

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
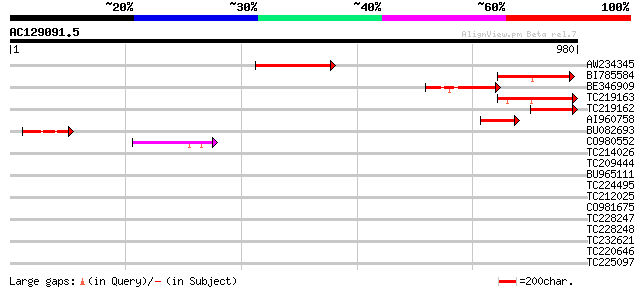
Query= AC129091.5 - phase: 0 /pseudo
(980 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW234345 217 2e-56
BI785584 191 1e-48
BE346909 169 4e-42
TC219163 161 1e-39
TC219162 128 1e-29
AI960758 112 1e-24
BU082693 112 1e-24
CO980552 95 1e-19
TC214026 similar to UP|Q6G0R7 (Q6G0R7) GTP-binding protein typA,... 34 0.36
TC209444 similar to GB|AAP21178.1|30102520|BT006370 At3g08040/T8... 33 0.80
BU965111 30 5.2
TC224495 weakly similar to GB|AAO63306.1|28950765|BT005242 At3g2... 30 5.2
TC212025 similar to UP|Q8H934 (Q8H934) Phototropin, partial (27%) 30 6.8
CO981675 30 6.8
TC228247 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific le... 30 6.8
TC228248 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific le... 30 6.8
TC232621 similar to UP|Q8LPN7 (Q8LPN7) AT3g19950/MPN9_19, partia... 29 8.9
TC220646 weakly similar to UP|Q70EW1 (Q70EW1) Diamine oxidase (C... 29 8.9
TC225097 homologue to UP|Q9M6R6 (Q9M6R6) RNA helicase, partial (... 29 8.9
>AW234345
Length = 418
Score = 217 bits (552), Expect = 2e-56
Identities = 101/137 (73%), Positives = 115/137 (83%)
Frame = +1
Query: 426 AFFQGRVRIFSGSNFEPVTNYEINVGSSISVPAFSATSCCSASVWHDTSKGEAMLKIIRV 485
AF +G V IFSG NF PV NY+I+VGS+I+ PAFS+TSCCSASVWHDTSK + +LKIIRV
Sbjct: 7 AFLRGEVHIFSGPNFAPVDNYQISVGSAIAAPAFSSTSCCSASVWHDTSKDQTILKIIRV 186
Query: 486 LPPPFPIGQEKATSSTWEHAISERFWFSLLVGVDWWDVVGCTQRAAEEGIVSVNGVIAVL 545
LP P Q KA SS WE AI+ERFW+SLLVGV+WWD VGCTQ AAE+GIVS+N VIAVL
Sbjct: 187 LPHAIPTSQVKANSSNWERAIAERFWWSLLVGVNWWDAVGCTQSAAEDGIVSLNSVIAVL 366
Query: 546 DADFHSLPTAQHRQQYC 562
DADFHSLP+AQHRQ YC
Sbjct: 367 DADFHSLPSAQHRQHYC 417
>BI785584
Length = 421
Score = 191 bits (486), Expect = 1e-48
Identities = 97/140 (69%), Positives = 110/140 (78%), Gaps = 6/140 (4%)
Frame = +1
Query: 843 GSYTVLPEVVEASLGPYMQNMPRLGDADDTGLLLRELQLHPPAEEWHQLNMFVRPCTDS- 901
G+Y+V PEVVEASLGP+MQNMPR AD GLLLREL+LHPPAEEWH+ NMF P +D
Sbjct: 1 GNYSVQPEVVEASLGPHMQNMPRPRGADAAGLLLRELELHPPAEEWHRRNMFGAPWSDPE 180
Query: 902 -----NDTPKPFRSNPLDSRSLESNDIDYGTNGLRPKKRRMIERDAAFGLNTSLGLGAYL 956
NDTPK S+PLD SLE D+ YGT+ L P+KRRM ERDAAFGLNTS+GLG YL
Sbjct: 181 DVDCVNDTPKLVNSDPLDFSSLEHCDVYYGTHRLWPRKRRMSERDAAFGLNTSVGLGGYL 360
Query: 957 GIMGSRRDVITTSWKTGLEG 976
GIMGSRRDV+T +WKTGLEG
Sbjct: 361 GIMGSRRDVVTATWKTGLEG 420
>BE346909
Length = 435
Score = 169 bits (429), Expect = 4e-42
Identities = 87/151 (57%), Positives = 108/151 (70%), Gaps = 23/151 (15%)
Frame = +3
Query: 720 GDCHFLHRLCQLLFFCFFFKRSQLARYKSGLRRTAETSL--------------------- 758
GDCHFLHRLCQLL FCFFF+R+QL RY + RT++T++
Sbjct: 3 GDCHFLHRLCQLLLFCFFFRRTQLPRY---INRTSDTNIQKPQSNTPAPGKVEEIAKPVS 173
Query: 759 --VRSNDGQTGRDRQIVPGSKGGEEPSPGPVRLGNGNAGQGYSVEEVKVIFQVLLDLCRR 816
V+S+DGQTGR G+KG EE G RLG+GNAGQGY+ EEVKV+F +L+DLCRR
Sbjct: 174 AVVKSDDGQTGRT-----GAKGAEEVPSGRSRLGSGNAGQGYTFEEVKVLFMMLMDLCRR 338
Query: 817 TSGLQHPLPVSQVGSSNIQVQLHYIEGSYTV 847
T+GLQHPLPVSQVGS+NIQV+LHYI+G+Y+V
Sbjct: 339 TAGLQHPLPVSQVGSNNIQVRLHYIDGNYSV 431
>TC219163
Length = 895
Score = 161 bits (408), Expect = 1e-39
Identities = 89/163 (54%), Positives = 102/163 (61%), Gaps = 26/163 (15%)
Frame = +2
Query: 844 SYTVLPEVVEASLGPY--------------------MQNMPRLGDADDTGLLLRELQLHP 883
+YTVLPEV + L P MQNMPR AD GLLL EL+LHP
Sbjct: 2 NYTVLPEVXKQPLAPICSD*XXCE*ILPLKINKKMAMQNMPRPRGADAAGLLLCELELHP 181
Query: 884 PAEEWHQLNMFVRPCTD------SNDTPKPFRSNPLDSRSLESNDIDYGTNGLRPKKRRM 937
PAEEWH+ NMF P +D +ND +PLDS SLE D+ YG NGL + RRM
Sbjct: 182 PAEEWHRRNMFGGPWSDPDVLDFANDALNLVSLHPLDSSSLEYCDVYYGANGLWSR*RRM 361
Query: 938 IERDAAFGLNTSLGLGAYLGIMGSRRDVITTSWKTGLEGVWYK 980
ERDAAFGLNT +GLGAYLGIMGSRRD +T WKTG+EG+WYK
Sbjct: 362 SERDAAFGLNTYVGLGAYLGIMGSRRDAVTALWKTGIEGIWYK 490
>TC219162
Length = 700
Score = 128 bits (321), Expect = 1e-29
Identities = 60/80 (75%), Positives = 67/80 (83%)
Frame = +3
Query: 901 SNDTPKPFRSNPLDSRSLESNDIDYGTNGLRPKKRRMIERDAAFGLNTSLGLGAYLGIMG 960
+ND PK NPLDS SLE+ D+ YG NGL P+KRRM ERDAAFGLNTS+GLGAYLGIMG
Sbjct: 24 ANDAPKLISLNPLDSSSLENCDVYYGANGLWPRKRRMSERDAAFGLNTSVGLGAYLGIMG 203
Query: 961 SRRDVITTSWKTGLEGVWYK 980
SRRDV+T WKTGLEG+WYK
Sbjct: 204 SRRDVVTALWKTGLEGIWYK 263
>AI960758
Length = 204
Score = 112 bits (279), Expect = 1e-24
Identities = 54/67 (80%), Positives = 62/67 (91%)
Frame = +2
Query: 815 RRTSGLQHPLPVSQVGSSNIQVQLHYIEGSYTVLPEVVEASLGPYMQNMPRLGDADDTGL 874
RRT+GLQHPLPVSQVGS+NIQV+LHYI+G+YTVLPEVVEA+LGP+MQNMPR AD GL
Sbjct: 2 RRTAGLQHPLPVSQVGSNNIQVRLHYIDGNYTVLPEVVEAALGPHMQNMPRPRGADAAGL 181
Query: 875 LLRELQL 881
LLREL+L
Sbjct: 182 LLRELEL 202
>BU082693
Length = 429
Score = 112 bits (279), Expect = 1e-24
Identities = 58/89 (65%), Positives = 64/89 (71%)
Frame = +2
Query: 22 MEEEEVEAGTVFDISLEQSPSTSLHKITVHESMRSNFSAVAWCAKLNVIACATETCVNGI 81
MEE+ V TVF I L+Q S LHK++V E R NFSAV+WC KLN IACA ETC I
Sbjct: 167 MEEDPVNPATVFSIRLKQPRSNLLHKMSVPELCR-NFSAVSWCGKLNAIACAAETCAR-I 340
Query: 82 PRSSVNPWFWIPIHIVIPERPTEIAAFNV 110
P S+ NP FWIPIHIVIPERPTE A FNV
Sbjct: 341 PSSTANPPFWIPIHIVIPERPTECAVFNV 427
>CO980552
Length = 685
Score = 95.1 bits (235), Expect = 1e-19
Identities = 85/171 (49%), Positives = 95/171 (54%), Gaps = 24/171 (14%)
Frame = -3
Query: 212 LRMQHRPNGFALVKG**NVGQVGL*LLMLSSQTVVCYTWPVC*LLVQLPL*YGWFCLDRE 271
LRM +GFALVK VGQV L +MLS QTVV V LL++ PL +G L E
Sbjct: 683 LRMAQHRSGFALVKDFWVVGQVALWPVMLS*QTVVPCM*QVYQLLIRPPLSFGRLHLGLE 504
Query: 272 MVWNFLPRQV-LQLQHLLAHPTGLVLPLYLPFCSVGKI------------------NLYH 312
MV L RQV L + HLLA PTG+VL L L VGKI LY
Sbjct: 503 MVSR*LQRQVPLVVSHLLALPTGMVLRL*LHIYLVGKIIYYLKQSKGENRQTKTLLMLYL 324
Query: 313 FTAQQFQIFQHM*VLKL-----HPLPSVVV*QG*PLIQRVVVL**LLL*LR 358
+ A QF+IFQHM*VLKL PL V+V*Q PLIQ V V **LL *LR
Sbjct: 323 YIAHQFRIFQHM*VLKLQLNLQQPLHGVLV*QQWPLIQLVRVQ**LL**LR 171
>TC214026 similar to UP|Q6G0R7 (Q6G0R7) GTP-binding protein typA, partial
(14%)
Length = 501
Score = 33.9 bits (76), Expect = 0.36
Identities = 28/76 (36%), Positives = 31/76 (39%), Gaps = 12/76 (15%)
Frame = +3
Query: 725 LHRLCQLLFFCFFFKRSQ---LARYKSGLRRTAETSLVRSNDG---------QTGRDRQI 772
LH+ L FF FFF S LA + LRR + RSN Q GRDR
Sbjct: 87 LHQKILLFFFFFFFSLSSFIPLALLLTRLRRRSRHCPCRSNHRLLPRPQPPPQPGRDRSC 266
Query: 773 VPGSKGGEEPSPGPVR 788
P P P PVR
Sbjct: 267 RPRQDHPHGPPPPPVR 314
>TC209444 similar to GB|AAP21178.1|30102520|BT006370 At3g08040/T8G24.8
{Arabidopsis thaliana;} , partial (37%)
Length = 931
Score = 32.7 bits (73), Expect = 0.80
Identities = 15/31 (48%), Positives = 18/31 (57%)
Frame = +1
Query: 263 YGWFCLDREMVWNFLPRQVLQLQHLLAHPTG 293
+GW CL MVW L RQ LQ+ L+ TG
Sbjct: 121 FGWQCLFLRMVWLLLGRQFLQVHLLIRTSTG 213
>BU965111
Length = 445
Score = 30.0 bits (66), Expect = 5.2
Identities = 11/35 (31%), Positives = 21/35 (59%)
Frame = +1
Query: 509 RFWFSLLVGVDWWDVVGCTQRAAEEGIVSVNGVIA 543
R+W LVG+ WW+ + C A + + S++ +I+
Sbjct: 136 RWWHWCLVGLGWWNHLHCLHLHAHDTLFSLSALIS 240
>TC224495 weakly similar to GB|AAO63306.1|28950765|BT005242 At3g26470
{Arabidopsis thaliana;} , partial (35%)
Length = 751
Score = 30.0 bits (66), Expect = 5.2
Identities = 15/43 (34%), Positives = 20/43 (45%)
Frame = -3
Query: 145 WTQPSRGPPNLVIDTNCWQREHEWRQETAVVTKWLSGASPLDG 187
WT+ SR P + C +R WR+ + SG SP DG
Sbjct: 293 WTRSSRACPFFLRGGGCGRRSSGWRRGPCGRARRRSGESPRDG 165
>TC212025 similar to UP|Q8H934 (Q8H934) Phototropin, partial (27%)
Length = 978
Score = 29.6 bits (65), Expect = 6.8
Identities = 13/33 (39%), Positives = 20/33 (60%)
Frame = +1
Query: 751 RRTAETSLVRSNDGQTGRDRQIVPGSKGGEEPS 783
+R AE LV D +TG+ + + + GGEEP+
Sbjct: 19 QRAAEWGLVLRTDTETGKPQGVAVRNSGGEEPN 117
>CO981675
Length = 810
Score = 29.6 bits (65), Expect = 6.8
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = -3
Query: 722 CHFLHRLCQLLFFCFFFKRSQLARYK 747
C+FLH L ++ F FFF QL ++
Sbjct: 169 CYFLHHLMRIFIFAFFFLFFQLGSFR 92
>TC228247 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin
PP2-like protein, partial (46%)
Length = 1124
Score = 29.6 bits (65), Expect = 6.8
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = +2
Query: 722 CHFLHRLCQLLFFCFFFK 739
CH LH Q + FCFFFK
Sbjct: 926 CHGLHAQLQSILFCFFFK 979
>TC228248 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin
PP2-like protein, partial (28%)
Length = 519
Score = 29.6 bits (65), Expect = 6.8
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = +2
Query: 722 CHFLHRLCQLLFFCFFFK 739
CH LH Q + FCFFFK
Sbjct: 359 CHGLHAQLQSILFCFFFK 412
>TC232621 similar to UP|Q8LPN7 (Q8LPN7) AT3g19950/MPN9_19, partial (33%)
Length = 1315
Score = 29.3 bits (64), Expect = 8.9
Identities = 28/95 (29%), Positives = 44/95 (45%), Gaps = 5/95 (5%)
Frame = -1
Query: 776 SKGGEEPSPGPVRLGNGNAGQGYSVEEVKVIFQVLLDLCRRTSGLQHPLPV----SQVGS 831
SKGGE +PG R +G S ++ + L + +S + PL V S+ G
Sbjct: 289 SKGGEAAAPGRTRGMVAAPARGKSGKKGLGGLGLGLGIGNSSSSRKPPLQVGQRRSEEGE 110
Query: 832 SNIQ-VQLHYIEGSYTVLPEVVEASLGPYMQNMPR 865
I+ V+LH+ + + PEV L P ++M R
Sbjct: 109 GEIETVRLHWWQKYWRGSPEVAPPPLSPPAEDM*R 5
>TC220646 weakly similar to UP|Q70EW1 (Q70EW1) Diamine oxidase (Copper amino
oxidase) , partial (23%)
Length = 707
Score = 29.3 bits (64), Expect = 8.9
Identities = 15/53 (28%), Positives = 28/53 (52%), Gaps = 2/53 (3%)
Frame = +2
Query: 171 ETAVVTKWLSGASPLDGPSFSVSVL--YHNQERFNFIGPKGLLLRMQHRPNGF 221
+ A V KWLS + +FS++++ ++ N + P+ ++L H NGF
Sbjct: 110 DKAAVLKWLSSGARTPRNAFSIALINGQIHELTVNLLSPRNVVLDKIHTGNGF 268
>TC225097 homologue to UP|Q9M6R6 (Q9M6R6) RNA helicase, partial (15%)
Length = 757
Score = 29.3 bits (64), Expect = 8.9
Identities = 14/34 (41%), Positives = 20/34 (58%)
Frame = +2
Query: 903 DTPKPFRSNPLDSRSLESNDIDYGTNGLRPKKRR 936
DTPKP ++SL D+D G +G + KKR+
Sbjct: 143 DTPKPIAKKKTKTQSLTDPDLD-GVSGKKTKKRK 241
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,710,411
Number of Sequences: 63676
Number of extensions: 750289
Number of successful extensions: 5402
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 5316
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5386
length of query: 980
length of database: 12,639,632
effective HSP length: 106
effective length of query: 874
effective length of database: 5,889,976
effective search space: 5147839024
effective search space used: 5147839024
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC129091.5


