

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127019.14 + phase: 0 /pseudo
(532 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
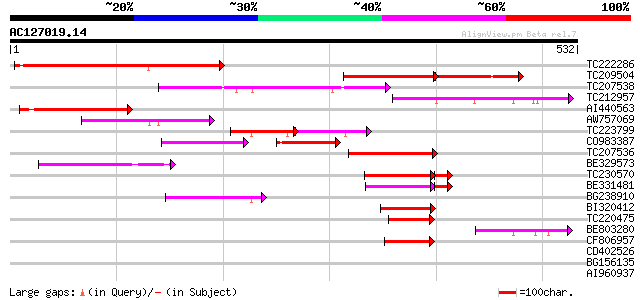
Score E
Sequences producing significant alignments: (bits) Value
TC222286 similar to UP|ML10_ARATH (Q9FKY5) MLO-like protein 10 (... 308 5e-84
TC209504 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (A... 146 8e-51
TC207538 similar to UP|MLO1_ARATH (O49621) MLO-like protein 1 (A... 170 1e-42
TC212957 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (A... 108 6e-24
AI440563 similar to PIR|T02582|T025 H. vulgare Mlo protein homol... 102 4e-22
AW757069 similar to GP|15982791|gb AT5g53760/MGN6_12 {Arabidopsi... 99 5e-21
TC223799 weakly similar to UP|ML12_ARATH (O80961) MLO-like prote... 69 5e-19
CO983387 62 1e-18
TC207536 similar to UP|MLO1_ARATH (O49621) MLO-like protein 1 (A... 79 5e-15
BE329573 76 4e-14
TC230570 similar to UP|MLO6_ARATH (Q94KB7) MLO-like protein 6 (A... 69 2e-12
BE331481 similar to PIR|T02582|T025 H. vulgare Mlo protein homol... 55 1e-08
BG238910 51 1e-06
BI320412 similar to GP|14091574|gb| membrane protein Mlo2 {Arabi... 49 7e-06
TC220475 similar to UP|ML11_ARATH (Q9FI00) MLO-like protein 11 (... 46 4e-05
BE803280 similar to GP|14091584|gb| membrane protein Mlo7 {Arabi... 44 2e-04
CF806957 42 7e-04
CD402526 similar to GP|13784991|gb| seven transmembrane protein ... 38 0.004
BG156135 similar to GP|14091578|gb| membrane protein Mlo4 {Arabi... 37 0.017
AI960937 33 0.32
>TC222286 similar to UP|ML10_ARATH (Q9FKY5) MLO-like protein 10 (AtMlo10),
partial (25%)
Length = 731
Score = 308 bits (788), Expect = 5e-84
Identities = 156/210 (74%), Positives = 175/210 (83%), Gaps = 13/210 (6%)
Frame = +3
Query: 5 VFSCLCLFYGGLSMAASDSRNSSRTLDQTPTWGVAAVCTVFILISIALEKSLHKLGTWLG 64
+FS LC +GGL+MAA +S +SSR LDQTPTW VAAVCTVFIL+SIALEKSLHK+GTWLG
Sbjct: 108 LFSWLC--FGGLAMAAGESSSSSRDLDQTPTWAVAAVCTVFILVSIALEKSLHKVGTWLG 281
Query: 65 KREKKALLEALEKVKAELMILGFISLLLTIGQNYIVRICISEKIANKMLPCPHKYIGGDK 124
+++KKALLEALEKVKAELMILGFISLLLT GQ+YIVRICI EK+A+ MLPCP+KY K
Sbjct: 282 QKKKKALLEALEKVKAELMILGFISLLLTFGQSYIVRICIPEKLADNMLPCPYKYKEDKK 461
Query: 125 KSNS-------------RYLAADTPSFKCSRKGHEPLLSVNGLHQLHILIFFLAVFHVFY 171
S+S RYLAADT SFKCSR+GHEPLLSVNGLHQLHIL+FFLAV HV Y
Sbjct: 462 ASDSEEEHRRKLLSYERRYLAADTTSFKCSREGHEPLLSVNGLHQLHILVFFLAVIHVLY 641
Query: 172 SAVTMLLGRLKIRGWKEWEEETLTHGYEFA 201
SA+TM+LGRLKIRGWK WE ET TH YEFA
Sbjct: 642 SAITMMLGRLKIRGWKAWEAETSTHNYEFA 731
>TC209504 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (AtMlo8),
partial (24%)
Length = 735
Score = 146 bits (369), Expect(2) = 8e-51
Identities = 66/89 (74%), Positives = 76/89 (85%)
Frame = +2
Query: 314 LDSNYSRMALEISERHIVVQGMPLVQGSDRYFWLGQPQLVLHLIHFALFQNSFQITYILW 373
L + + MA+EI+ERH VVQG+PLVQGSDRYFW G+PQLVLHLIHFALFQN+FQITY LW
Sbjct: 20 LQAALANMAIEITERHAVVQGIPLVQGSDRYFWFGRPQLVLHLIHFALFQNAFQITYFLW 199
Query: 374 TWYSFGLENCFRLDYKLAIIKVAFGLYML 402
WYSFGL NCF DYKLA++KVA GL +L
Sbjct: 200 IWYSFGLRNCFHADYKLAVVKVALGLGVL 286
Score = 72.8 bits (177), Expect(2) = 8e-51
Identities = 50/85 (58%), Positives = 57/85 (66%), Gaps = 2/85 (2%)
Frame = +3
Query: 400 YMLL*HRWVQG*KNQYLMNKHPRR*RNGT*L*KRSM-*S*ESPL*EPWMVAPLLDQQCSP 458
YMLL RW Q *KNQYL NKH RR*RNGT L*KRS *+ E P EPWM A L+ QQ +
Sbjct: 315 YMLLLLRWSQE*KNQYLTNKHRRR*RNGTWL*KRSKE*NLEIPRCEPWMEAALI-QQYTL 491
Query: 459 LDQHFTVSRLLATRHDL-QPLMNKM 482
L H+TV++L T+ L Q M KM
Sbjct: 492 LAPHYTVTKLPVTQLTLYQTTMTKM 566
>TC207538 similar to UP|MLO1_ARATH (O49621) MLO-like protein 1 (AtMlo1) (MLO
protein homolog 1) (AtMLO-H1), partial (42%)
Length = 737
Score = 170 bits (431), Expect = 1e-42
Identities = 98/235 (41%), Positives = 138/235 (58%), Gaps = 17/235 (7%)
Frame = +1
Query: 140 CSRKGHEPLLSVNGLHQLHILIFFLAVFHVFYSAVTMLLGRLKIRGWKEWEEETLTHGYE 199
CSRK PLLSV LH LHI IF LAV HV +S +T++ G +IR WK WE+ YE
Sbjct: 40 CSRKHKVPLLSVEALHHLHIFIFVLAVVHVSFSVLTVVFGGARIRQWKHWEDSIAKQNYE 219
Query: 200 FANDAAKLRLTH--ETSFVRAHTSFWTRV-----YIGCFFRQFYRSVRKVDYQTLRNGFI 252
+ K ++TH + F+R + + + ++ F +QFY SV K DY TLR+GFI
Sbjct: 220 -TDRVLKPKVTHVHQHDFIRGRFAGFGKDSAIVGWLLSFLKQFYGSVTKSDYVTLRHGFI 396
Query: 253 NVHLAPGSKFNFQKYIKRSLEDDFKVVVGVSPVLWASAVVFLLLNIDAY----------I 302
H KFNF KY+ R+LEDDFK VVG+S LW V+FLLLNI+ + +
Sbjct: 397 MTHCRTNPKFNFHKYMIRALEDDFKQVVGISWYLWLFVVIFLLLNINGWHTYFWIAFIPV 576
Query: 303 IFRMAYSNMGILDSNYSRMALEISERHIVVQGMPLVQGSDRYFWLGQPQLVLHLI 357
I +A L+ +++A E++E+H ++G +VQ SD +FW +P++VL LI
Sbjct: 577 ILLLAVGTK--LEHIITQLAHEVAEKHAAIEGDLVVQPSDDHFWFHRPRVVLFLI 735
>TC212957 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (AtMlo8),
partial (15%)
Length = 780
Score = 108 bits (270), Expect = 6e-24
Identities = 86/196 (43%), Positives = 110/196 (55%), Gaps = 26/196 (13%)
Frame = +1
Query: 360 ALFQNSFQITYILWTWYSFGLENCFRLDYKLAIIKVAFGL------------YMLL*HRW 407
ALFQN+FQITYI WY G E L K G YMLL* RW
Sbjct: 1 ALFQNAFQITYIXXIWYXXG*ETVSVLTTSXX**K*L*GF*CYASAAISPFHYMLL*LRW 180
Query: 408 VQG*KNQYLMNKHPRR*RNGT*L*KRS---M*S*ESPL*EPWMVAPL-LDQQCSPLDQHF 463
VQG*K QYL +K R *RNGT L +RS * *ES + E WM APL + QQC+PL H+
Sbjct: 181 VQG*KQQYLTSKQTRL*RNGTWLRRRSREEQ*R*ESRVHESWMEAPLVILQQCTPLAPHY 360
Query: 464 TVSRLLAT-----RHDLQPLMNKMNMNYLIMS--*WK---LT*L*ELIMMSKKQKIGIII 513
TVS+LLAT + + + MNMN +++S W+ L*ELIM + ++ I
Sbjct: 361 TVSKLLATQPAPHQQRTRIKIKIMNMNPMVLSCLRWRRKQQASL*ELIMATNNKQNIDKI 540
Query: 514 VREKPIMMISLSSSMS 529
VREKP +++ ++S +S
Sbjct: 541 VREKPTVVVKVNSHLS 588
>AI440563 similar to PIR|T02582|T025 H. vulgare Mlo protein homolog
[imported] - Arabidopsis thaliana, partial (15%)
Length = 376
Score = 102 bits (254), Expect = 4e-22
Identities = 56/106 (52%), Positives = 69/106 (64%)
Frame = +1
Query: 10 CLFYGGLSMAASDSRNSSRTLDQTPTWGVAAVCTVFILISIALEKSLHKLGTWLGKREKK 69
CL GG M S+ L+ TPTW VA VC V + ISI +E L +LG WL K+ KK
Sbjct: 64 CLECGG*KM----SKVLQAKLEATPTWAVAVVCFVMLAISILIEHILEELGKWLKKKHKK 231
Query: 70 ALLEALEKVKAELMILGFISLLLTIGQNYIVRICISEKIANKMLPC 115
AL EALEKVK ELM+LGFISLLL + Q++I ICI + +A+ PC
Sbjct: 232 ALHEALEKVKGELMLLGFISLLLVMFQDHISNICIPKSVASTWHPC 369
>AW757069 similar to GP|15982791|gb AT5g53760/MGN6_12 {Arabidopsis thaliana},
partial (22%)
Length = 443
Score = 99.0 bits (245), Expect = 5e-21
Identities = 58/141 (41%), Positives = 75/141 (53%), Gaps = 16/141 (11%)
Frame = +2
Query: 68 KKALLEALEKVKAELMILGFISLLLTIGQNYIVRICISEKIANKML-PCPHKYIGGDKKS 126
+K LL ALEK+K ELM+LGFISLLLT I ICI K N PC I + +
Sbjct: 5 RKPLLAALEKMKEELMLLGFISLLLTATSRMIANICIPSKFYNSAFAPCTRSEIDEEMED 184
Query: 127 NSR-----YLAADTPSF----------KCSRKGHEPLLSVNGLHQLHILIFFLAVFHVFY 171
N +A+ P ++G+EP +S GL QLH IF +AV H+ Y
Sbjct: 185 NGSEERKLLMASSYPHLVRRMLNGINSSTCKEGYEPFVSYEGLEQLHRFIFVMAVTHISY 364
Query: 172 SAVTMLLGRLKIRGWKEWEEE 192
S +TMLL +KI W+ WE+E
Sbjct: 365 SCLTMLLAIVKIHSWRVWEDE 427
>TC223799 weakly similar to UP|ML12_ARATH (O80961) MLO-like protein 12
(AtMlo12) (AtMlo18), partial (25%)
Length = 449
Score = 68.9 bits (167), Expect(2) = 5e-19
Identities = 35/70 (50%), Positives = 46/70 (65%), Gaps = 6/70 (8%)
Frame = +1
Query: 208 RLTHETSFVRAHTSFWTR----VYIGCFFRQFYRSVRKVDYQTLRNGFINVHLAPG--SK 261
R +T+F R H S W+R ++I FFRQFY S+ KVDY LR+GF+ HL P +K
Sbjct: 7 RFARDTTFGRRHLSSWSRSPGSLWIVSFFRQFYGSLNKVDYMALRHGFLVAHLTPANEAK 186
Query: 262 FNFQKYIKRS 271
F+FQ YIKR+
Sbjct: 187 FDFQNYIKRT 216
Score = 43.9 bits (102), Expect(2) = 5e-19
Identities = 28/79 (35%), Positives = 43/79 (53%), Gaps = 8/79 (10%)
Frame = +2
Query: 269 KRSLEDDFKVVVGVSPVLWASAVVFLLLNIDA-YIIFRMAYSNMGI-------LDSNYSR 320
+ L++DF VVG++P +W AV+ LL N Y F + + + I L +
Sbjct: 209 REHLDEDFAAVVGITPTIWFFAVLILLTNTHGWYSYFWIPFIPVIIILLVGTKLQMIITE 388
Query: 321 MALEISERHIVVQGMPLVQ 339
MAL+I +R VV+G PLV+
Sbjct: 389 MALKIQDRGEVVKGAPLVE 445
>CO983387
Length = 856
Score = 62.4 bits (150), Expect(2) = 1e-18
Identities = 33/82 (40%), Positives = 47/82 (57%)
Frame = -1
Query: 143 KGHEPLLSVNGLHQLHILIFFLAVFHVFYSAVTMLLGRLKIRGWKEWEEETLTHGYEFAN 202
+G LLS G+ + IF+LA HV S +T LG KIR + WE ET T Y+FA
Sbjct: 802 RGKVSLLSREGVREQQYFIFYLARCHVVSSFLTFGLGLAKIRRSESWEGETRTLEYQFAY 623
Query: 203 DAAKLRLTHETSFVRAHTSFWT 224
D + +LT +T F + H ++W+
Sbjct: 622 DPRRYQLTGQTPFGKRHLNYWS 557
Score = 49.3 bits (116), Expect(2) = 1e-18
Identities = 27/60 (45%), Positives = 38/60 (63%)
Frame = -3
Query: 251 FINVHLAPGSKFNFQKYIKRSLEDDFKVVVGVSPVLWASAVVFLLLNIDAYIIFRMAYSN 310
F NV LA S F+FQKYI+R+LE DF VVVG+ +W +V+++ N +A YS+
Sbjct: 551 FCNV-LAKESNFDFQKYIERALEKDFGVVVGLRWWIWIFSVLYIFFNANARFCSAAFYSH 375
Score = 33.5 bits (75), Expect = 0.24
Identities = 19/50 (38%), Positives = 27/50 (54%)
Frame = -1
Query: 347 LGQPQLVLHLIHFALFQNSFQITYILWTWYSFGLENCFRLDYKLAIIKVA 396
+G P+L+LHLI F LFQ FG+ +CF + + II+VA
Sbjct: 202 IGCPKLLLHLISFILFQ------------IRFGIRSCFHQETENIIIRVA 89
>TC207536 similar to UP|MLO1_ARATH (O49621) MLO-like protein 1 (AtMlo1) (MLO
protein homolog 1) (AtMLO-H1), partial (28%)
Length = 874
Score = 79.0 bits (193), Expect = 5e-15
Identities = 29/83 (34%), Positives = 55/83 (65%)
Frame = +3
Query: 319 SRMALEISERHIVVQGMPLVQGSDRYFWLGQPQLVLHLIHFALFQNSFQITYILWTWYSF 378
+++A E++E+H ++G +VQ SD +FW +P +VL LIHF LFQN+F+I + W W ++
Sbjct: 51 TQLAQEVAEKHAAIEGDLVVQPSDEHFWFHRPHVVLFLIHFILFQNAFEIAFFFWIWVTY 230
Query: 379 GLENCFRLDYKLAIIKVAFGLYM 401
G ++C + + ++ G+++
Sbjct: 231 GFDSCIMGQVRYIVPRLVIGVFI 299
>BE329573
Length = 400
Score = 75.9 bits (185), Expect = 4e-14
Identities = 47/130 (36%), Positives = 73/130 (56%), Gaps = 2/130 (1%)
Frame = +3
Query: 28 RTLDQTPTWGVAAVCTVFILISIALEKSLHKLGTWLGKREKKALLEALEKVKAELMILGF 87
R+L +TPT+ VA V TV + +S + +L KL WL + ++K+LL AL+K+K ELM+ G
Sbjct: 9 RSLAETPTYAVATVITVLVSLSFLFQGTLKKLVKWLDRTKRKSLLSALDKIKEELMLFGL 188
Query: 88 ISLLLTIGQNYIVRICI-SEKIANKMLPCPHKYIGGDKKSNSRYLA-ADTPSFKCSRKGH 145
+SLL+ ++ +IC+ S +++ PC +K S R++ PS K KG
Sbjct: 189 LSLLMGHWIIFVAKICVKSSVLSSTFFPC-----AMEKNSVKRFVGLGSEPSNKTVLKGQ 353
Query: 146 EPLLSVNGLH 155
NGLH
Sbjct: 354 ----MNNGLH 371
>TC230570 similar to UP|MLO6_ARATH (Q94KB7) MLO-like protein 6 (AtMlo6),
partial (24%)
Length = 957
Score = 69.3 bits (168), Expect(2) = 2e-12
Identities = 27/66 (40%), Positives = 44/66 (65%)
Frame = +3
Query: 334 GMPLVQGSDRYFWLGQPQLVLHLIHFALFQNSFQITYILWTWYSFGLENCFRLDYKLAII 393
G+PLVQ D FW +P L+L+LI+F LFQN+FQ+ + W+ FG+++CF + +I
Sbjct: 3 GVPLVQPGDHLFWFNRPGLILYLINFVLFQNAFQLAFFAWSALQFGIKSCFHSHTEDVVI 182
Query: 394 KVAFGL 399
++ G+
Sbjct: 183 RITMGV 200
Score = 20.8 bits (42), Expect(2) = 2e-12
Identities = 10/17 (58%), Positives = 11/17 (63%)
Frame = +1
Query: 399 LYMLL*HRWVQG*KNQY 415
L L *HRWVQ * +Y
Sbjct: 235 LSTL**HRWVQQ*SQRY 285
>BE331481 similar to PIR|T02582|T025 H. vulgare Mlo protein homolog
[imported] - Arabidopsis thaliana, partial (9%)
Length = 424
Score = 55.5 bits (132), Expect(2) = 1e-08
Identities = 22/65 (33%), Positives = 38/65 (57%)
Frame = +2
Query: 335 MPLVQGSDRYFWLGQPQLVLHLIHFALFQNSFQITYILWTWYSFGLENCFRLDYKLAIIK 394
+P VQ D FW +P L+L+LI+F LFQN+F + + W +++CF + +I+
Sbjct: 2 VPSVQPGDHLFWFNRPGLILYLINFGLFQNAFHLAFFAWAALQCEIKSCFHSLTEEVVIR 181
Query: 395 VAFGL 399
+ G+
Sbjct: 182 ITMGV 196
Score = 21.9 bits (45), Expect(2) = 1e-08
Identities = 10/17 (58%), Positives = 11/17 (63%)
Frame = +1
Query: 399 LYMLL*HRWVQG*KNQY 415
L L *HRWVQ * +Y
Sbjct: 232 LSTLY*HRWVQQ*SQRY 282
>BG238910
Length = 310
Score = 51.2 bits (121), Expect = 1e-06
Identities = 33/102 (32%), Positives = 50/102 (48%), Gaps = 7/102 (6%)
Frame = +2
Query: 147 PLLSVNGLHQLHILIFFLAV-FHVFYSAVTMLLGRLKIRGWKEWEEETLTHGYEFAND-A 204
PLLS+ LH LHI IF + + V +S +T + G +IR WK WE+ YE
Sbjct: 5 PLLSLEALHHLHIFIFGPRLSYTVSFSLLTGVFGGARIRQWKHWEDSIAKQNYETGRVLK 184
Query: 205 AKLRLTHETSFVRAHTSFWTR-----VYIGCFFRQFYRSVRK 241
K+ H+ F+R + + + ++ F +Q Y SV K
Sbjct: 185 PKVTQVHQHDFIRGRFAGFDKDSAIVGWVLSFLKQVYGSVDK 310
>BI320412 similar to GP|14091574|gb| membrane protein Mlo2 {Arabidopsis
thaliana}, partial (12%)
Length = 213
Score = 48.5 bits (114), Expect = 7e-06
Identities = 19/51 (37%), Positives = 34/51 (66%)
Frame = +1
Query: 349 QPQLVLHLIHFALFQNSFQITYILWTWYSFGLENCFRLDYKLAIIKVAFGL 399
+P+L+L LIH LFQN+FQ+ + W+ Y F +++CF + +I++ G+
Sbjct: 1 RPRLLLFLIHLVLFQNAFQLAFFSWSTYEFSVKSCFHETTEDNVIRLVTGV 153
>TC220475 similar to UP|ML11_ARATH (Q9FI00) MLO-like protein 11 (AtMlo11),
partial (24%)
Length = 749
Score = 46.2 bits (108), Expect = 4e-05
Identities = 18/43 (41%), Positives = 27/43 (61%)
Frame = +3
Query: 356 LIHFALFQNSFQITYILWTWYSFGLENCFRLDYKLAIIKVAFG 398
LIHF LFQN+F++ W W+ FG +CF ++ L I++ G
Sbjct: 3 LIHFILFQNAFELASFFWFWWQFGYYSCFIRNHLLLYIRLILG 131
>BE803280 similar to GP|14091584|gb| membrane protein Mlo7 {Arabidopsis
thaliana}, partial (3%)
Length = 332
Score = 43.9 bits (102), Expect = 2e-04
Identities = 40/103 (38%), Positives = 62/103 (59%), Gaps = 12/103 (11%)
Frame = +1
Query: 438 *ESPL*EPWMVAPLLD-QQCSP-LDQHFTVSRLLAT--RHDLQPLMNKMNMNYLIMS*W- 492
*ES + E WM APL+ QQC+P L H+TVS+L AT R ++ ++ MNMN +++S +
Sbjct: 16 *ESRVHESWMEAPLVILQQCTPPLFPHYTVSKLPATQQRTRIKIMIMIMNMNPMMLSCFL 195
Query: 493 ----KLT*L*ELIMMS---KKQKIGIIIVREKPIMMISLSSSM 528
+L L*E + + K+ K IVREKP + + ++S +
Sbjct: 196 CRRKQLASL*EWTVATNNNKQNKNIDNIVREKPTVAVKVNSHL 324
>CF806957
Length = 688
Score = 42.0 bits (97), Expect = 7e-04
Identities = 17/47 (36%), Positives = 29/47 (61%)
Frame = +2
Query: 352 LVLHLIHFALFQNSFQITYILWTWYSFGLENCFRLDYKLAIIKVAFG 398
L+L LIH FQN+FQ+ + W+ Y F +++CF + +I++ G
Sbjct: 11 LLLFLIHLVPFQNAFQLAFFSWSTYEFSVKSCFHETTEDNVIRLVTG 151
>CD402526 similar to GP|13784991|gb| seven transmembrane protein Mlo9 {Zea
mays}, partial (6%)
Length = 226
Score = 38.1 bits (87), Expect(2) = 0.004
Identities = 14/36 (38%), Positives = 26/36 (71%)
Frame = -1
Query: 262 FNFQKYIKRSLEDDFKVVVGVSPVLWASAVVFLLLN 297
++F Y+ RS++++F+ +VGVS +LW A+ + LN
Sbjct: 154 YDFHNYMLRSMDEEFRDIVGVSVLLWVYAICCIFLN 47
Score = 20.4 bits (41), Expect(2) = 0.004
Identities = 9/23 (39%), Positives = 11/23 (47%)
Frame = -3
Query: 243 DYQTLRNGFINVHLAPGSKFNFQ 265
DY LR GFI + S N +
Sbjct: 218 DYMALRLGFITCSIESWSSHNIR 150
>BG156135 similar to GP|14091578|gb| membrane protein Mlo4 {Arabidopsis
thaliana}, partial (5%)
Length = 414
Score = 37.4 bits (85), Expect = 0.017
Identities = 19/56 (33%), Positives = 28/56 (49%), Gaps = 2/56 (3%)
Frame = +1
Query: 168 HVFYSAVTMLLGRLKIRGWKEWEEETLTHGYEFANDAAK--LRLTHETSFVRAHTS 221
H+ YS + + L +KI W+ WE E T + D ++ RL +FV HTS
Sbjct: 4 HITYSFIAVALAMIKIYSWRTWENEAKTIAVQSIQDTSQGTSRLRRLNTFVFHHTS 171
>AI960937
Length = 320
Score = 33.1 bits (74), Expect = 0.32
Identities = 17/40 (42%), Positives = 26/40 (64%), Gaps = 2/40 (5%)
Frame = +3
Query: 2 FKRVFSCLC--LFYGGLSMAASDSRNSSRTLDQTPTWGVA 39
++ +F CL L +++A+S+S S+ LDQTPTW VA
Sbjct: 201 YRNLFLCLASWLCCCCVAVASSESSIGSKDLDQTPTWAVA 320
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.338 0.147 0.474
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,702,327
Number of Sequences: 63676
Number of extensions: 380341
Number of successful extensions: 3423
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 3373
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3409
length of query: 532
length of database: 12,639,632
effective HSP length: 102
effective length of query: 430
effective length of database: 6,144,680
effective search space: 2642212400
effective search space used: 2642212400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC127019.14


