

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126794.6 - phase: 0
(926 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
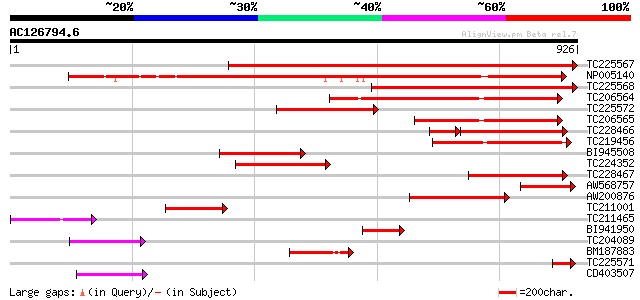
Sequences producing significant alignments: (bits) Value
TC225567 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp pro... 1042 0.0
NP005140 heat shock protein 663 0.0
TC225568 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp pro... 630 0.0
TC206564 homologue to UP|Q9LF37 (Q9LF37) ClpB heat shock protein... 370 e-102
TC225572 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp pro... 295 8e-80
TC206565 similar to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-l... 259 3e-69
TC228466 similar to UP|ERD1_ARATH (P42762) ERD1 protein, chlorop... 180 7e-66
TC219456 homologue to UP|Q9LLI0 (Q9LLI0) ClpB, partial (24%) 216 4e-56
BI945508 homologue to PIR|T51523|T51 clpB heat shock protein-lik... 199 6e-51
TC224352 homologue to UP|Q9LF37 (Q9LF37) ClpB heat shock protein... 195 9e-50
TC228467 weakly similar to UP|ERD1_ARATH (P42762) ERD1 protein, ... 164 2e-40
AW568757 similar to SP|P35100|CLPA_ ATP-dependent clp protease A... 152 7e-37
AW200876 weakly similar to GP|4105131|gb| ClpC protease {Spinaci... 146 4e-35
TC211001 homologue to UP|Q9LF37 (Q9LF37) ClpB heat shock protein... 140 3e-33
TC211465 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp prote... 93 5e-19
BI941950 similar to PIR|T51523|T51 clpB heat shock protein-like ... 82 1e-15
TC204089 70 4e-12
BM187883 homologue to GP|9651530|gb| ClpB {Phaseolus lunatus}, p... 70 4e-12
TC225571 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp prote... 68 2e-11
CD403507 65 1e-10
>TC225567 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (61%)
Length = 2077
Score = 1042 bits (2695), Expect = 0.0
Identities = 530/569 (93%), Positives = 555/569 (97%)
Frame = +3
Query: 358 KKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYR 417
KKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYR
Sbjct: 6 KKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYR 185
Query: 418 KHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQY 477
KHIEKDPALERRFQPVKVPEPTV ETIQILKGLRERYEIHHKLRYTDEALVAAA+LS+QY
Sbjct: 186 KHIEKDPALERRFQPVKVPEPTVDETIQILKGLRERYEIHHKLRYTDEALVAAAQLSYQY 365
Query: 478 ISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGE 537
ISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEAR L+KEVRQI+KEK+EAVRNQ+FEKAGE
Sbjct: 366 ISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARELDKEVRQIIKEKEEAVRNQDFEKAGE 545
Query: 538 LRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDE 597
LRD+EMDLK QIS L+EK KEM+KAE+EAGD G +VTE DIQHIV+SWTGIPV+KVS DE
Sbjct: 546 LRDREMDLKAQISTLVEKGKEMSKAETEAGDEGPIVTEADIQHIVSSWTGIPVEKVSTDE 725
Query: 598 SDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELA 657
SDRLLKME+TLHKR+IGQ EAV+AISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELA
Sbjct: 726 SDRLLKMEETLHKRVIGQDEAVKAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELA 905
Query: 658 KALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVL 717
KALA+YYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVL
Sbjct: 906 KALAAYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVL 1085
Query: 718 FDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFD 777
FDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGR+IGFD
Sbjct: 1086FDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRKIGFD 1265
Query: 778 LDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFE 837
LDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVF+
Sbjct: 1266LDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFD 1445
Query: 838 RLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSV 897
RLK K+IEL VTERFR+RVV+EGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSV
Sbjct: 1446RLKVKDIELQVTERFRDRVVEEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSV 1625
Query: 898 IVDADSDGNVIVLNGSTGAPDSLPDALPV 926
IVD DSDGNVIVLNGS+GAP+SLP+ LPV
Sbjct: 1626IVDVDSDGNVIVLNGSSGAPESLPETLPV 1712
>NP005140 heat shock protein
Length = 2736
Score = 663 bits (1711), Expect = 0.0
Identities = 373/874 (42%), Positives = 539/874 (60%), Gaps = 61/874 (6%)
Frame = +1
Query: 97 ERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVE 156
E+FT K + + A E A GH + + LI + GI + S G + AR
Sbjct: 10 EKFTHKTNEALASAHELAMSSGHAQLTPIHLAHALISDPNGIFVLAINSAGGGEESARA- 186
Query: 157 VEKIIGRGSGFVAV------EIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREG 210
VE+++ + + E+P + R + + + G + + L+LG+L +
Sbjct: 187 VERVLNQALKKLPCQSPPPDEVPASTNLVRAIRRAQAAQKSRGDTRLAVDQLILGILEDS 366
Query: 211 EGVAARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLA 270
+ +L+ G + ++V ++ G+ G V S S + L+ YG +L + A
Sbjct: 367 Q--IGDLLKEAGVAVAKVESEVDKLRGKE----GKKVESASGDTNFQALKTYGRDLVEQA 528
Query: 271 EEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETI 330
GKLDPV+GR +I RV +IL RRTKNNP L+GEPGVGKTA+ EGLAQRI GDVP +
Sbjct: 529 --GKLDPVIGRDEEIRRVVRILSRRTKNNPVLVGEPGVGKTAVVEGLAQRIVRGDVPSNL 702
Query: 331 EGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSD-EIILFIDEVHTLIGAGAAEGA 389
++I LDMG LVAG KYRGEFEERLK +++E+++++ ++ILFIDE+H ++GAG EG+
Sbjct: 703 ADVRLIALDMGALVAGAKYRGEFEERLKAVLKEVEEAEGKVILFIDEIHLVLGAGRTEGS 882
Query: 390 IDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKG 449
+DAAN+ KP LARG+L+CIGATTL+EYRK++EKD A ERRFQ V V EP+V +TI IL+G
Sbjct: 883 MDAANLFKPMLARGQLRCIGATTLEEYRKYVEKDAAFERRFQQVFVAEPSVVDTISILRG 1062
Query: 450 LRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEA 509
L+ERYE HH +R D ALV AA+LS++YI+ R LPDKAIDL+DEA + VR+Q PEE
Sbjct: 1063LKERYEGHHGVRIQDRALVMAAQLSNRYITGRHLPDKAIDLVDEACANVRVQLDSQPEEI 1242
Query: 510 RGLEK-------EVRQIVKEKDEAVRNQEFEKAGELRD-------------KEMDLKTQI 549
LE+ E+ + KEKD+A + + E EL D KE + +I
Sbjct: 1243DNLERKRMQLEVELHALEKEKDKASKARLVEVRKELDDLRDKLQPLMMKYRKEKERVDEI 1422
Query: 550 SALIEKNKEMNKAESEA--------------GDVGALVTEV------------------- 576
L +K +E+ A EA G + + T +
Sbjct: 1423RRLKKKREELLFALQEAERRYDLARAADLRYGAIQEVETAIQQLEGSTEENLMLTETVGP 1602
Query: 577 -DIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKN 635
I +V+ WTGIPV ++ +E +RL+ + D LH R++GQ +AV A++ A+ R+R GL
Sbjct: 1603EQIAEVVSRWTGIPVTRLGQNEKERLIGLGDRLHSRVVGQDQAVNAVAEAVLRSRAGLGR 1782
Query: 636 PNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGY 695
P +P SF+F GPTGVGK+ELAKALA F +E ++R+DMSE+ME+H+VS+LIG+PPGY
Sbjct: 1783PQQPTGSFLFLGPTGVGKTELAKALAEQLFDNENQLVRIDMSEYMEQHSVSRLIGAPPGY 1962
Query: 696 VGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTL 755
VG+ EGGQLTEAVRRRPY+VVLFDE+EKAH VFN +LQ+L+DGRLTD +GRTVDF+NT+
Sbjct: 1963VGHEEGGQLTEAVRRRPYSVVLFDEVEKAHTSVFNTLLQVLDDGRLTDGQGRTVDFRNTV 2142
Query: 756 LIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIV 815
+IMTSN+G+ + G + + V +E+++ FRPE LNRLDE++V
Sbjct: 2143IIMTSNLGAEHLLSG----------LSGKCTMQVARDRVMQEVRRQFRPELLNRLDEIVV 2292
Query: 816 FRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIM 875
F L+ +++++A + +K+V RL K I L+VT+ + ++ E Y+P YGARP+RR +
Sbjct: 2293FDPLSHDQLRKVARLQMKDVASRLAEKGIALAVTDAALDYILSESYDPVYGARPIRRWLE 2472
Query: 876 RLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIV 909
+ + ++ ++ EI E +V +DA +G +V
Sbjct: 2473KKVVTELSRMLVREEIDENSTVYIDAGPNGGELV 2574
>TC225568 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (36%)
Length = 1287
Score = 630 bits (1625), Expect = 0.0
Identities = 322/336 (95%), Positives = 332/336 (97%)
Frame = +2
Query: 591 DKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTG 650
+KVS DESDRLLKME+TLHKR+IGQ EAV+AISRAIRRARVGLKNPNRPIASFIFSGPTG
Sbjct: 5 EKVSTDESDRLLKMEETLHKRVIGQDEAVKAISRAIRRARVGLKNPNRPIASFIFSGPTG 184
Query: 651 VGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRR 710
VGKSELAKALA+YYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRR
Sbjct: 185 VGKSELAKALAAYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRR 364
Query: 711 RPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKG 770
RPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKG
Sbjct: 365 RPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKG 544
Query: 771 GRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADI 830
GR+IGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADI
Sbjct: 545 GRKIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADI 724
Query: 831 MLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLARE 890
MLKEVF+RLK KEI+LSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLARE
Sbjct: 725 MLKEVFDRLKAKEIDLSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLARE 904
Query: 891 IKEGDSVIVDADSDGNVIVLNGSTGAPDSLPDALPV 926
IKEGDSVIVDADSDGNVIVLNGS+GAPDSL +ALPV
Sbjct: 905 IKEGDSVIVDADSDGNVIVLNGSSGAPDSLEEALPV 1012
>TC206564 homologue to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-like,
partial (40%)
Length = 1497
Score = 370 bits (949), Expect = e-102
Identities = 191/383 (49%), Positives = 273/383 (70%), Gaps = 2/383 (0%)
Frame = +2
Query: 522 EKDEAVRNQEFEKAGELRDKEMD-LKTQI-SALIEKNKEMNKAESEAGDVGALVTEVDIQ 579
E +A R + +A EL+ ++ L+ Q+ SA E ++ MN +S + VT DI
Sbjct: 8 EIQQAEREYDLNRAAELKYGSLNSLQRQLESAEKELDEYMNSGKSMLREE---VTGNDIA 178
Query: 580 HIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRP 639
IV+ WTGIPV K+ E ++LL +E+ LHKR++GQ AV+AI+ AI+R+R GL +P+RP
Sbjct: 179 EIVSKWTGIPVSKLQQSEREKLLHLEEVLHKRVVGQDPAVKAIAEAIQRSRAGLSDPHRP 358
Query: 640 IASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYT 699
IASF+F GPTGVGK+ELAKALA+Y F +EEA++R+DMSE+ME+H VS+LIG+PPGYVGY
Sbjct: 359 IASFMFMGPTGVGKTELAKALAAYLFNTEEALVRIDMSEYMEKHAVSRLIGAPPGYVGYE 538
Query: 700 EGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMT 759
EGGQLTE VRRRPY V+LFDEIEKAH DVFN+ LQIL+DGR+TDS+GRTV F NT++IMT
Sbjct: 539 EGGQLTEIVRRRPYAVILFDEIEKAHADVFNVFLQILDDGRVTDSQGRTVSFTNTVIIMT 718
Query: 760 SNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQL 819
SNVGS I + D D K+ +Y IK V + + FRPEF+NR+DE IVF+ L
Sbjct: 719 SNVGSQYI------LNTDDDTTPKELAYETIKQRVMDAARSIFRPEFMNRVDEYIVFQPL 880
Query: 820 TKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLE 879
+ ++ I + L+ V +R+ +++++ VT+ + + GY+P+YGARP++R I + +E
Sbjct: 881 DREQISSIVRLQLERVQKRIADRKMKIQVTDAAVQLLGSLGYDPNYGARPVKRVIQQNVE 1060
Query: 880 DSMAEKMLAREIKEGDSVIVDAD 902
+ +A+ +L E KE D++++D +
Sbjct: 1061NELAKGILRGEFKEEDAILIDTE 1129
>TC225572 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (18%)
Length = 511
Score = 295 bits (754), Expect = 8e-80
Identities = 149/167 (89%), Positives = 159/167 (94%)
Frame = +3
Query: 436 PEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAG 495
PEPTV ETIQILKGLRERYEIHHKL YTD+ALVAAA+LSHQYISDRFLPDKAIDLIDEAG
Sbjct: 9 PEPTVNETIQILKGLRERYEIHHKLHYTDDALVAAAQLSHQYISDRFLPDKAIDLIDEAG 188
Query: 496 SRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEK 555
SRVRLQHAQLPEEAR L+KEVRQIVKEK+E+VRNQ+FEKAGELRDKEMDLK QISALIEK
Sbjct: 189 SRVRLQHAQLPEEARELDKEVRQIVKEKEESVRNQDFEKAGELRDKEMDLKAQISALIEK 368
Query: 556 NKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDESDRLL 602
KEM+KAESEAGD G +VTEVDIQHIV+SWTGIPV+KVS DESDRLL
Sbjct: 369 GKEMSKAESEAGDEGPMVTEVDIQHIVSSWTGIPVEKVSTDESDRLL 509
>TC206565 similar to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-like, partial
(25%)
Length = 1078
Score = 259 bits (663), Expect = 3e-69
Identities = 127/241 (52%), Positives = 178/241 (73%)
Frame = +2
Query: 662 SYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEI 721
SY F +EEA++R+DMSE+ME+HTVS+LIG+PPGYVGY EGGQLTE VRRRPY V+LFDEI
Sbjct: 2 SYLFNTEEALVRIDMSEYMEKHTVSRLIGAPPGYVGYEEGGQLTETVRRRPYAVILFDEI 181
Query: 722 EKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYD 781
EKAH DVFN+ LQIL+DGR+TDS+GRTV F NT++IMTSNVGS I + D D
Sbjct: 182 EKAHSDVFNVFLQILDDGRVTDSQGRTVSFTNTVIIMTSNVGSQYI------LNTDDDTV 343
Query: 782 EKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKT 841
K+S+Y IK V + + FRPEF+NR+DE IVF+ L + ++ I + L+ V +R+
Sbjct: 344 PKESAYETIKQRVMDAARSIFRPEFMNRVDEYIVFQPLDRNQISSIVRLQLERVQKRIAD 523
Query: 842 KEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDA 901
+++++ VTE + + GY+P+YGARP++R I + +E+ +A+ +L E KE D+++VD
Sbjct: 524 RKMKIQVTEAAIQLLGSLGYDPNYGARPVKRVIQQNVENELAKGILRGEFKEEDTILVDT 703
Query: 902 D 902
+
Sbjct: 704 E 706
>TC228466 similar to UP|ERD1_ARATH (P42762) ERD1 protein, chloroplast
precursor, partial (23%)
Length = 936
Score = 180 bits (456), Expect(2) = 7e-66
Identities = 90/176 (51%), Positives = 130/176 (73%), Gaps = 1/176 (0%)
Frame = +2
Query: 737 EDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRR-IGFDLDYDEKDSSYNRIKSLVT 795
+DG+LTDS+GR V FKN L++MTSNVGSS I KG IGF L D+K +SYN +KS+V
Sbjct: 155 QDGQLTDSQGRRVSFKNALVVMTSNVGSSAIAKGRHNSIGF-LIPDDKTTSYNGLKSMVI 331
Query: 796 EELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRER 855
EEL+ YFRPE LNR+DE++VF+ L K ++ +I D++L+++ +R+ + + + V+E +
Sbjct: 332 EELRSYFRPELLNRIDEVVVFQPLEKSQLLQILDLLLQDMKKRVLSLGVHVKVSEAVKNL 511
Query: 856 VVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVLN 911
V +GYNP+YGARPLRRAI L+ED ++E L E K+GD+V++D D++GN V N
Sbjct: 512 VCQQGYNPTYGARPLRRAITSLIEDPLSEAFLYGECKQGDTVLIDLDANGNPFVTN 679
Score = 90.5 bits (223), Expect(2) = 7e-66
Identities = 40/51 (78%), Positives = 48/51 (93%)
Frame = +1
Query: 686 SKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQIL 736
SKLIGSPPGYVGY EGG LTEA+RR+P+T++L DEIEKAHPD+FN++LQIL
Sbjct: 1 SKLIGSPPGYVGYGEGGVLTEAIRRKPFTLLLLDEIEKAHPDIFNILLQIL 153
>TC219456 homologue to UP|Q9LLI0 (Q9LLI0) ClpB, partial (24%)
Length = 871
Score = 216 bits (550), Expect = 4e-56
Identities = 112/227 (49%), Positives = 160/227 (70%)
Frame = +1
Query: 691 SPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVD 750
+PPGYVGY EGGQLTE VRRRPY+VVLFDEIEKAH DVFN++LQ+L+DGR+TDS+GRTV
Sbjct: 1 APPGYVGYEEGGQLTEVVRRRPYSVVLFDEIEKAHHDVFNILLQLLDDGRITDSQGRTVS 180
Query: 751 FKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRL 810
F N ++IMTSN+GS I R D+K + Y+++K V E +Q F PEF+NR+
Sbjct: 181 FTNCVVIMTSNIGSHYILDTLRS-----TQDDKTAVYDQMKRQVVELARQTFHPEFMNRI 345
Query: 811 DEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPL 870
DE IVF+ L ++ +I ++ ++ V RLK K+I+L TE+ + + G++P++GARP+
Sbjct: 346 DEYIVFQPLDSEQISKIVELQMERVKNRLKQKKIDLHYTEKAVKLLGVLGFDPNFGARPV 525
Query: 871 RRAIMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVLNGSTGAP 917
+R I +L+E+ +A +L + KE DS+IVDAD + L+G +P
Sbjct: 526 KRVIQQLVENEIAMGVLRGDFKEEDSIIVDAD-----VTLSGKERSP 651
>BI945508 homologue to PIR|T51523|T51 clpB heat shock protein-like -
Arabidopsis thaliana, partial (14%)
Length = 432
Score = 199 bits (505), Expect = 6e-51
Identities = 97/141 (68%), Positives = 119/141 (83%), Gaps = 1/141 (0%)
Frame = +1
Query: 343 LVAGTKYRGEFEERLKKLMEEIKQSD-EIILFIDEVHTLIGAGAAEGAIDAANILKPALA 401
L+AG KYRGEFE+RLK +++E+ +SD + ILFIDE+HT++GAGA GA+DA N+LKP L
Sbjct: 10 LIAGAKYRGEFEDRLKAVLKEVTESDGQTILFIDEIHTVVGAGATNGAMDAGNLLKPMLG 189
Query: 402 RGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLR 461
RGEL+CIGATTLDEYRK+IEKDPALERRFQ V V +PTV +TI IL+GLRERYE+HH +R
Sbjct: 190 RGELRCIGATTLDEYRKYIEKDPALERRFQQVYVDQPTVEDTISILRGLRERYELHHGVR 369
Query: 462 YTDEALVAAAELSHQYISDRF 482
+D ALV AA LS +YIS RF
Sbjct: 370 ISDSALVEAAILSDRYISGRF 432
>TC224352 homologue to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-like,
partial (17%)
Length = 505
Score = 195 bits (495), Expect = 9e-50
Identities = 94/155 (60%), Positives = 123/155 (78%)
Frame = +1
Query: 369 EIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALER 428
+ ILFIDE+HT++GAGA+ GA+DA N+LKP L RGEL+CIGATTLDEYRK+IEKDPALER
Sbjct: 4 QTILFIDEIHTVVGAGASNGAMDAGNLLKPMLGRGELRCIGATTLDEYRKYIEKDPALER 183
Query: 429 RFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAI 488
RFQ V V +P+V +TI IL+GLRERYE+HH +R +D ALV AA LS +YIS RFLPDKAI
Sbjct: 184 RFQQVYVDQPSVEDTISILRGLRERYELHHGVRISDSALVDAAILSDRYISGRFLPDKAI 363
Query: 489 DLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEK 523
DL+DEA ++++++ P + + V ++ E+
Sbjct: 364 DLVDEAAAKLKMEITSKPTALDEINRSVLKLEMER 468
>TC228467 weakly similar to UP|ERD1_ARATH (P42762) ERD1 protein, chloroplast
precursor, partial (17%)
Length = 799
Score = 164 bits (415), Expect = 2e-40
Identities = 84/164 (51%), Positives = 120/164 (72%), Gaps = 1/164 (0%)
Frame = +1
Query: 749 VDFKNTLLIMTSNVGSSVIEKGGRR-IGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFL 807
V FKN L++MTSNVGSS I KG IGF L D+K +SYN +KS+V EEL+ YFRPE L
Sbjct: 1 VSFKNALVVMTSNVGSSAITKGRHNSIGF-LIPDDKKTSYNGLKSMVIEELRTYFRPELL 177
Query: 808 NRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGA 867
NR+DE++VF+ L K ++ +I D++L+++ +R+ + I + V+E + V +GYNP+YGA
Sbjct: 178 NRIDEVVVFQPLEKSQLLQILDVLLQDMKKRVLSLGIHVKVSEAVKNLVCQQGYNPTYGA 357
Query: 868 RPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVLN 911
RPLRRAI L+ED ++E +L E K+GD+V+VD D++GN V N
Sbjct: 358 RPLRRAITSLIEDPLSEALLYGECKQGDTVLVDLDANGNPFVTN 489
>AW568757 similar to SP|P35100|CLPA_ ATP-dependent clp protease ATP-binding
subunit clpA homolog chloroplast precursor. [Garden
pea], partial (9%)
Length = 271
Score = 152 bits (384), Expect = 7e-37
Identities = 78/90 (86%), Positives = 83/90 (91%)
Frame = +2
Query: 835 VFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEG 894
VF+RLK KEI+LSVTERFRERVVDEGYNPSYGARPL RAIM+LLEDSMAEKMLAREI EG
Sbjct: 2 VFQRLKAKEIDLSVTERFRERVVDEGYNPSYGARPLTRAIMQLLEDSMAEKMLAREI*EG 181
Query: 895 DSVIVDADSDGNVIVLNGSTGAPDSLPDAL 924
DSVIVD+DS+GNVIVLNG GAPDSL D L
Sbjct: 182 DSVIVDSDSEGNVIVLNGMFGAPDSLEDVL 271
>AW200876 weakly similar to GP|4105131|gb| ClpC protease {Spinacia oleracea},
partial (12%)
Length = 487
Score = 146 bits (369), Expect = 4e-35
Identities = 87/162 (53%), Positives = 105/162 (64%)
Frame = +2
Query: 654 SELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPY 713
+E + L++ E +I LDM M+ H+ +LI +P GY GG L EAVRR Y
Sbjct: 2 TEKDRCLSANLLPYVEGIILLDMCGDMDGHSGLQLIDTPTGYSASYVGG*LLEAVRRHTY 181
Query: 714 TVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRR 773
TV +I+ AHP VFNM LQILE+GRL+DS + DF NTLLIMTSN G V EK GR
Sbjct: 182 TV*RICDIDNAHPAVFNMKLQILEEGRLSDSLR*SEDFTNTLLIMTSNAGGRVHEKRGRI 361
Query: 774 IGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIV 815
I FDLDYDEKDS RI S VT+ELK + PE LNR ++MIV
Sbjct: 362 I*FDLDYDEKDSMLYRIMS*VTDELKPHCMPESLNRENDMIV 487
>TC211001 homologue to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-like,
partial (16%)
Length = 456
Score = 140 bits (352), Expect = 3e-33
Identities = 66/101 (65%), Positives = 81/101 (79%)
Frame = +2
Query: 255 KMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIA 314
K LE+YG +LT +A+ GKLDPV+GR +I R QIL RRTKNNP LIGEPGVGKTAI+
Sbjct: 152 KYEALEKYGKDLTAMAKAGKLDPVIGRDDEIRRCIQILSRRTKNNPVLIGEPGVGKTAIS 331
Query: 315 EGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEE 355
EGLAQRI GDVP+ + +++I+LDMG L+AG KYRGEFE+
Sbjct: 332 EGLAQRIVQGDVPQALMNRRLISLDMGALIAGAKYRGEFED 454
>TC211465 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (15%)
Length = 593
Score = 93.2 bits (230), Expect = 5e-19
Identities = 61/143 (42%), Positives = 81/143 (55%), Gaps = 1/143 (0%)
Frame = +2
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD 60
M+R LAQS+NVPGLVA RH KG+ + +RS +MM RT R+S +SGLRT N LD
Sbjct: 176 MARVLAQSVNVPGLVAEHRH-GQQKGSGKLKRSTKMMSALRTNGLRMSGFSGLRTFNPLD 352
Query: 61 SMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFT-EKAIKVIMLAQEEARRLGH 119
+MLRPG DFHSKV + +A R +RCV ++ F E + + + L
Sbjct: 353 TMLRPGIDFHSKVSIATSSRQA---RATRCVAQSHV*AFH*ESNQS*LC*PRRKQDVLVT 523
Query: 120 NFVGTEQILLGLIGEGTGIAAKV 142
+ + L +GTGIAAKV
Sbjct: 524 ILLAQNKFYWVLSAKGTGIAAKV 592
Score = 40.8 bits (94), Expect = 0.003
Identities = 16/33 (48%), Positives = 27/33 (81%)
Frame = +1
Query: 177 RAKRVLELSLEEARQLGHNYIGSEHLLLGLLRE 209
+ +++ L+ EEAR+LGHN++G+E +LLGL+ E
Sbjct: 469 KQSKLIMLAQEEARRLGHNFVGTEQILLGLIGE 567
>BI941950 similar to PIR|T51523|T51 clpB heat shock protein-like -
Arabidopsis thaliana, partial (13%)
Length = 421
Score = 81.6 bits (200), Expect = 1e-15
Identities = 35/68 (51%), Positives = 53/68 (77%)
Frame = +2
Query: 577 DIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNP 636
DI IV+ WTGIP+ K+ + ++LL +E+ LHKR++GQ AV+A++ AI+R+R GL +P
Sbjct: 215 DIADIVSKWTGIPISKLQQSDREKLLYLEEELHKRVVGQDPAVKAVAEAIQRSRAGLSDP 394
Query: 637 NRPIASFI 644
+RPIASF+
Sbjct: 395 HRPIASFM 418
>TC204089
Length = 1141
Score = 70.1 bits (170), Expect = 4e-12
Identities = 43/128 (33%), Positives = 71/128 (54%), Gaps = 3/128 (2%)
Frame = +2
Query: 98 RFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEV 157
+++E+AIK + + EAR+L + GTE +L+G++ EGT AAK ++ GI L R E
Sbjct: 362 KWSERAIKSYAMGELEARKLKYPNTGTEALLMGILVEGTSKAAKFSRANGITLLKVREET 541
Query: 158 EKIIGRGS--GFVAVEIPFTPRAKRVLELSLEEARQLGH-NYIGSEHLLLGLLREGEGVA 214
++G+ F P T A++ L+ ++EE + G I + HLLLG+ + E
Sbjct: 542 VGLLGKSDLFFFSPEHPPLTEPAQKALDWAIEEKLKSGEGGEINATHLLLGIWSQKESAG 721
Query: 215 ARVLENLG 222
++L LG
Sbjct: 722 QQILATLG 745
>BM187883 homologue to GP|9651530|gb| ClpB {Phaseolus lunatus}, partial (14%)
Length = 427
Score = 70.1 bits (170), Expect = 4e-12
Identities = 37/105 (35%), Positives = 71/105 (67%)
Frame = +3
Query: 457 HHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEV 516
HH ++ +D ALV+AA L+ +YI++RFLPDKAIDL+DEA ++++++ P E +++ +
Sbjct: 3 HHGVKISDSALVSAAVLADRYITERFLPDKAIDLVDEAAAKLKMEITSKPTELDEIDRAI 182
Query: 517 RQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNK 561
++ EK +++N + +KA +++ L+ +S L +K KE+ +
Sbjct: 183 LKLEMEK-LSLKN-DTDKAS--KERLSKLENDLSLLKQKQKELTE 305
>TC225571 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (4%)
Length = 654
Score = 67.8 bits (164), Expect = 2e-11
Identities = 33/38 (86%), Positives = 36/38 (93%)
Frame = +3
Query: 887 LAREIKEGDSVIVDADSDGNVIVLNGSTGAPDSLPDAL 924
LAREIKEGDSVIVD+DS+GNVIVLNGS+GAPDSL D L
Sbjct: 3 LAREIKEGDSVIVDSDSEGNVIVLNGSSGAPDSLEDVL 116
>CD403507
Length = 633
Score = 65.5 bits (158), Expect = 1e-10
Identities = 43/120 (35%), Positives = 66/120 (54%), Gaps = 4/120 (3%)
Frame = -2
Query: 109 LAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGS--G 166
+ + EAR+L + GTE IL+G++ EGT AAK L++ GI L AR E +++G+
Sbjct: 632 MGELEARKLKYPNTGTEAILMGILVEGTSNAAKFLRANGITLLKAREETVELLGKSDLFF 453
Query: 167 FVAVEIPFTPRAKRVLELSLEE-ARQLGH-NYIGSEHLLLGLLREGEGVAARVLENLGAD 224
F P T A++ L+ ++EE + G I HLLLG+ + E ++L LG D
Sbjct: 452 FSPEHPPLTEPAQKALDWAIEEKLKSAGEGGEINVTHLLLGIWSQEESAGQQILATLGFD 273
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.136 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,281,213
Number of Sequences: 63676
Number of extensions: 293943
Number of successful extensions: 1595
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 1567
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1582
length of query: 926
length of database: 12,639,632
effective HSP length: 106
effective length of query: 820
effective length of database: 5,889,976
effective search space: 4829780320
effective search space used: 4829780320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC126794.6


