

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.7 + phase: 0
(359 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
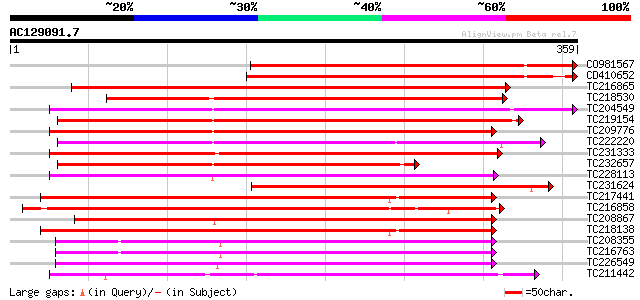
Score E
Sequences producing significant alignments: (bits) Value
CO981567 321 3e-88
CD410652 303 6e-83
TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 283 8e-77
TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partia... 276 1e-74
TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-l... 250 8e-67
TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 249 1e-66
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 240 6e-64
TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like prot... 234 3e-62
TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, parti... 233 8e-62
TC232657 weakly similar to UP|O22580 (O22580) Receptor-like seri... 231 3e-61
TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 231 4e-61
TC231624 similar to UP|KPEL_DROME (Q05652) Probable serine/threo... 231 5e-61
TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic em... 227 6e-60
TC216858 homologue to UP|Q76FZ8 (Q76FZ8) Brassinosteroid recepto... 226 9e-60
TC208867 similar to UP|Q9LXT2 (Q9LXT2) Serine/threonine-specific... 225 3e-59
TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis r... 224 4e-59
TC208355 UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, complete 223 1e-58
TC216763 UP|Q9LKY3 (Q9LKY3) Pti1 kinase-like protein, complete 221 3e-58
TC226549 UP|Q84P43 (Q84P43) Protein kinase Pti1, complete 217 7e-57
TC211442 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicite... 216 1e-56
>CO981567
Length = 812
Score = 321 bits (823), Expect = 3e-88
Identities = 161/207 (77%), Positives = 181/207 (86%)
Frame = -2
Query: 153 HEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTTGYLAPEYA 212
HEE HIVHRDIKASN+LLD+DFNPKIGDFG+AKLFPDDITHISTRIAGTTGYLAPEYA
Sbjct: 811 HEEHVPHIVHRDIKASNILLDRDFNPKIGDFGLAKLFPDDITHISTRIAGTTGYLAPEYA 632
Query: 213 LGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKWLALVDPEM 272
+GGQLT KADVYSFGVLILEIISGKSS+RTNW GS+K LLEWAWQL+EE K L LVDP+M
Sbjct: 631 MGGQLTMKADVYSFGVLILEIISGKSSARTNWGGSNKFLLEWAWQLYEEGKLLELVDPDM 452
Query: 273 EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLNDKQLTAPGLFNYDAGET 332
EFPEKEVI+Y+KVA FCTQAAA RRP+M+QVVDMLSK ++LN+KQLTAPGLF D+G +
Sbjct: 451 VEFPEKEVIRYMKVAFFCTQAAASRRPMMSQVVDMLSKNMRLNEKQLTAPGLFQ-DSGAS 275
Query: 333 SQKKSNPESLVYHTSSTQPSITEVTAR 359
SQKKS+ ES Y SS SIT++ R
Sbjct: 274 SQKKSSFESTGYQFSSNPSSITQLAPR 194
>CD410652
Length = 621
Score = 303 bits (777), Expect = 6e-83
Identities = 158/210 (75%), Positives = 178/210 (84%), Gaps = 1/210 (0%)
Frame = -1
Query: 151 YLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTTGYLAPE 210
+LHEEL+ IVHRDIKASNVLLD+DFNPKIGDFG+AKLFPDDITHISTRIAGTTGYLAPE
Sbjct: 621 FLHEELSPPIVHRDIKASNVLLDRDFNPKIGDFGLAKLFPDDITHISTRIAGTTGYLAPE 442
Query: 211 YALGGQLTKKADVYSFGVLILEIISGKSSS-RTNWDGSHKSLLEWAWQLHEEEKWLALVD 269
YALGGQLTKKAD+YSFGVLILEIISG+SS+ RTN GSHK LLEWAWQL+EE K L VD
Sbjct: 441 YALGGQLTKKADIYSFGVLILEIISGRSSARRTNGGGSHKFLLEWAWQLYEERKLLEFVD 262
Query: 270 PEMEEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLNDKQLTAPGLFNYDA 329
+MEEFPE+EVI+Y+KVALFCTQ+AA RRPLM QVVDMLSK IQLN+K+LTAPG F +
Sbjct: 261 QDMEEFPEEEVIRYMKVALFCTQSAANRRPLMIQVVDMLSKAIQLNEKELTAPGFFT-NE 85
Query: 330 GETSQKKSNPESLVYHTSSTQPSITEVTAR 359
GE+S+ SNP S +IT+VT R
Sbjct: 84 GESSRDDSNPVSSFI-------TITQVTPR 16
>TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (28%)
Length = 1358
Score = 283 bits (724), Expect = 8e-77
Identities = 149/279 (53%), Positives = 192/279 (68%), Gaps = 1/279 (0%)
Frame = +1
Query: 40 LGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNVKHSNLVELVG 99
+ NKIG GGFG VY+G DG IAVK LS S+QG REFL EI +S ++H +LV+L G
Sbjct: 1 VANKIGEGGFGPVYKGCFSDGTLIAVKQLSSKSRQGNREFLNEIGMISALQHPHLVKLYG 180
Query: 100 FCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLAYLHEELTQH 159
C++G +VYEY+EN +L AL G + +K+ W R IC+G A+GLAYLHEE
Sbjct: 181 CCVEGDQLLLVYEYMENNSLARALFGAEEHQIKLDWTTRYKICVGIARGLAYLHEESRLK 360
Query: 160 IVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTTGYLAPEYALGGQLTK 219
IVHRDIKA+NVLLD+D NPKI DFG+AKL +D THISTRIAGT GY+APEYA+ G LT
Sbjct: 361 IVHRDIKATNVLLDQDLNPKISDFGLAKLDEEDNTHISTRIAGTFGYMAPEYAMHGYLTD 540
Query: 220 KADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKWLALVDPEME-EFPEK 278
KADVYSFG++ LEII+G+S++ S+LEWA L E+ + LVD + EF ++
Sbjct: 541 KADVYSFGIVALEIINGRSNTIHRQKEESFSVLEWAHLLREKGDIMDLVDRRLGLEFNKE 720
Query: 279 EVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLNDK 317
E + IKVAL CT A RP M+ VV ML +I ++++
Sbjct: 721 EALVMIKVALLCTNVTAALRPTMSSVVSMLEGKIVVDEE 837
>TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partial (25%)
Length = 1190
Score = 276 bits (706), Expect = 1e-74
Identities = 139/254 (54%), Positives = 184/254 (71%)
Frame = +3
Query: 62 KIAVKPLSVGSKQGVREFLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHT 121
+IAVK LSVGS QG +F+TEI T+S V+H NLV+L G CI+G R +VYEY+EN +L
Sbjct: 3 EIAVKQLSVGSHQGKSQFITEIATISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQ 182
Query: 122 ALLGKKSLSVKMKWRERSTICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIG 181
AL GK + + W R IC+G A+GL YLHEE IVHRD+KASN+LLD + PKI
Sbjct: 183 ALFGK---CLTLNWSTRYDICLGVARGLTYLHEESRLRIVHRDVKASNILLDYELIPKIS 353
Query: 182 DFGMAKLFPDDITHISTRIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSR 241
DFG+AKL+ D THIST +AGT GYLAPEYA+ G LT+KADV+SFGV+ LE++SG+ +S
Sbjct: 354 DFGLAKLYDDKKTHISTGVAGTIGYLAPEYAMRGHLTEKADVFSFGVVALELVSGRPNSD 533
Query: 242 TNWDGSHKSLLEWAWQLHEEEKWLALVDPEMEEFPEKEVIKYIKVALFCTQAAARRRPLM 301
++ +G LLEWAWQLHE+ + LVD + EF E+EV + + +AL CTQ + RP M
Sbjct: 534 SSLEGEKVYLLEWAWQLHEKNCIIDLVDDRLSEFNEEEVKRVVGIALLCTQTSPTLRPSM 713
Query: 302 TQVVDMLSKEIQLN 315
++VV MLS +I+++
Sbjct: 714 SRVVAMLSGDIEVS 755
>TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-like protein,
partial (54%)
Length = 2032
Score = 250 bits (638), Expect = 8e-67
Identities = 143/337 (42%), Positives = 198/337 (58%), Gaps = 3/337 (0%)
Frame = +3
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
F + AT+ + NK+G GGFG VY+GTL G+ +AVK LS S QG EF E+
Sbjct: 792 FDFSTIEAATNKFSADNKLGEGGFGEVYKGTLSSGQVVAVKRLSKSSGQGGEEFKNEVVV 971
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGT 145
++ ++H NLV L+GFC+QG + +VYEYV N +L L + ++ W R I G
Sbjct: 972 VAKLQHRNLVRLLGFCLQGEEKILVYEYVPNKSLDYILFDPEK-QRELDWGRRYKIIGGI 1148
Query: 146 AKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST-RIAGTT 204
A+G+ YLHE+ I+HRD+KASN+LLD D NPKI DFGMA++F D T +T RI GT
Sbjct: 1149 ARGIQYLHEDSRLRIIHRDLKASNILLDGDMNPKISDFGMARIFGVDQTQGNTSRIVGTY 1328
Query: 205 GYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKW 264
GY+APEYA+ G+ + K+DVYSFGVL++EI+SGK +S + LL +AWQL ++
Sbjct: 1329 GYMAPEYAMHGEFSVKSDVYSFGVLLMEILSGKKNSSFYQTDGAEDLLSYAWQLWKDGTP 1508
Query: 265 LALVDPEM-EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML-SKEIQLNDKQLTAP 322
L L+DP + E + + EVI+ I + L C Q RP M +V ML S + L T P
Sbjct: 1509 LELMDPILRESYNQNEVIRSIHIGLLCVQEDPADRPTMATIVLMLDSNTVTLPTP--TQP 1682
Query: 323 GLFNYDAGETSQKKSNPESLVYHTSSTQPSITEVTAR 359
F + + + K P S SI+E+ R
Sbjct: 1683 AFFVHSGTDPNMPKELPFDQSIPMSVNDMSISEMDPR 1793
>TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (31%)
Length = 1363
Score = 249 bits (637), Expect = 1e-66
Identities = 133/297 (44%), Positives = 193/297 (64%), Gaps = 2/297 (0%)
Frame = +1
Query: 31 LSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNVK 90
++ ATDN+ L NKIG GGFG VY+G L G++IAVK LS S QG+ EF+TE+K ++ ++
Sbjct: 328 ITAATDNFLLNNKIGEGGFGPVYKGKLVGGQEIAVKRLSSLSGQGITEFITEVKLIAKLQ 507
Query: 91 HSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLA 150
H NLV+L+G CI+G + +VYEYV NG+L++ + + S + W R I +G A+GL
Sbjct: 508 HRNLVKLLGCCIKGQEKLLVYEYVVNGSLNSFIFDQIK-SKLLDWPRRFNIILGIARGLL 684
Query: 151 YLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST-RIAGTTGYLAP 209
YLH++ I+HRD+KASNVLLD+ NPKI DFGMA+ F D T +T R+ GT GY+AP
Sbjct: 685 YLHQDSRLRIIHRDLKASNVLLDEKLNPKISDFGMARAFGGDQTEGNTNRVVGTYGYMAP 864
Query: 210 EYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKWLALVD 269
EYA G + K+DV+SFG+L+LEI+ G + + +L+ +AW L +E+ L L+D
Sbjct: 865 EYAFDGNFSIKSDVFSFGILLLEIVCGIKNKSLCHENQTLNLVGYAWALWKEQNALQLID 1044
Query: 270 PEMEE-FPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLNDKQLTAPGLF 325
+++ EV++ I V+L C Q RP MT V+ ML E+ + + + PG F
Sbjct: 1045SGIKDSCVIPEVLRCIHVSLLCVQQYPEDRPTMTSVIQMLGSEMDMVEPK--EPGFF 1209
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein
, partial (47%)
Length = 1305
Score = 240 bits (613), Expect = 6e-64
Identities = 131/285 (45%), Positives = 178/285 (61%), Gaps = 2/285 (0%)
Frame = +2
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
F + AT + NK+G GGFG VY+G L G+++AVK LS S QG EF E++
Sbjct: 44 FDFSTIEAATQKFSEANKLGEGGFGEVYKGLLPSGQEVAVKRLSKISGQGGEEFKNEVEI 223
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGT 145
++ ++H NLV L+GFC++G + +VYE+V N +L L + + W R I G
Sbjct: 224 VAKLQHRNLVRLLGFCLEGEEKILVYEFVANKSLDYILFDPEK-QKSLDWTRRYKIVEGI 400
Query: 146 AKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST-RIAGTT 204
A+G+ YLHE+ I+HRD+KASNVLLD D NPKI DFGMA++F D T +T RI GT
Sbjct: 401 ARGIQYLHEDSRLKIIHRDLKASNVLLDGDMNPKISDFGMARIFGVDQTQANTNRIVGTY 580
Query: 205 GYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKW 264
GY++PEYA+ G+ + K+DVYSFGVLILEIISGK +S + LL +AW+L ++E
Sbjct: 581 GYMSPEYAMHGEYSAKSDVYSFGVLILEIISGKRNSSFYETDVAEDLLSYAWKLWKDEAP 760
Query: 265 LALVDPEM-EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
L L+D + E + EVI+ I + L C Q RP M VV ML
Sbjct: 761 LELMDQSLRESYTRNEVIRCIHIGLLCVQEDPIDRPTMASVVLML 895
>TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like protein kinase,
RLK3 precursor, partial (35%)
Length = 1304
Score = 234 bits (598), Expect = 3e-62
Identities = 131/315 (41%), Positives = 189/315 (59%), Gaps = 6/315 (1%)
Frame = +3
Query: 31 LSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNVK 90
+ AT+ + +IG GGFG VY+G L DGR+IAVK LS S QG EF EI ++ ++
Sbjct: 204 IEAATNKFSYEKRIGEGGFGVVYKGVLPDGREIAVKKLSKSSGQGANEFKNEILLIAKLQ 383
Query: 91 HSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLA 150
H NLV L+GFC++ + ++YE+V N +L L S ++ W ER I G A+G++
Sbjct: 384 HRNLVTLLGFCLEEHEKMLIYEFVSNKSLDYFLFDSHR-SKQLNWSERYKIIEGIAQGIS 560
Query: 151 YLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP-DDITHISTRIAGTTGYLAP 209
YLHE ++HRD+K SNVLLD + NPKI DFGMA++ D + + RI GT GY++P
Sbjct: 561 YLHEHSRLKVIHRDLKPSNVLLDSNMNPKISDFGMARIVAIDQLQGKTNRIVGTYGYMSP 740
Query: 210 EYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKWLALVD 269
EYA+ GQ ++K+DV+SFGV++LEIIS K ++R+ + H LL + W+ +E L + D
Sbjct: 741 EYAMHGQFSEKSDVFSFGVIVLEIISAKRNTRSVF-SDHDDLLSYTWEQWMDEAPLNIFD 917
Query: 270 PEME-EF-PEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSK---EIQLNDKQLTAPGL 324
+E EF EV+K I++ L C Q RP +TQV+ L+ E+ L K + G+
Sbjct: 918 QSIEAEFCDHSEVVKCIQIGLLCVQEKPDDRPKITQVISYLNSSITELPLPKKPIRQSGI 1097
Query: 325 FNYDAGETSQKKSNP 339
A S S P
Sbjct: 1098VQKIAVGESSSGSTP 1142
>TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partial (38%)
Length = 1009
Score = 233 bits (595), Expect = 8e-62
Identities = 121/289 (41%), Positives = 180/289 (61%), Gaps = 2/289 (0%)
Frame = +3
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
F+ EL LAT + N + GGFG+V++G L DG+ IAVK + S QG +EF +E++
Sbjct: 6 FTFSELQLATGGFSQANFLAEGGFGSVHRGVLPDGQVIAVKQYKLASTQGDKEFCSEVEV 185
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGT 145
LS +H N+V L+GFC+ R +VYEY+ NG+L + + +K ++W R I +G
Sbjct: 186 LSCAQHRNVVMLIGFCVDDGRRLLVYEYICNGSLDSHIYRRKQNV--LEWSARQKIAVGA 359
Query: 146 AKGLAYLHEELTQH-IVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTT 204
A+GL YLHEE IVHRD++ +N+LL DF +GDFG+A+ PD + TR+ GT
Sbjct: 360 ARGLRYLHEECRVGCIVHRDMRPNNILLTHDFEALVGDFGLARWQPDGDMGVETRVIGTF 539
Query: 205 GYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKW 264
GYLAPEYA GQ+T+KADVYSFG+++LE+++G+ + N + L EWA L E++
Sbjct: 540 GYLAPEYAQSGQITEKADVYSFGIVLLELVTGRKAVDINRPKGQQCLSEWARPLLEKQAT 719
Query: 265 LALVDPEMEE-FPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEI 312
L+DP + + ++EV + +K + C RP M+QV+ ML +I
Sbjct: 720 YKLIDPSLRNCYVDQEVYRMLKCSSLCIGRDPHLRPRMSQVLRMLEGDI 866
>TC232657 weakly similar to UP|O22580 (O22580) Receptor-like serine/threonine
kinase, partial (35%)
Length = 1208
Score = 231 bits (590), Expect = 3e-61
Identities = 115/229 (50%), Positives = 158/229 (68%)
Frame = +3
Query: 31 LSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNVK 90
L AT+ ++ NK+G+GG G+VY+G + DG +A+K LS + Q F TE+ +S +
Sbjct: 189 LEKATNYFNEANKLGQGGSGSVYKGVMPDGNTVAIKRLSYNTTQWAEHFFTEVNLISGIH 368
Query: 91 HSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLA 150
H NLV+L+G I GP +VYEYV N +LH +++ S + W R I +G A+G+A
Sbjct: 369 HKNLVKLLGCSITGPESLLVYEYVPNQSLHDHFSVRRT-SQPLTWEMRQKIILGIAEGMA 545
Query: 151 YLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTTGYLAPE 210
YLHEE I+HRDIK SN+LL++DF PKI DFG+A+LFP+D +HIST IAGT GY+APE
Sbjct: 546 YLHEESHVRIIHRDIKLSNILLEEDFTPKIADFGLARLFPEDKSHISTAIAGTLGYMAPE 725
Query: 211 YALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLH 259
Y + G+LT+KADVYSFGVL++EI+SGK S + S SLL+ W L+
Sbjct: 726 YIVRGKLTEKADVYSFGVLVIEIVSGKKISSYIMNSS--SLLQTVWSLY 866
>TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (53%)
Length = 1492
Score = 231 bits (589), Expect = 4e-61
Identities = 126/320 (39%), Positives = 183/320 (56%), Gaps = 36/320 (11%)
Frame = +1
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
F+ + +AT+++ NK+G+GGFG VY+G L DG+ IAVK LS S QG EF E+
Sbjct: 118 FNFDTIRVATEDFSDSNKLGQGGFGAVYRGRLSDGQMIAVKRLSRESSQGDTEFKNEVLL 297
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKK------------------ 127
++ ++H NLV L+GFC++G R ++YEYV N +L + G+
Sbjct: 298 VAKLQHRNLVRLLGFCLEGKERLLIYEYVPNKSLDYFIFGRS**VHN*TVALYFTSGSGQ 477
Query: 128 -----------------SLSVKMKWRERSTICIGTAKGLAYLHEELTQHIVHRDIKASNV 170
+ ++ W R I G A+GL YLHE+ I+HRD+KASN+
Sbjct: 478 RLNIHIPLKLMAVYADPTKKAQLNWEMRYKIITGVARGLLYLHEDSHLRIIHRDLKASNI 657
Query: 171 LLDKDFNPKIGDFGMAKLFPDDITHIST-RIAGTTGYLAPEYALGGQLTKKADVYSFGVL 229
LL+++ NPKI DFGMA+L D T +T RI GT GY+APEYA+ GQ + K+DV+SFGVL
Sbjct: 658 LLNEEMNPKIADFGMARLVLMDQTQANTNRIVGTYGYMAPEYAMHGQFSMKSDVFSFGVL 837
Query: 230 ILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKWLALVDPEMEEFPEKEVIKYIKVALF 289
+LEIISG +S + + LL +AW+ E + +VDP + E+++ I + L
Sbjct: 838 VLEIISGHKNSGIRHGENVEDLLSFAWRNWREGTAVKIVDPSLNNNSRNEMLRCIHIGLL 1017
Query: 290 CTQAAARRRPLMTQVVDMLS 309
C Q RP MT ++ ML+
Sbjct: 1018CVQENLADRPTMTTIMLMLN 1077
>TC231624 similar to UP|KPEL_DROME (Q05652) Probable serine/threonine-protein
kinase pelle , partial (7%)
Length = 841
Score = 231 bits (588), Expect = 5e-61
Identities = 113/195 (57%), Positives = 151/195 (76%), Gaps = 4/195 (2%)
Frame = +3
Query: 154 EELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTTGYLAPEYAL 213
+E +IVHRDIKASN+L D +FNPKIGDFG+AKLFPD++TH+STR+AGT GYLAPEYAL
Sbjct: 3 DEAQPNIVHRDIKASNILXDGNFNPKIGDFGLAKLFPDNVTHVSTRVAGTVGYLAPEYAL 182
Query: 214 GGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKWLALVDPEME 273
GQLTKKADVYSFG+L+LEIISGKSSS ++ + L+EWAW+L E + L LVD E+
Sbjct: 183 LGQLTKKADVYSFGILMLEIISGKSSSIAAFEEDYLVLVEWAWKLKGENRLLDLVDSELS 362
Query: 274 EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLNDKQLTAPGLFNYDA---- 329
E+ E V +++ VALFCTQ+AA+ RP M QV++MLSKE+ LN+K LT PG++ + +
Sbjct: 363 EYDESVVYRFLIVALFCTQSAAQHRPSMKQVLEMLSKEVHLNEKALTEPGIYRWHSNGKK 542
Query: 330 GETSQKKSNPESLVY 344
G + + S+ +++ Y
Sbjct: 543 GGSLNETSSSQAIKY 587
>TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic embryogenesis
receptor-like kinase 3 {Arabidopsis thaliana;} , partial
(54%)
Length = 1008
Score = 227 bits (579), Expect = 6e-60
Identities = 121/294 (41%), Positives = 187/294 (63%), Gaps = 5/294 (1%)
Frame = +2
Query: 20 LDNIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGV-RE 78
L ++ FS +EL +ATDN+ + +GRGGFG VY+G L DG +AVK L QG +
Sbjct: 101 LGQLKRFSLRELQVATDNFSNKHILGRGGFGKVYKGRLADGSLVAVKRLKEERTQGG*LQ 280
Query: 79 FLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRER 138
F TE++ +S H NL+ L GFC+ R +VY Y+ NG++ + L ++ + W ER
Sbjct: 281 FQTEVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERQESQPPLGWPER 460
Query: 139 STICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST 198
I +G+A+GLAYLH+ I+HRD+KA+N+LLD++F +GDFG+AKL TH++T
Sbjct: 461 KRIALGSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTT 640
Query: 199 RIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSS---SRTNWDGSHKSLLEWA 255
+ GT G++APEY G+ ++K DV+ +GV++LE+I+G+ + +R D LL+W
Sbjct: 641 AVRGTIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLAND-DDVMLLDWV 817
Query: 256 WQLHEEEKWLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
L ++ K LVD +++ + ++EV + I+VAL CTQ + RP M++VV ML
Sbjct: 818 KGLLKDRKLETLVDADLQGSYNDEEVEQLIQVALLCTQGSPMERPKMSEVVRML 979
>TC216858 homologue to UP|Q76FZ8 (Q76FZ8) Brassinosteroid receptor, partial
(29%)
Length = 1320
Score = 226 bits (577), Expect = 9e-60
Identities = 130/309 (42%), Positives = 190/309 (61%), Gaps = 4/309 (1%)
Frame = +1
Query: 9 VSVFFCCAGYPLDNIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPL 68
+S+F PL R + +L AT+ +H + IG GGFG VY+ LKDG +A+K L
Sbjct: 4 LSIFLATFEKPL---RKLTFADLLDATNGFHNDSLIGSGGFGDVYKAQLKDGSVVAIKKL 174
Query: 69 SVGSKQGVREFLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKS 128
S QG REF E++T+ +KH NLV L+G+C G R +VYEY++ G+L L +K
Sbjct: 175 IHVSGQGDREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDQKK 354
Query: 129 LSVKMKWRERSTICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKL 188
+K+ W R I IG A+GLA+LH HI+HRD+K+SNVLLD++ ++ DFGMA+L
Sbjct: 355 AGIKLNWAIRRKIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARL 534
Query: 189 FPDDITHIS-TRIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGS 247
TH+S + +AGT GY+ PEY + + K DVYS+GV++LE+++GK + + D
Sbjct: 535 MSAMDTHLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPT-DSADFG 711
Query: 248 HKSLLEWAWQLHEEEKWLALVDPE-MEEFP--EKEVIKYIKVALFCTQAAARRRPLMTQV 304
+L+ W Q H + K + DPE M+E P E E+++++K+A+ C RRP M QV
Sbjct: 712 DNNLVGWVKQ-HAKLKISDIFDPELMKEDPNLEMELLQHLKIAVSCLDDRPWRRPTMIQV 888
Query: 305 VDMLSKEIQ 313
+ M KEIQ
Sbjct: 889 MAMF-KEIQ 912
>TC208867 similar to UP|Q9LXT2 (Q9LXT2) Serine/threonine-specific protein
kinase-like protein, partial (72%)
Length = 1118
Score = 225 bits (573), Expect = 3e-59
Identities = 119/273 (43%), Positives = 168/273 (60%), Gaps = 6/273 (2%)
Frame = +1
Query: 42 NKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNVKHSNLVELVGFC 101
N IG GGFG VY+G L DGRK+A+K + KQG EF E++ LS + L+ L+G+C
Sbjct: 10 NVIGHGGFGLVYRGVLNDGRKVAIKFMDQAGKQGEEEFKVEVELLSRLHSPYLLALLGYC 189
Query: 102 IQGPNRTVVYEYVENGNLHTALLGKKS---LSVKMKWRERSTICIGTAKGLAYLHEELTQ 158
++ +VYE++ NG L L + VK+ W R I + AKGL YLHE ++
Sbjct: 190 SDSNHKLLVYEFMANGGLQEHLYPVSNSIITPVKLDWETRLRIALEAAKGLEYLHEHVSP 369
Query: 159 HIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDI-THISTRIAGTTGYLAPEYALGGQL 217
++HRD K+SN+LLDK F+ K+ DFG+AKL PD H+STR+ GT GY+APEYAL G L
Sbjct: 370 PVIHRDFKSSNILLDKKFHAKVSDFGLAKLGPDRAGGHVSTRVLGTQGYVAPEYALTGHL 549
Query: 218 TKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQ-LHEEEKWLALVDPEME-EF 275
T K+DVYS+GV++LE+++G+ L+ WA L + EK + ++DP +E ++
Sbjct: 550 TTKSDVYSYGVVLLELLTGRVPVDMKRPPGEGVLVSWALPLLTDREKVVKIMDPSLEGQY 729
Query: 276 PEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
KEV++ +A C Q A RPLM VV L
Sbjct: 730 SMKEVVQVAAIAAMCVQPEADYRPLMADVVQSL 828
>TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis receptor
kinase 1, partial (60%)
Length = 1444
Score = 224 bits (572), Expect = 4e-59
Identities = 119/294 (40%), Positives = 185/294 (62%), Gaps = 5/294 (1%)
Frame = +1
Query: 20 LDNIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVR-E 78
L ++ FS +EL +ATD++ N +GRGGFG VY+G L DG +AVK L G +
Sbjct: 103 LGQLKRFSLRELQVATDSFSNKNILGRGGFGKVYKGRLADGSLVAVKRLKEERTPGGELQ 282
Query: 79 FLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRER 138
F TE++ +S H NL+ L GFC+ R +VY Y+ NG++ + L + + W R
Sbjct: 283 FQTEVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPPYQEPLDWPTR 462
Query: 139 STICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST 198
+ +G+A+GL+YLH+ I+HRD+KA+N+LLD++F +GDFG+AKL TH++T
Sbjct: 463 KRVALGSARGLSYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTT 642
Query: 199 RIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSS---SRTNWDGSHKSLLEWA 255
+ GT G++APEY G+ ++K DV+ +G+++LE+I+G+ + +R D LL+W
Sbjct: 643 AVRGTIGHIAPEYLSTGKSSEKTDVFGYGIMLLELITGQRAFDLARLAND-DDVMLLDWV 819
Query: 256 WQLHEEEKWLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
L +E+K LVDP+++ + E EV + I+VAL CTQ + RP M++VV ML
Sbjct: 820 KGLLKEKKLEMLVDPDLQTNYIETEVEQLIQVALLCTQGSPMDRPKMSEVVRML 981
>TC208355 UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, complete
Length = 1536
Score = 223 bits (567), Expect = 1e-58
Identities = 123/286 (43%), Positives = 168/286 (58%), Gaps = 7/286 (2%)
Frame = +2
Query: 30 ELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNV 89
EL TDN+ IG G +G VYQ TLK+GR + +K L S Q EFL+++ +S +
Sbjct: 380 ELKPLTDNFGSKCFIGEGAYGKVYQATLKNGRAVVIKKLD-SSNQPEHEFLSQVSIVSRL 556
Query: 90 KHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVK-----MKWRERSTICIG 144
KH N+VELV +C+ GP R + YEY G+LH L G+K + + W +R I +G
Sbjct: 557 KHENVVELVNYCVDGPFRALAYEYAPKGSLHDILHGRKGVKGAQPGPVLSWAQRVKIAVG 736
Query: 145 TAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHI-STRIAGT 203
A+GL YLHE+ HI+HR IK+SN+LL D KI DF ++ PD + STR+ GT
Sbjct: 737 AARGLEYLHEKAEIHIIHRYIKSSNILLFDDDVAKIADFDLSNQAPDAAARLHSTRVLGT 916
Query: 204 TGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEK 263
GY APEYA+ GQLT K+DVYSFGV++LE+++G+ +SL+ WA E+K
Sbjct: 917 FGYHAPEYAMTGQLTSKSDVYSFGVILLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDK 1096
Query: 264 WLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
VD ++ E+P K V K VA C Q A RP M+ +V L
Sbjct: 1097VKQCVDVRLKGEYPSKSVAKMAAVAALCVQYEAEFRPNMSIIVKAL 1234
>TC216763 UP|Q9LKY3 (Q9LKY3) Pti1 kinase-like protein, complete
Length = 1634
Score = 221 bits (564), Expect = 3e-58
Identities = 121/286 (42%), Positives = 168/286 (58%), Gaps = 7/286 (2%)
Frame = +1
Query: 30 ELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNV 89
EL TDN+ IG G +G VYQ TLK+G + +K L S Q +EFL+++ +S +
Sbjct: 370 ELKSLTDNFGSKYFIGEGAYGKVYQATLKNGHAVVIKKLD-SSNQPEQEFLSQVSIVSRL 546
Query: 90 KHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVK-----MKWRERSTICIG 144
KH N+VELV +C+ GP R + YEY G+LH L G+K + + W +R I +G
Sbjct: 547 KHENVVELVNYCVDGPFRALAYEYAPKGSLHDILHGRKGVKGAQPGPVLSWAQRVKIAVG 726
Query: 145 TAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHI-STRIAGT 203
A+GL YLHE+ HI+HR IK+SN+LL D K+ DF ++ PD + STR+ GT
Sbjct: 727 AARGLEYLHEKAEIHIIHRYIKSSNILLFDDDVAKVADFDLSNQAPDAAARLHSTRVLGT 906
Query: 204 TGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEK 263
GY APEYA+ GQLT K+DVYSFGV++LE+++G+ +SL+ WA E+K
Sbjct: 907 FGYHAPEYAMTGQLTSKSDVYSFGVILLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDK 1086
Query: 264 WLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
VD ++ E+P K V K VA C Q A RP M+ +V L
Sbjct: 1087VKQCVDVRLKGEYPSKSVAKMAAVAALCVQYEAEFRPNMSIIVKAL 1224
>TC226549 UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1627
Score = 217 bits (552), Expect = 7e-57
Identities = 120/287 (41%), Positives = 173/287 (59%), Gaps = 8/287 (2%)
Frame = +3
Query: 30 ELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSK-QGVREFLTEIKTLSN 88
EL TDN+ IG G +G VY TL +G+ +AVK L V S+ + EFLT++ +S
Sbjct: 384 ELKEKTDNFGSKALIGEGSYGRVYYATLNNGKAVAVKKLDVSSEPESNNEFLTQVSMVSR 563
Query: 89 VKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLS-----VKMKWRERSTICI 143
+K+ N VEL G+C++G R + YE+ G+LH L G+K + + W +R I +
Sbjct: 564 LKNGNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAV 743
Query: 144 GTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHI-STRIAG 202
A+GL YLHE++ I+HRDI++SNVL+ +D+ KI DF ++ PD + STR+ G
Sbjct: 744 DAARGLEYLHEKVQPPIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTRVLG 923
Query: 203 TTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEE 262
T GY APEYA+ GQLT+K+DVYSFGV++LE+++G+ +SL+ WA E+
Sbjct: 924 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 1103
Query: 263 KWLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
K VDP+++ E+P K V K VA C Q A RP M+ VV L
Sbjct: 1104KVKQCVDPKLKGEYPPKGVAKLGAVAALCVQYEAEFRPNMSIVVKAL 1244
>TC211442 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicited protein
264, partial (73%)
Length = 1047
Score = 216 bits (551), Expect = 1e-56
Identities = 128/321 (39%), Positives = 189/321 (58%), Gaps = 11/321 (3%)
Frame = +2
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKD-------GRKIAVKPLSVGSKQGVRE 78
F+ +EL AT+++ N +G GGFG VY+G + D + +AVK L + QG RE
Sbjct: 35 FTLEELREATNSFSWSNMLGEGGFGPVYKGFVDDKLRSGLKAQTVAVKRLDLDGLQGHRE 214
Query: 79 FLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRER 138
+L EI L ++H +LV+L+G+C + +R ++YEY+ G+L L + S M W R
Sbjct: 215 WLAEIIFLGQLRHPHLVKLIGYCYEDEHRLLMYEYMPRGSLENQLF--RRYSAAMPWSTR 388
Query: 139 STICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPD-DITHIS 197
I +G AKGLA+LHE + +++RD KASN+LLD DF K+ DFG+AK P+ + TH++
Sbjct: 389 MKIALGAAKGLAFLHEA-DKPVIYRDFKASNILLDSDFTAKLSDFGLAKDGPEGEDTHVT 565
Query: 198 TRIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQ 257
TRI GT GY APEY + G LT K+DVYS+GV++LE+++G+ + KSL+EWA
Sbjct: 566 TRIMGTQGYAAPEYIMTGHLTTKSDVYSYGVVLLELLTGRRVVDKSRSNEGKSLVEWARP 745
Query: 258 -LHEEEKWLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLN 315
L +++K ++D +E +FP K +K +A C RP M+ V+ +L L
Sbjct: 746 LLRDQKKVYNIIDRRLEGQFPMKGAMKVAMLAFKCLSHHPNARPTMSDVIKVLE---PLQ 916
Query: 316 DKQLTAPGLFNYDA-GETSQK 335
D G F Y A ET K
Sbjct: 917 DYDDVFIGPFVYVAVSETGDK 979
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,845,516
Number of Sequences: 63676
Number of extensions: 215529
Number of successful extensions: 2897
Number of sequences better than 10.0: 1013
Number of HSP's better than 10.0 without gapping: 2151
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2178
length of query: 359
length of database: 12,639,632
effective HSP length: 98
effective length of query: 261
effective length of database: 6,399,384
effective search space: 1670239224
effective search space used: 1670239224
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC129091.7


