangrin, partial (68%)
Length = 1482
Score = 30.4 bits (67), Expect = 1.1
Identities = 14/47 (29%), Positives = 26/47 (54%)
Frame = +2
Query: 20 GISVLKKSSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAI 66
GI VL ++ KK D+Y I+ +GD ++ + + L +D +A+
Sbjct: 521 GICVLIQNKSEKKGDMYEAIYSFYFGDYGHISVQGSYLTYEDTYLAV 661
>TC228439 similar to UP|Q93VK5 (Q93VK5) At1g31800/68069_m00159, partial (66%)
Length = 1543
Score = 29.3 bits (64), Expect = 2.4
Identities = 19/72 (26%), Positives = 35/72 (48%), Gaps = 6/72 (8%)
Frame = +3
Query: 232 LEQVLQGKLPGSDEIKSRNRAEELLVELSANLPQPKFMDDLGLDDDLL------KGIDGL 285
++ VL + P +++K ++ E PQP + L+DD+L +G D
Sbjct: 666 VDSVLGDQYPTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSLEDDVLGEYPIKRGEDIF 845
Query: 286 LNVWSPTRSRRL 297
++VW+ RS +L
Sbjct: 846 ISVWNLHRSPKL 881
>TC234088 homologue to UP|Q94C23 (Q94C23) AT4g13510/T6G15_60, partial (8%)
Length = 408
Score = 28.1 bits (61), Expect = 5.3
Identities = 13/43 (30%), Positives = 23/43 (53%)
Frame = +3
Query: 28 SKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFW 70
S GK + +++ K+PYGD P ++ Q+ P + Y+W
Sbjct: 123 SSGKTKSQKNIMIKIPYGDNPR*QSFQ*QICPSVPR-RLFYYW 248
>TC204670 similar to UP|Q43499 (Q43499) Peroxidase precursor , partial (46%)
Length = 661
Score = 27.3 bits (59), Expect = 9.1
Identities = 17/80 (21%), Positives = 40/80 (49%)
Frame = -1
Query: 168 QVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANKALNLALCLMRQSRYEE 227
Q+S+ + +++L + + +NY + K I D LAL ++ + R+E
Sbjct: 334 QLSLWKHHSKVLDQIAEFHANSSNY------YSKINQI*FDYPY---LALLVLNRHRFEG 182
Query: 228 AYLILEQVLQGKLPGSDEIK 247
+++E+ ++G+ G D ++
Sbjct: 181 IVIVVEEEMKGRWKGQDHLR 122
>TC219748 similar to GB|AAQ62420.1|34146824|BT010419 At3g12170 {Arabidopsis
thaliana;} , partial (91%)
Length = 1074
Score = 27.3 bits (59), Expect = 9.1
Identities = 36/151 (23%), Positives = 59/151 (38%), Gaps = 25/151 (16%)
Frame = +1
Query: 4 QKEEKQKGHLCVLEMEGISVLKKSSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLV----- 58
+K K+K V E E I + LY V+ +K + +L
Sbjct: 46 RKMGKKKKRSRVSEDEEIESDTNPFDKNENSLYQVLGVEKTASQQEIKKAYYKLALRLHP 225
Query: 59 DKDP--EVAIVYFWKAINA----GDKVDSALKDMAVVMKQLDRAEEAIEAIKS-FRGLCN 111
DK+P E A F + N GD+ A+ D + + A + ++ +K FR +
Sbjct: 226 DKNPGDEEAKAKFQQLQNVIAILGDEEKRAVYDQTGFVDDAELAGDVVQNLKEYFRAMYK 405
Query: 112 KHSQESLD-------------NVLLDLYKKC 129
K ++ ++ N L+DLYKKC
Sbjct: 406 KVTEADIEEFEANYRGSDIEKNDLIDLYKKC 498
>TC227551 homologue to UP|O65879 (O65879) Translation initiation factor,
complete
Length = 1632
Score = 27.3 bits (59), Expect = 9.1
Identities = 12/36 (33%), Positives = 22/36 (60%)
Frame = +2
Query: 247 KSRNRAEELLVELSANLPQPKFMDDLGLDDDLLKGI 282
++ N AEE+ E + + +++G+ DDLL+GI
Sbjct: 140 RAANPAEEMDFETTEGVKAIASFEEMGIKDDLLRGI 247
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,340,729
Number of Sequences: 63676
Number of extensions: 122848
Number of successful extensions: 610
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 608
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 610
length of query: 312
length of database: 12,639,632
effective HSP length: 97
effective length of query: 215
effective length of database: 6,463,060
effective search space: 1389557900
effective search space used: 1389557900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC130200.5


