

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.3 + phase: 0
(170 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
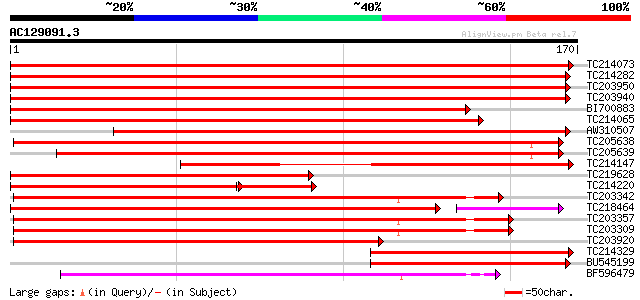
Score E
Sequences producing significant alignments: (bits) Value
TC214073 322 7e-89
TC214282 305 9e-84
TC203950 263 4e-71
TC203940 257 2e-69
BI700883 256 3e-69
TC214065 248 1e-66
AW310507 199 7e-52
TC205638 similar to GB|AAM61587.1|21537246|AY085029 aluminum-ind... 184 2e-47
TC205639 similar to GB|AAM61587.1|21537246|AY085029 aluminum-ind... 172 7e-44
TC214147 166 7e-42
TC219628 154 2e-38
TC214220 120 4e-36
TC203342 similar to UP|Q6UK15 (Q6UK15) Al-induced protein, complete 141 2e-34
TC218464 weakly similar to UP|TSJT_TOBAC (P24805) Stem-specific ... 131 5e-34
TC203357 homologue to UP|O64438 (O64438) ARG10, partial (71%) 139 5e-34
TC203309 homologue to UP|O64438 (O64438) ARG10, complete 139 5e-34
TC203920 similar to UP|Q6UK15 (Q6UK15) Al-induced protein, parti... 127 3e-30
TC214329 similar to UP|Q9LE80 (Q9LE80) Arabidopsis thaliana geno... 126 6e-30
BU545199 96 1e-20
BF596479 similar to GP|2970051|dbj ARG10 {Vigna radiata}, partia... 93 5e-20
>TC214073
Length = 1110
Score = 322 bits (824), Expect = 7e-89
Identities = 151/169 (89%), Positives = 161/169 (94%)
Frame = +3
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G LHNLS LNKQYGLSKG NEAMFIIEAYRTLRDRGPYPADQVLK LEGSFAFVIYD+K
Sbjct: 405 LGGLHNLSMLNKQYGLSKGTNEAMFIIEAYRTLRDRGPYPADQVLKELEGSFAFVIYDNK 584
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
DGTVF ASGS+GHIGLYWGIA DGSV+ISENLEL+KASCAKSFAPFP GC+FHSEHGL+N
Sbjct: 585 DGTVFVASGSNGHIGLYWGIAGDGSVIISENLELIKASCAKSFAPFPAGCMFHSEHGLMN 764
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATWG 169
FEHPT+KMKAMPRIDSEGVMCGANFNVDSQS+ Q+MPRVGSEANWATWG
Sbjct: 765 FEHPTQKMKAMPRIDSEGVMCGANFNVDSQSKIQVMPRVGSEANWATWG 911
>TC214282
Length = 876
Score = 305 bits (780), Expect = 9e-84
Identities = 143/168 (85%), Positives = 155/168 (92%)
Frame = +1
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G L NLS LNKQYGLSKG NEAMFI EAYRTLRDRGPYPADQVLK LEGSF FVIYD+K
Sbjct: 43 LGGLXNLSMLNKQYGLSKGTNEAMFITEAYRTLRDRGPYPADQVLKELEGSFGFVIYDNK 222
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
DGTVF ASGS+G IGLYWG+AADGS+VISENLEL+KASCAKSFAPFP GC+FHSEHGL++
Sbjct: 223 DGTVFVASGSNGQIGLYWGVAADGSIVISENLELIKASCAKSFAPFPNGCMFHSEHGLMS 402
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATW 168
EHPTKKMKAMPRIDSEG MCGANFNVDSQ++ Q+MPRVGSEANWATW
Sbjct: 403 IEHPTKKMKAMPRIDSEGFMCGANFNVDSQTKIQVMPRVGSEANWATW 546
>TC203950
Length = 1217
Score = 263 bits (671), Expect = 4e-71
Identities = 121/168 (72%), Positives = 145/168 (86%)
Frame = +2
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G+L+NLS LNKQYGLSK +EAMF+IEAY+TLRDRGPYPADQV+K L+GSFAFV+YD K
Sbjct: 311 LGSLNNLSLLNKQYGLSKSTDEAMFVIEAYKTLRDRGPYPADQVVKELDGSFAFVVYDSK 490
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
G+VFAA GSDG + LYWGIAADGSVVIS++LE++K CAKSFAPFPTGC+FHSE GL++
Sbjct: 491 VGSVFAALGSDGGVKLYWGIAADGSVVISDDLEVIKEGCAKSFAPFPTGCMFHSEGGLMS 670
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATW 168
FEHP K+KAMPR+DSEG MCGANF VD +R +PRVGS++NW W
Sbjct: 671 FEHPMNKLKAMPRVDSEGAMCGANFKVDKFARVNSIPRVGSQSNWMEW 814
>TC203940
Length = 932
Score = 257 bits (657), Expect = 2e-69
Identities = 119/168 (70%), Positives = 142/168 (83%)
Frame = +3
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G+L+NL LNKQYGL+KG NEAMF+IEAY+TLRDRGPYPADQV+K L+GSF FV+YD K
Sbjct: 321 LGSLNNLCSLNKQYGLAKGTNEAMFVIEAYKTLRDRGPYPADQVVKDLDGSFGFVVYDSK 500
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
G+VFAA GSDG I LYWGIAAD SVVIS++L+++K CAKSFAPFP GC+FHSE GL++
Sbjct: 501 VGSVFAALGSDGGIKLYWGIAADESVVISDDLDVMKEGCAKSFAPFPPGCMFHSEGGLMS 680
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATW 168
FEHP K+KAMPRIDSEG +CGANF VD +R +PRVGS+ANW W
Sbjct: 681 FEHPMNKLKAMPRIDSEGAICGANFKVDKYARVNSIPRVGSQANWMEW 824
>BI700883
Length = 422
Score = 256 bits (655), Expect = 3e-69
Identities = 121/138 (87%), Positives = 129/138 (92%)
Frame = +1
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G LHNLS LNKQYGLSKG NEA FIIEAYRTLRDRGPYP DQVLK LEGSF FVIYD+K
Sbjct: 7 LGGLHNLSMLNKQYGLSKGTNEARFIIEAYRTLRDRGPYPVDQVLKELEGSFGFVIYDNK 186
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
DG+VF ASGS+GHIGLYWGIAADGSV ISENLEL+KASCAKSFAPFPTGC+FHSEHGL+N
Sbjct: 187 DGSVFVASGSNGHIGLYWGIAADGSVTISENLELIKASCAKSFAPFPTGCMFHSEHGLMN 366
Query: 121 FEHPTKKMKAMPRIDSEG 138
FEHPT+KMKAMPRIDSEG
Sbjct: 367 FEHPTQKMKAMPRIDSEG 420
>TC214065
Length = 801
Score = 248 bits (632), Expect = 1e-66
Identities = 116/142 (81%), Positives = 128/142 (89%)
Frame = +3
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G LHNLS LNKQYGLSKG NEAMFI EAYRTLRDRGPYPADQVLK LEGSF FVIYD+K
Sbjct: 366 LGGLHNLSMLNKQYGLSKGTNEAMFITEAYRTLRDRGPYPADQVLKELEGSFGFVIYDNK 545
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
DGTVF ASGS+G IG YWG+AADGS+VISENLEL+KASCAKSFAPFP GC+FHSEHGL++
Sbjct: 546 DGTVFVASGSNGQIGFYWGVAADGSIVISENLELIKASCAKSFAPFPNGCMFHSEHGLMS 725
Query: 121 FEHPTKKMKAMPRIDSEGVMCG 142
EHPTKKMKAMPRI ++G+ G
Sbjct: 726 IEHPTKKMKAMPRIATKGLCVG 791
>AW310507
Length = 412
Score = 199 bits (505), Expect = 7e-52
Identities = 94/137 (68%), Positives = 109/137 (78%)
Frame = -1
Query: 32 TLRDRGPYPADQVLKGLEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISEN 91
+LR GPY A VLK LE F FVIYD DGTVF S+ IGLYW +AA GS+VISE+
Sbjct: 412 SLRHMGPYIAAHVLKELEARFEFVIYDTTDGTVFVPLRSNAQIGLYWRVAAHGSIVISED 233
Query: 92 LELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCGANFNVDSQS 151
L+L+KASCA SFAP P GC+FHSEHGL++ EHPTKKMKAMPRIDS+G MCGA FNV SQ+
Sbjct: 232 LQLIKASCATSFAPSPNGCMFHSEHGLMSIEHPTKKMKAMPRIDSQGFMCGATFNVHSQT 53
Query: 152 RNQMMPRVGSEANWATW 168
+ Q+ PRVGSEA+WATW
Sbjct: 52 KIQVTPRVGSEAHWATW 2
>TC205638 similar to GB|AAM61587.1|21537246|AY085029 aluminum-induced
protein-like {Arabidopsis thaliana;} , partial (98%)
Length = 1107
Score = 184 bits (466), Expect = 2e-47
Identities = 84/167 (50%), Positives = 116/167 (69%), Gaps = 2/167 (1%)
Frame = +3
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G+L N++ L +QYGL+K E + IIEAYRTLRDRGPYPA QV++ +G FAF++YD
Sbjct: 300 GHLENVANLKQQYGLNKTATEVIIIIEAYRTLRDRGPYPAAQVVRDFQGKFAFILYDSGS 479
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNF 121
T F A+ +DG + WG ADG+++ S+ E+V SC KS APFP G F + GL +F
Sbjct: 480 KTAFVAADADGSVPFVWGTDADGNLIFSDETEIVTKSCGKSSAPFPKGFFFSTSGGLSSF 659
Query: 122 EHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQM--MPRVGSEANWA 166
EHP ++K +PR+DS G +CGA F VD++++ + MPRVGS ANW+
Sbjct: 660 EHPLNEVKPVPRVDSSGQVCGATFKVDAEAKKEATGMPRVGSAANWS 800
>TC205639 similar to GB|AAM61587.1|21537246|AY085029 aluminum-induced
protein-like {Arabidopsis thaliana;} , partial (61%)
Length = 730
Score = 172 bits (436), Expect = 7e-44
Identities = 79/154 (51%), Positives = 107/154 (69%), Gaps = 2/154 (1%)
Frame = +3
Query: 15 GLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKDGTVFAASGSDGHI 74
GL+K E + IIEAYRTLRDRGPYPA QV++ +G FAF++YD T F A+ +DG +
Sbjct: 3 GLNKTATEVIIIIEAYRTLRDRGPYPAAQVVRDFQGKFAFILYDSGSKTAFVAADADGSV 182
Query: 75 GLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKKMKAMPRI 134
WG ADG++V S+ E+V SC KS+APFP G F + GL +FEHP ++K +PR+
Sbjct: 183 PFVWGTDADGNLVFSDETEIVTKSCGKSYAPFPKGFFFSTSGGLSSFEHPLNEVKPVPRV 362
Query: 135 DSEGVMCGANFNVDSQSRNQM--MPRVGSEANWA 166
DS G +CGA F VD++++ + MPRVGS ANW+
Sbjct: 363 DSSGQVCGATFKVDAEAKKEATGMPRVGSAANWS 464
>TC214147
Length = 487
Score = 166 bits (419), Expect = 7e-42
Identities = 81/118 (68%), Positives = 88/118 (73%)
Frame = +2
Query: 52 FAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCL 111
FAFVIYD+KDGTVF ASGS+GHI LYWGIA GC+
Sbjct: 2 FAFVIYDNKDGTVFVASGSNGHIELYWGIA---------------------------GCM 100
Query: 112 FHSEHGLLNFEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATWG 169
FHSEHGL+NFEHPT+KMKAMPRIDSEGVMCGANFNVDSQS+ Q+MPRVGSEANWATWG
Sbjct: 101 FHSEHGLMNFEHPTQKMKAMPRIDSEGVMCGANFNVDSQSKIQVMPRVGSEANWATWG 274
>TC219628
Length = 670
Score = 154 bits (390), Expect = 2e-38
Identities = 75/91 (82%), Positives = 84/91 (91%)
Frame = +1
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G+L+NLS+L KQYGLSKG N+AMFIIEAYRTLRDRGPYPADQVLK LEGSF FVIYD+K
Sbjct: 397 LGSLNNLSKLIKQYGLSKGTNDAMFIIEAYRTLRDRGPYPADQVLKELEGSFGFVIYDNK 576
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISEN 91
DGTVF ASGS+G I L+WG+AADGSVVISEN
Sbjct: 577 DGTVFTASGSNGQIELFWGVAADGSVVISEN 669
>TC214220
Length = 714
Score = 120 bits (300), Expect(2) = 4e-36
Identities = 59/70 (84%), Positives = 61/70 (86%)
Frame = +2
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G LHNLS LNKQYGLSKG NEA FI EAYRTLRDRGPYPADQVLK LEGSF FVIYD+K
Sbjct: 332 LGGLHNLSMLNKQYGLSKGTNEARFITEAYRTLRDRGPYPADQVLKELEGSFGFVIYDNK 511
Query: 61 DGTVFAASGS 70
DGTVF AS S
Sbjct: 512 DGTVFVASVS 541
Score = 47.8 bits (112), Expect(2) = 4e-36
Identities = 20/24 (83%), Positives = 23/24 (95%)
Frame = +3
Query: 69 GSDGHIGLYWGIAADGSVVISENL 92
GS+G IGLYWG+AADGS+VISENL
Sbjct: 642 GSNGQIGLYWGVAADGSIVISENL 713
>TC203342 similar to UP|Q6UK15 (Q6UK15) Al-induced protein, complete
Length = 965
Score = 141 bits (355), Expect = 2e-34
Identities = 69/148 (46%), Positives = 96/148 (64%), Gaps = 1/148 (0%)
Frame = +3
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G L NL L +QYGL+K NE + +IEAY+ LRDR PYPA+ V+ L GSFAF+++D
Sbjct: 282 GALDNLGNLRQQYGLAKSTNEVLLVIEAYKALRDRAPYPANHVVGHLSGSFAFIVFDKST 461
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSE-HGLLN 120
T+F AS G + LYWGI ADG V +++ EL+K +C KS A FP GC + + GL+
Sbjct: 462 STLFVASDQYGKVPLYWGITADGYVAFADDAELLKGACGKSLASFPQGCFYSTAVGGLMC 641
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVD 148
+E+P K+ A+P + E + GA F V+
Sbjct: 642 YENPKNKITAVPANEEE--IWGATFKVE 719
>TC218464 weakly similar to UP|TSJT_TOBAC (P24805) Stem-specific protein
TSJT1, partial (71%)
Length = 1012
Score = 131 bits (329), Expect(2) = 5e-34
Identities = 59/129 (45%), Positives = 86/129 (65%)
Frame = +2
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
MG L N+++L YGL + EAM +IEAY+ LRDR PYP DQV K L+G FAF+I+D K
Sbjct: 332 MGALANIAELRHHYGLPRQATEAMIVIEAYKALRDRAPYPPDQVAKHLDGKFAFIIFDAK 511
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
T+F A +G + WG+A DGS+V S++ +++ C ++ A FP GC+F + GL +
Sbjct: 512 TYTLFIARDREGSVKFQWGMARDGSLVCSDDPTIIREGCGQACAAFPPGCIFINGSGLTS 691
Query: 121 FEHPTKKMK 129
F+HP K++
Sbjct: 692 FDHPLHKVQ 718
Score = 29.6 bits (65), Expect(2) = 5e-34
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +3
Query: 135 DSEGVMCGANFNVDSQSRNQMMPRVGSEANWA 166
D G + F VD ++ +PR GS ANWA
Sbjct: 735 DDSGNILSVYFQVDLYTKIPSIPRTGSAANWA 830
>TC203357 homologue to UP|O64438 (O64438) ARG10, partial (71%)
Length = 1203
Score = 139 bits (351), Expect = 5e-34
Identities = 69/151 (45%), Positives = 97/151 (63%), Gaps = 1/151 (0%)
Frame = +2
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G L NL L +QYGL+K NE + +IEAY+ LRDR PYPA++V+ L GSFAF+++D
Sbjct: 500 GALDNLGSLRQQYGLAKSANEVILVIEAYKALRDRAPYPANRVVCHLSGSFAFIVFDKST 679
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSE-HGLLN 120
T+F AS G + LYWGI ADG V +++ +L+K SC KS A FP GC + + GL
Sbjct: 680 STLFVASDQAGKVPLYWGITADGYVAFADDADLLKGSCGKSLASFPQGCFYSTAVGGLRC 859
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQS 151
+E+P K+ A+P + E + GA F V+ +
Sbjct: 860 YENPKNKITAVPAEEEE--IWGAFFKVEGST 946
>TC203309 homologue to UP|O64438 (O64438) ARG10, complete
Length = 1052
Score = 139 bits (351), Expect = 5e-34
Identities = 69/151 (45%), Positives = 97/151 (63%), Gaps = 1/151 (0%)
Frame = +2
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G L NL L +QYGL+K NE + +IEAY+ LRDR PYPA++V+ L GSFAF+++D
Sbjct: 371 GALDNLGSLRQQYGLAKSANEVVLVIEAYKALRDRAPYPANRVVCHLSGSFAFIVFDKST 550
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSE-HGLLN 120
T+F AS G + LYWGI ADG V +++ +L+K SC KS A FP GC + + GL
Sbjct: 551 STLFVASDQAGKVPLYWGITADGYVAFADDADLLKGSCGKSLASFPQGCFYSTAVGGLRC 730
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQS 151
+E+P K+ A+P + E + GA F V+ +
Sbjct: 731 YENPKNKITAIPAEEEE--IWGAFFKVEGSA 817
>TC203920 similar to UP|Q6UK15 (Q6UK15) Al-induced protein, partial (81%)
Length = 686
Score = 127 bits (318), Expect = 3e-30
Identities = 57/111 (51%), Positives = 75/111 (67%)
Frame = +1
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G L NL L +QYGL+K NE + +IEAY+ LRDR PYPA+ V+ L GSFAF+++D
Sbjct: 235 GALDNLGNLRQQYGLAKSTNEVLLVIEAYKALRDRAPYPANHVVGHLSGSFAFIVFDKST 414
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLF 112
T+F AS G + LYWGI ADG V +++ EL+K +C KS A FP GC +
Sbjct: 415 STLFVASDQYGKVPLYWGITADGYVAFADDAELLKGACGKSLASFPQGCFY 567
>TC214329 similar to UP|Q9LE80 (Q9LE80) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MJK13 (AT3g15450/MJK13_11)
(MJK13.11 protein), partial (23%)
Length = 846
Score = 126 bits (316), Expect = 6e-30
Identities = 55/61 (90%), Positives = 60/61 (98%)
Frame = +2
Query: 109 GCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATW 168
GC+FHSEHGL+NFEHPT+KMKAMPRIDSEGVMCGANFNVDSQS+ Q+MPRVGSEANWATW
Sbjct: 245 GCMFHSEHGLMNFEHPTQKMKAMPRIDSEGVMCGANFNVDSQSKIQVMPRVGSEANWATW 424
Query: 169 G 169
G
Sbjct: 425 G 427
>BU545199
Length = 409
Score = 95.5 bits (236), Expect = 1e-20
Identities = 40/60 (66%), Positives = 48/60 (79%)
Frame = -2
Query: 109 GCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATW 168
GC+FHSE GL++FEHP K+KAMPRIDSEG +CGANF VD +R +PRVGS+ANW W
Sbjct: 204 GCMFHSEGGLMSFEHPMNKLKAMPRIDSEGAICGANFKVDKYARVNSIPRVGSQANWMEW 25
>BF596479 similar to GP|2970051|dbj ARG10 {Vigna radiata}, partial (58%)
Length = 415
Score = 93.2 bits (230), Expect = 5e-20
Identities = 53/133 (39%), Positives = 77/133 (57%), Gaps = 1/133 (0%)
Frame = +2
Query: 16 LSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKDGTVFAASGSDGHIG 75
L+K NE + +I+AY +R R PYPA+ V L G+ AF+I D T+ AS G +
Sbjct: 2 LAKSPNEVVLVIDAY*AIRHRSPYPANLVAFHLTGTSAFIIIDKFTSTLIVASDHSGKVP 181
Query: 76 LYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEH-GLLNFEHPTKKMKAMPRI 134
LY GI ADG V +++ +L+K SC S A F GC + + GL +E+P + A+P +
Sbjct: 182 LYCGITADGYVSCADDADLLKGSCGTSLAYFSQGCFYSTAV*GLRCYENPNNTITAIPAV 361
Query: 135 DSEGVMCGANFNV 147
+ E + C A FNV
Sbjct: 362 EDE-IWC-AYFNV 394
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,285,103
Number of Sequences: 63676
Number of extensions: 85895
Number of successful extensions: 407
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 402
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 403
length of query: 170
length of database: 12,639,632
effective HSP length: 90
effective length of query: 80
effective length of database: 6,908,792
effective search space: 552703360
effective search space used: 552703360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC129091.3


