

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
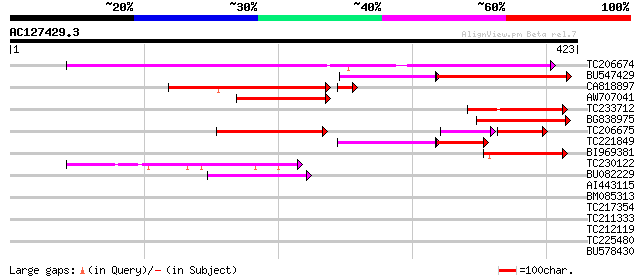
Query= AC127429.3 - phase: 0 /pseudo
(423 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC206674 similar to UP|FMO3_HUMAN (P31513) Dimethylaniline monoo... 271 5e-73
BU547429 172 9e-49
CA818897 similar to GP|16555354|gb flavin-containing monooxygena... 159 5e-41
AW707041 108 5e-24
TC233712 similar to UP|O23024 (O23024) T1G11.14 protein (Flavin-... 105 5e-23
BG838975 similar to GP|16555354|gb| flavin-containing monooxygen... 99 5e-21
TC206675 84 9e-17
TC221849 64 1e-16
BI969381 84 2e-16
TC230122 51 9e-07
BU082229 similar to GP|9757850|dbj dimethylaniline monooxygenase... 47 2e-05
AI443115 similar to GP|10177621|db phytoene dehydrogenase-like {... 32 0.41
BM085313 similar to GP|10177621|db phytoene dehydrogenase-like {... 32 0.41
TC217354 similar to UP|Q9SBB5 (Q9SBB5) CTF2A, partial (50%) 32 0.70
TC211333 similar to UP|Q9SXE0 (Q9SXE0) T3P18.13, partial (18%) 30 2.7
TC212119 weakly similar to UP|Q9LDR3 (Q9LDR3) Arabidopsis thalia... 29 3.5
TC225480 UP|Q9XE94 (Q9XE94) Geranylgeranyl hydrogenase, complete 28 6.0
BU578430 28 7.8
>TC206674 similar to UP|FMO3_HUMAN (P31513) Dimethylaniline monooxygenase
[N-oxide-forming] 3 (Hepatic flavin-containing
monooxygenase 3) (FMO 3) (Dimethylaniline oxidase 3)
(FMO form 2) (FMO II) , partial (4%)
Length = 1470
Score = 271 bits (692), Expect = 5e-73
Identities = 146/377 (38%), Positives = 219/377 (57%), Gaps = 12/377 (3%)
Frame = +3
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEF 102
+I+GAG SG+A A L ++ +P ++LER +C ASLW+ TYDRL LHL KQVCELP + F
Sbjct: 57 IIIGAGTSGIATAGCLTKQSIPYIMLEREDCFASLWQKYTYDRLHLHLRKQVCELPHLPF 236
Query: 103 PSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSEG-KAGMVTE 161
P +P Y ++QFI+YL +Y +F+I+P + V E+D G WRV+++ ++G + E
Sbjct: 237 PKSYPHYVPRKQFIDYLGNYVNHFEIKPLYQRAVELVEYDGWKGIWRVKAQNRRSGELEE 416
Query: 162 FVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEV 221
+ ++L+VA+GE AE +P+I+G++ F G + H++ YK+G EF+ K VLVVG GNSGME+
Sbjct: 417 YAGKYLVVASGETAEPRLPQIQGLESFNGKVIHSTAYKNGNEFKNKHVLVVGSGNSGMEI 596
Query: 222 CLDLCNHDAAPSIVVRDSPTKRYAGKINFW-----------VVHVVAQVATRATGGSHPS 270
LDL N A PSI+VR SP + + ++ V V + ++ G
Sbjct: 597 ALDLSNFGAKPSIIVR-SPVHFLSRDMMYYASLMLNYLSLSTVEKVLVMVSKVVYGDLSE 773
Query: 271 HCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPRCGCPCQD*KRRHQGIKRLKRYTVE 330
+ + + T++ A K D G ++ + IK ++ V
Sbjct: 774 YGIPYPSEGPFTMKMKYA--------KFPIIDVGTVKKIKSREIQVLPAEIKSIRGNEVL 929
Query: 331 FADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTK 390
F DG + FD+I+ TG+K + WLK EDG+P+ FPN WKG NGLY VG ++
Sbjct: 930 FQDGKSYTFDSIVFCTGFKRSTQKWLKGGDDLLNEDGFPKNSFPNHWKGRNGLYCVGLSR 1109
Query: 391 RGLLGASMDAKNIAEDI 407
RG GA+MDA+ +A DI
Sbjct: 1110RGFFGANMDAQLVANDI 1160
>BU547429
Length = 665
Score = 172 bits (436), Expect(2) = 9e-49
Identities = 79/101 (78%), Positives = 88/101 (86%)
Frame = -2
Query: 319 QGIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWK 378
+GIKRL R VEF DG ENFDA++LATGYKSNVP WLK MFS++DG+PRKPFPNGWK
Sbjct: 325 RGIKRLARNAVEFVDGKVENFDAMVLATGYKSNVPSWLKGSDMFSEKDGFPRKPFPNGWK 146
Query: 379 GENGLYAVGFTKRGLLGASMDAKNIAEDIERCWKAEAKHIF 419
GENGLYAVGFTKRGLLGAS+DAK IAEDIE WKAEAKH++
Sbjct: 145 GENGLYAVGFTKRGLLGASIDAKRIAEDIEHSWKAEAKHVW 23
Score = 39.7 bits (91), Expect(2) = 9e-49
Identities = 25/75 (33%), Positives = 38/75 (50%)
Frame = -1
Query: 247 KINFWVVHVVAQVATRATGGSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTGPR 306
+INFW +HV+AQV GS + V +A +H ++ S+ S Q+ V KDT
Sbjct: 551 EINFWFIHVLAQVVPHTFCGSIFASNVTSHAXRHSSIWTSSSKTWSSRAQELVRKDTSFG 372
Query: 307 CGCPCQD*KRRHQGI 321
C D K ++ G+
Sbjct: 371 CWDTHSDQKWKN*GL 327
>CA818897 similar to GP|16555354|gb flavin-containing monooxygenase YUCCA2
{Arabidopsis thaliana}, partial (26%)
Length = 445
Score = 159 bits (402), Expect(2) = 5e-41
Identities = 78/123 (63%), Positives = 95/123 (76%), Gaps = 2/123 (1%)
Frame = +1
Query: 119 LESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSEG--KAGMVTEFVCRWLIVATGENAE 176
L++Y+ +FDI+P F++TV+ AEFD WRV++ G K E+VC+WLIVATGE AE
Sbjct: 1 LKAYADHFDIKPVFSQTVVSAEFDHVCQLWRVKTRGVIKKEDTAEYVCQWLIVATGECAE 180
Query: 177 AVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDLCNHDAAPSIVV 236
VVP+IEG+ EF G I HTS YKSG F GK VLVVGCGNSGMEVCLDLCNH+A PS+VV
Sbjct: 181 EVVPQIEGMGEFEGQIVHTSKYKSGSMFCGKNVLVVGCGNSGMEVCLDLCNHNARPSLVV 360
Query: 237 RDS 239
RD+
Sbjct: 361 RDT 369
Score = 26.9 bits (58), Expect(2) = 5e-41
Identities = 10/15 (66%), Positives = 13/15 (86%)
Frame = +3
Query: 245 AGKINFWVVHVVAQV 259
A +INFW +HV+AQV
Sbjct: 393 AREINFWFIHVLAQV 437
>AW707041
Length = 365
Score = 108 bits (270), Expect = 5e-24
Identities = 51/70 (72%), Positives = 58/70 (82%)
Frame = +1
Query: 170 ATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDLCNHD 229
ATGENAE V+PEIEG+ EF G + H YKSGE FRGKKVLVVGCGNSGME+ LDLCNH
Sbjct: 7 ATGENAECVMPEIEGLSEFKGDVIHACDYKSGERFRGKKVLVVGCGNSGMELSLDLCNHH 186
Query: 230 AAPSIVVRDS 239
++PS+VVR S
Sbjct: 187 SSPSMVVRSS 216
>TC233712 similar to UP|O23024 (O23024) T1G11.14 protein (Flavin-containing
monooxygenase YUCCA3) (At1g04610), partial (21%)
Length = 741
Score = 105 bits (261), Expect = 5e-23
Identities = 47/75 (62%), Positives = 58/75 (76%)
Frame = +3
Query: 342 IILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAK 401
++LATGY SNVP WLK+ F+ DG PR PFPNGW+G+ GLYAVGFT++GL GAS+DA
Sbjct: 18 VVLATGYHSNVPSWLKENDFFTS-DGTPRNPFPNGWRGKGGLYAVGFTRKGLSGASLDAI 194
Query: 402 NIAEDIERCWKAEAK 416
N+A DI + WK E K
Sbjct: 195 NVAHDIAKNWKEENK 239
>BG838975 similar to GP|16555354|gb| flavin-containing monooxygenase YUCCA2
{Arabidopsis thaliana}, partial (13%)
Length = 428
Score = 98.6 bits (244), Expect = 5e-21
Identities = 46/70 (65%), Positives = 52/70 (73%)
Frame = -2
Query: 349 KSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAEDIE 408
KSN P+ +K + E +PRKPFPNGWKGENG Y VGFTK GLLGAS+DAK IAEDIE
Sbjct: 376 KSNXPFLVKGQXHV**ERWFPRKPFPNGWKGENGXYXVGFTKXGLLGASIDAKRIAEDIE 197
Query: 409 RCWKAEAKHI 418
WK EA H+
Sbjct: 196 HXWKXEATHV 167
Score = 32.3 bits (72), Expect = 0.41
Identities = 15/38 (39%), Positives = 21/38 (54%), Gaps = 1/38 (2%)
Frame = -3
Query: 333 DGSTENFDAIILATG-YKSNVPYWLKDKGMFSKEDGYP 369
DG E D + K+ P+WLK MFS++DG+P
Sbjct: 426 DGKVEXLDXXHXSNW*TKAMXPFWLKGSXMFSEKDGFP 313
>TC206675
Length = 957
Score = 84.3 bits (207), Expect = 9e-17
Identities = 41/83 (49%), Positives = 60/83 (71%)
Frame = +2
Query: 155 KAGMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGC 214
++G V E+ +L++A+GE AE VP+IEG++ F G + H++ Y +G+EF+ K VLVV
Sbjct: 152 RSGEVEEYAGWYLVLASGETAEPRVPQIEGLESFNGKVIHSTGYNNGKEFKDKLVLVV*S 331
Query: 215 GNSGMEVCLDLCNHDAAPSIVVR 237
GNSGME+ LDL + A PSI+VR
Sbjct: 332 GNSGMEIALDLSDFGAKPSIIVR 400
Score = 57.0 bits (136), Expect(2) = 2e-11
Identities = 22/37 (59%), Positives = 29/37 (77%)
Frame = +2
Query: 365 EDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAK 401
E+G+P+ FPN WKG NGLY VG ++RG GA+MDA+
Sbjct: 785 ENGFPKSSFPNHWKGRNGLYCVGLSRRGFFGANMDAQ 895
Score = 29.3 bits (64), Expect(2) = 2e-11
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +1
Query: 322 KRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMF 362
K ++ V F DG + FD+I+ T +K + WL+ +F
Sbjct: 652 KSIRGDEVLFRDGKSRPFDSIVFCTRFKRSTQKWLQGVMIF 774
>TC221849
Length = 452
Score = 64.3 bits (155), Expect(2) = 1e-16
Identities = 29/39 (74%), Positives = 32/39 (81%)
Frame = +1
Query: 319 QGIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLK 357
+GIKRL R VEF DG ENFDA++LATGYKSNVP WLK
Sbjct: 328 RGIKRLARNAVEFVDGKVENFDAMVLATGYKSNVPSWLK 444
Score = 39.7 bits (91), Expect(2) = 1e-16
Identities = 26/77 (33%), Positives = 39/77 (49%)
Frame = +3
Query: 245 AGKINFWVVHVVAQVATRATGGSHPSHCVMVNARQHRTVRFGSASVGSP*TQKTVWKDTG 304
A +INFW +HV+AQV GS + V +A +H ++ S+ S Q+ V KDT
Sbjct: 96 AREINFWFIHVLAQVVPHTFCGSIFASNVTSHAWRHSSIWTSSSKTWSSRAQELVRKDTS 275
Query: 305 PRCGCPCQD*KRRHQGI 321
C D K ++ G+
Sbjct: 276 FGCWDTHSDQKWKN*GL 326
>BI969381
Length = 602
Score = 83.6 bits (205), Expect = 2e-16
Identities = 39/65 (60%), Positives = 47/65 (72%), Gaps = 2/65 (3%)
Frame = -2
Query: 354 YWL--KDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCW 411
+WL +G F ++GYP+ PFP+GWKG GLYAVGFTKRGL GAS DA IA+DI + W
Sbjct: 556 FWLCYVQEGEFFSKNGYPKMPFPHGWKGNAGLYAVGFTKRGLSGASSDAVKIAQDIGQVW 377
Query: 412 KAEAK 416
K E K
Sbjct: 376 KNETK 362
>TC230122
Length = 779
Score = 51.2 bits (121), Expect = 9e-07
Identities = 48/213 (22%), Positives = 95/213 (44%), Gaps = 37/213 (17%)
Frame = +3
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEF 102
+I+GAG SGL Y+ + G ++ E + + LW+ T + +L KQ+ + M+F
Sbjct: 150 VIIGAGISGLVACKYVIEFGFNPIVFEVDDGVGGLWR-HTMESTKLQNNKQMFQ--FMDF 320
Query: 103 ---PSGFPTYPTKQQFIEYLESYSKNFDIRPW--FNETVMHAEF---------------- 141
PS P+ +Q ++Y+ SY+++F + P+ FN V+ ++
Sbjct: 321 PWPPSVKEDNPSHKQVLDYVNSYAEHFSLIPYIRFNSKVIDIDYVGGESSEEMKSWELWG 500
Query: 142 -----DATLGFWRVRSEGKAGMVTE-FVCRWLIVATGE-NAEAVVPEI---EGVDEFVGS 191
+ G W + + + E ++++ G+ + +PE +G + F G
Sbjct: 501 GNGRPFCSKGTWHIAVQHTKNLSIEVHEAEFVVLCIGKYSGFPNIPEFPPGKGPEVFNGK 680
Query: 192 IRHTSLYK------SGEEFRGKKVLVVGCGNSG 218
+ H+ Y + E +GK+V ++G SG
Sbjct: 681 VMHSMDYSNLDNDTAAELVKGKRVTIIGSLKSG 779
>BU082229 similar to GP|9757850|dbj dimethylaniline monooxygenase
(N-oxide-forming)-like protein {Arabidopsis thaliana},
partial (22%)
Length = 423
Score = 46.6 bits (109), Expect = 2e-05
Identities = 29/79 (36%), Positives = 46/79 (57%), Gaps = 1/79 (1%)
Frame = +2
Query: 148 WRVRSEGK-AGMVTEFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRG 206
W VRS+ K + E V ++VATG + +P I+G+ + H+ +Y+S E FRG
Sbjct: 83 WVVRSKDKNSEEEVEQVFDAVVVATGHYSNPRLPCIQGMAIWKRKQMHSHIYRSPEPFRG 262
Query: 207 KKVLVVGCGNSGMEVCLDL 225
+ V+VVG SG E+ ++L
Sbjct: 263 EIVVVVGNSFSGQEISMEL 319
>AI443115 similar to GP|10177621|db phytoene dehydrogenase-like
{Arabidopsis thaliana}, partial (20%)
Length = 421
Score = 32.3 bits (72), Expect = 0.41
Identities = 15/32 (46%), Positives = 21/32 (64%)
Frame = +2
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCI 74
L++G G +GL AAYL + G+ ILER + I
Sbjct: 29 LVIGGGHNGLTAAAYLARGGLSVAILERRHVI 124
>BM085313 similar to GP|10177621|db phytoene dehydrogenase-like {Arabidopsis
thaliana}, partial (19%)
Length = 426
Score = 32.3 bits (72), Expect = 0.41
Identities = 15/32 (46%), Positives = 21/32 (64%)
Frame = +1
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCI 74
LI+G G +GL AAYL + G+ +LER + I
Sbjct: 70 LIIGGGHNGLTAAAYLARGGLSVAVLERRHVI 165
>TC217354 similar to UP|Q9SBB5 (Q9SBB5) CTF2A, partial (50%)
Length = 859
Score = 31.6 bits (70), Expect = 0.70
Identities = 15/32 (46%), Positives = 23/32 (71%)
Frame = +3
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCI 74
++VGAG +GLA A L + GV SL+LE++ +
Sbjct: 120 VVVGAGIAGLATALSLHRLGVRSLVLEQAQSL 215
>TC211333 similar to UP|Q9SXE0 (Q9SXE0) T3P18.13, partial (18%)
Length = 828
Score = 29.6 bits (65), Expect = 2.7
Identities = 16/38 (42%), Positives = 23/38 (60%)
Frame = +3
Query: 329 VEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKED 366
V F DG + DAII TGYK + P+ L+ G+ + +D
Sbjct: 135 VAFEDGFSVYADAIIHCTGYKYHFPF-LETNGLVTVDD 245
>TC212119 weakly similar to UP|Q9LDR3 (Q9LDR3) Arabidopsis thaliana genomic
DNA, chromosome 3, P1 clone: MJK13 (MJK13.8 protein)
(At3g15420), partial (25%)
Length = 493
Score = 29.3 bits (64), Expect = 3.5
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = -2
Query: 80 LKTYDRLRLHLPKQVCELPLMEFPSGF 106
L +DR++LHLP C +P + P+ F
Sbjct: 381 LDPHDRVKLHLPPYSC*MPKSQLPNSF 301
>TC225480 UP|Q9XE94 (Q9XE94) Geranylgeranyl hydrogenase, complete
Length = 1746
Score = 28.5 bits (62), Expect = 6.0
Identities = 14/32 (43%), Positives = 20/32 (61%), Gaps = 2/32 (6%)
Frame = +1
Query: 44 IVGAGPSGLAVAAYLKQKGVPSLILER--SNC 73
+VG GP+G A A L + GV + ++ER NC
Sbjct: 166 VVGGGPAGGAAAETLAKGGVETFLIERKMDNC 261
>BU578430
Length = 402
Score = 28.1 bits (61), Expect = 7.8
Identities = 10/26 (38%), Positives = 18/26 (68%)
Frame = +3
Query: 117 EYLESYSKNFDIRPWFNETVMHAEFD 142
E LESY + + +RP+ N+ +MH ++
Sbjct: 231 EELESYCQMWSLRPFINDEIMHQAWE 308
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,473,236
Number of Sequences: 63676
Number of extensions: 312911
Number of successful extensions: 1387
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1374
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1383
length of query: 423
length of database: 12,639,632
effective HSP length: 100
effective length of query: 323
effective length of database: 6,272,032
effective search space: 2025866336
effective search space used: 2025866336
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC127429.3


