

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
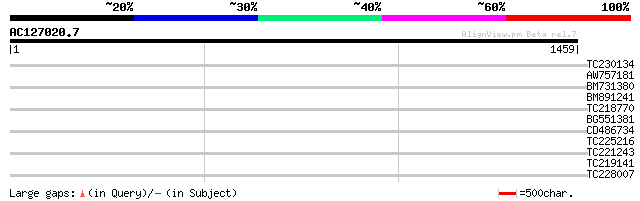
Query= AC127020.7 + phase: 0 /pseudo
(1459 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Pro... 35 0.24
AW757181 34 0.41
BM731380 32 1.6
BM891241 32 2.7
TC218770 UP|Q9XIU5 (Q9XIU5) GmMYB29B2 protein, complete 32 2.7
BG551381 31 3.5
CD486734 homologue to GP|16419191|gb| glucosamine-6-phosphate de... 31 4.6
TC225216 homologue to UP|Q6PW04 (Q6PW04) Membrane-anchored endo-... 30 6.0
TC221243 similar to UP|DNLI_ARATH (Q42572) DNA ligase (Polydeox... 30 6.0
TC219141 30 7.8
TC228007 similar to UP|Q9LT13 (Q9LT13) Gb|AAF01580.1, partial (51%) 30 7.8
>TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23),
partial (8%)
Length = 581
Score = 35.0 bits (79), Expect = 0.24
Identities = 20/73 (27%), Positives = 41/73 (55%), Gaps = 7/73 (9%)
Frame = -2
Query: 484 FVFCLFMIVIFL*TAPGEAGVLLYFGVIL*IVILLIFQEITLLLGLLIVLR-------VL 536
F+F LF+I+IF+ ++ +++ I+IL++ T +GL I++R +
Sbjct: 517 FIFILFVIIIFI-------SIIFIVIIVVVIIILIVVIAFTSTIGLGIIIRISFFPAIIT 359
Query: 537 *DLLVIMIILIAV 549
++L+I II++ V
Sbjct: 358 REILIIFIIIVTV 320
>AW757181
Length = 422
Score = 34.3 bits (77), Expect = 0.41
Identities = 32/136 (23%), Positives = 51/136 (36%)
Frame = +3
Query: 79 YLFQFHHKVDDARVLEEGPWLYDNFHIVLDRIAPSAVPNLVPLNHIEFWVQVHGLPFGFV 138
YL QF + + G + + +I++ R P+ + N + + WVQ LP
Sbjct: 15 YLVQFTSEEGYKNAMFNGS*MVADHYIIV*RWKPTFIQNAKKM*KVAEWVQFLLLPMKLY 194
Query: 139 QPKVGQGIGSFLGTLKAYDVRNTIHSSYMRIKVAIDVTVPLKKEWRVRASNGNFVTVNFK 198
S LGT+ D +IHS ++V L K V + +
Sbjct: 195 HDVFL*TTWSLLGTILNID*CTSIHSKARFSPTCVEVV--LNKTKLVSKFEARWNVYFVE 368
Query: 199 YEKLGVFCHRCGVLGH 214
L + C +CG GH
Sbjct: 369 *NGLHLICFQCGTYGH 416
>BM731380
Length = 433
Score = 32.3 bits (72), Expect = 1.6
Identities = 18/41 (43%), Positives = 30/41 (72%)
Frame = +2
Query: 504 VLLYFGVIL*IVILLIFQEITLLLGLLIVLRVL*DLLVIMI 544
VLL ++L +V+LL+ Q + LLL L++L+VL LLV+++
Sbjct: 293 VLLLLVLLLLLVLLLLLQVLLLLLVFLLLLQVLLLLLVLLL 415
>BM891241
Length = 407
Score = 31.6 bits (70), Expect = 2.7
Identities = 19/37 (51%), Positives = 21/37 (56%)
Frame = -3
Query: 1277 G*TGKSYPHQRYMEVWVSKISPLLI*LC*VNRVGS*L 1313
G* G+ + R VW SKISP I*LC N G *L
Sbjct: 375 G*NGRLFACPRLRGVWGSKISPNSI*LCWANGYGP*L 265
>TC218770 UP|Q9XIU5 (Q9XIU5) GmMYB29B2 protein, complete
Length = 1316
Score = 31.6 bits (70), Expect = 2.7
Identities = 28/98 (28%), Positives = 43/98 (43%), Gaps = 16/98 (16%)
Frame = -1
Query: 643 FRIFPLLMLKLWW-------LRLLITTQFTSILLL---------HLGPMLLSVIFAMKMR 686
F + LL L L W +L+ T F S+ +L HL P + + +
Sbjct: 509 FGLVALLELGLDWSDFSSLFFKLVCQTFFISLSVLPGNLAAIADHLFPNISCSLMIVSSS 330
Query: 687 GILNRASRISLLILGRNILLIRSFRSFPHVRKTCLYGR 724
I+ + LLI G L+ RS + FPH+R+ +GR
Sbjct: 329 SIV----KFPLLISGLK*LIQRSLQLFPHLRRPACFGR 228
>BG551381
Length = 230
Score = 31.2 bits (69), Expect = 3.5
Identities = 16/65 (24%), Positives = 27/65 (40%)
Frame = -3
Query: 223 FELDSDDGVRNWGPYLKPVTQRVGTAATNRWLQDPIPAVTPSHNSDVAAASVGRKTQTAG 282
F SD+ NW P K ++ ++ WL+ +P + SH S + + + G
Sbjct: 222 FRTFSDNPSTNWTPLFKSTMSKLQPLSSGCWLRSSLPPFSVSHISSPISLLLRAQLHCHG 43
Query: 283 TGVVT 287
G T
Sbjct: 42 GGTAT 28
>CD486734 homologue to GP|16419191|gb| glucosamine-6-phosphate deaminase
{Salmonella typhimurium LT2}, partial (20%)
Length = 366
Score = 30.8 bits (68), Expect = 4.6
Identities = 15/25 (60%), Positives = 17/25 (68%)
Frame = +3
Query: 330 LLPSSSTGTSASNLLPDRPIVLGLP 354
LL S T ++ SN PDRP VLGLP
Sbjct: 234 LLVISLTASTRSNQQPDRPFVLGLP 308
>TC225216 homologue to UP|Q6PW04 (Q6PW04) Membrane-anchored
endo-1,4-beta-glucanase , complete
Length = 2204
Score = 30.4 bits (67), Expect = 6.0
Identities = 19/68 (27%), Positives = 36/68 (52%), Gaps = 8/68 (11%)
Frame = +3
Query: 648 LLMLKLWWLRLLITTQFTSILLLHLGPML------LSVIFAMKMRGILN--RASRISLLI 699
LL L++WW+ ++ +S+ L P+L LS++ M++ G LN R S +LI
Sbjct: 444 LLRLRIWWVGTMMPGMLSSLTFLLRFPLLC*VGV*LSIVQNMRLLGSLNMLRRSLSGVLI 623
Query: 700 LGRNILLI 707
+ + ++
Sbjct: 624 IFSRVSIV 647
>TC221243 similar to UP|DNLI_ARATH (Q42572) DNA ligase
(Polydeoxyribonucleotide synthase [ATP]) , partial (22%)
Length = 847
Score = 30.4 bits (67), Expect = 6.0
Identities = 19/65 (29%), Positives = 33/65 (50%)
Frame = +3
Query: 655 WLRLLITTQFTSILLLHLGPMLLSVIFAMKMRGILNRASRISLLILGRNILLIRSFRSFP 714
W LL++T ++ + LL++ MK + R R SL + +N+LL+ + FP
Sbjct: 99 WYLLLLSTVVGNVQEFMVPSSLLAMTMIMKNFKVFAR*ERDSLKLCLKNVLLVFVLK*FP 278
Query: 715 HVRKT 719
+ R T
Sbjct: 279 NQRHT 293
>TC219141
Length = 703
Score = 30.0 bits (66), Expect = 7.8
Identities = 26/95 (27%), Positives = 43/95 (44%), Gaps = 11/95 (11%)
Frame = +3
Query: 525 LLLGLLIVLRVL*D--LLVIMIILIAVVKMPRGIFYDNFHLNLQGLGA---------FLA 573
L+LG+L VL + L + I LIA + N +L G+GA FL
Sbjct: 186 LVLGVLTVLFPTEEGPTLQVAISLIATIYFIHERLKSNIRASLYGVGAFGISWLLGTFLM 365
Query: 574 ILMTSSIQVRKGGTPLDLLGLLTAFVKLFLTLVYL 608
+ + I + KG +++ L +V L+++ YL
Sbjct: 366 VSVIPPITILKGPRAFEVISSLITYVLLWVSSTYL 470
>TC228007 similar to UP|Q9LT13 (Q9LT13) Gb|AAF01580.1, partial (51%)
Length = 1707
Score = 30.0 bits (66), Expect = 7.8
Identities = 17/58 (29%), Positives = 28/58 (47%)
Frame = +2
Query: 228 DDGVRNWGPYLKPVTQRVGTAATNRWLQDPIPAVTPSHNSDVAAASVGRKTQTAGTGV 285
DD ++ PYL PV + ++ + +DP+PA+ S A GR+ Q G +
Sbjct: 674 DDPSSSYSPYLPPVLEEPSSSFSEAADEDPLPAIEGLQIS--GEAFPGRELQACGYSI 841
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.351 0.158 0.526
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 66,366,892
Number of Sequences: 63676
Number of extensions: 995231
Number of successful extensions: 8510
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 6273
Number of HSP's successfully gapped in prelim test: 261
Number of HSP's that attempted gapping in prelim test: 2030
Number of HSP's gapped (non-prelim): 6843
length of query: 1459
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1350
effective length of database: 5,698,948
effective search space: 7693579800
effective search space used: 7693579800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 65 (29.6 bits)
Medicago: description of AC127020.7


