

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
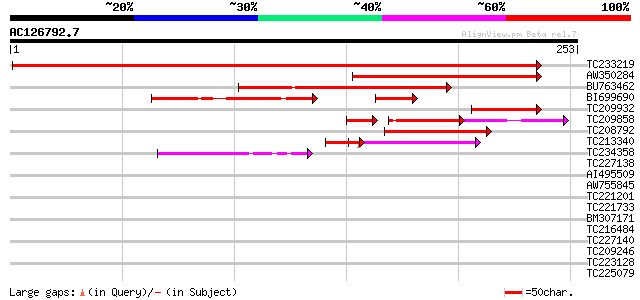
Query= AC126792.7 - phase: 0
(253 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC233219 weakly similar to UP|Q8LKI9 (Q8LKI9) NBS/LRR resistance... 369 e-102
AW350284 similar to GP|22946224|gb| CG31868-PA {Drosophila melan... 150 7e-37
BU763462 weakly similar to GP|9757923|dbj| laccase (diphenol oxi... 86 2e-17
BI699690 52 8e-10
TC209932 54 9e-08
TC209858 37 2e-07
TC208792 homologue to UP|Q9S834 (Q9S834) ATP-dependent Clp prote... 50 8e-07
TC213340 40 3e-05
TC234358 43 1e-04
TC227138 similar to GB|AAO44062.1|28466907|BT004796 At1g51160 {A... 35 0.002
AI495509 39 0.002
AW755845 35 0.043
TC221201 33 0.095
TC221733 similar to UP|Q94BV7 (Q94BV7) AT4g05020/T32N4_4, partia... 32 0.36
BM307171 29 1.8
TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabin... 28 5.2
TC227140 similar to GB|AAO44062.1|28466907|BT004796 At1g51160 {A... 28 5.2
TC209246 27 6.8
TC223128 similar to UP|WR27_ARATH (Q9FLX8) Probable WRKY transcr... 27 8.9
TC225079 similar to UP|Q8L8M0 (Q8L8M0) ARF GAP-like zinc finger-... 27 8.9
>TC233219 weakly similar to UP|Q8LKI9 (Q8LKI9) NBS/LRR resistance
protein-like protein (Fragment), partial (74%)
Length = 781
Score = 369 bits (946), Expect = e-102
Identities = 176/236 (74%), Positives = 203/236 (85%)
Frame = +3
Query: 2 IAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA 61
+AYIRAI+DMY+ STSVRTQ G ++DFPITIGLHQGSTLSPY+FTL+LDVLTE IQE+
Sbjct: 36 VAYIRAIQDMYDRVSTSVRTQGGESDDFPITIGLHQGSTLSPYLFTLILDVLTEQIQEIV 215
Query: 62 PRCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTLE 121
RCMLFADD+VL+G+SRE++N RLETWR+ALE +GF LSRSK+EYMEC F+ RR S E
Sbjct: 216 SRCMLFADDIVLLGESREKLNERLETWRRALETHGFRLSRSKSEYMECKFNKRRRVSNSE 395
Query: 122 VKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKKVPFKLKG 181
VK+GDHIIPQVTRFKYL S +Q+DGEIE DV+HRIQ W+KWR+AS VLCD KVP KLKG
Sbjct: 396 VKIGDHIIPQVTRFKYLGSVIQDDGEIEGDVNHRIQA*WMKWRKASGVLCDAKVPIKLKG 575
Query: 182 KFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTRHDRIRNDIIRER 237
KFYRTAVR A+LYGTECWAVKSQHEN+V VA RMLRWM GKTR D+IRN+ ER
Sbjct: 576 KFYRTAVRPAILYGTECWAVKSQHENKVGVAXXRMLRWMCGKTRQDKIRNEAXXER 743
>AW350284 similar to GP|22946224|gb| CG31868-PA {Drosophila melanogaster},
partial (1%)
Length = 767
Score = 150 bits (378), Expect = 7e-37
Identities = 68/84 (80%), Positives = 74/84 (87%)
Frame = -1
Query: 154 HRIQVGWLKWRRASCVLCDKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAE 213
HRIQ GW+KWR+ S VLCD KVP KLKGKFYRTAVR A+LYGTECWAVKSQHEN+V VAE
Sbjct: 590 HRIQAGWMKWRKTSGVLCDAKVPIKLKGKFYRTAVRPAILYGTECWAVKSQHENKVGVAE 411
Query: 214 MRMLRWMSGKTRHDRIRNDIIRER 237
MRMLRWM GKTR D+IRN+ IRER
Sbjct: 410 MRMLRWMCGKTRQDKIRNEAIRER 339
>BU763462 weakly similar to GP|9757923|dbj| laccase (diphenol oxidase)
{Arabidopsis thaliana}, partial (4%)
Length = 421
Score = 85.5 bits (210), Expect = 2e-17
Identities = 44/95 (46%), Positives = 62/95 (64%)
Frame = +1
Query: 103 KTEYMECNFSGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLK 162
KT+YM NFS + ++ LEVK+GD IP V++FKY E +QN I D +H++Q+G LK
Sbjct: 1 KTKYMHSNFSKK*EENELEVKMGD-AIP*VSKFKYFEPILQNSW*INEDGTHKMQLGCLK 177
Query: 163 WRRASCVLCDKKVPFKLKGKFYRTAVRSALLYGTE 197
+AS + CD KVP +K +FY T + +LYG E
Sbjct: 178 EAKASRINCDHKVPTNIKAQFYCTVIHPNILYGNE 282
>BI699690
Length = 407
Score = 51.6 bits (122), Expect(2) = 8e-10
Identities = 30/74 (40%), Positives = 46/74 (61%)
Frame = -1
Query: 64 CMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTLEVK 123
CML ADD+ + +S G + W+Q+ F LSRSKT+YM C+FS ++ + E
Sbjct: 347 CMLSADDIFFIEESHNMRFGH-KLWKQS-----FHLSRSKTKYMYCSFSKKQEEYK*E-- 192
Query: 124 VGDHIIPQVTRFKY 137
G+ +IPQV++FK+
Sbjct: 191 DGEDVIPQVSKFKF 150
Score = 28.5 bits (62), Expect(2) = 8e-10
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = -3
Query: 164 RRASCVLCDKKVPFKLKGK 182
+R V+CD KVP KLKGK
Sbjct: 57 KRQLGVICDHKVPIKLKGK 1
>TC209932
Length = 829
Score = 53.5 bits (127), Expect = 9e-08
Identities = 24/31 (77%), Positives = 27/31 (86%)
Frame = +2
Query: 207 NQVSVAEMRMLRWMSGKTRHDRIRNDIIRER 237
N+V VAEMRMLRWM GKTR D+IRN+ IRER
Sbjct: 224 NKVGVAEMRMLRWMCGKTRQDKIRNEAIRER 316
>TC209858
Length = 1340
Score = 37.0 bits (84), Expect(3) = 2e-07
Identities = 18/34 (52%), Positives = 23/34 (66%)
Frame = -3
Query: 170 LCDKKVPFKLKGKFYRTAVRSALLYGTECWAVKS 203
LC + VP K KGKFY +R A+ GT+C A+KS
Sbjct: 285 LC-QNVPLKTKGKFYHPNIRHAM*CGTKCRAIKS 187
Score = 29.3 bits (64), Expect(3) = 2e-07
Identities = 19/48 (39%), Positives = 25/48 (51%)
Frame = -1
Query: 202 KSQHENQVSVAEMRMLRWMSGKTRHDRIRNDIIRERERESGGSTYSRK 249
K+ +NQ++V EMR+L WM R D R+S STY RK
Sbjct: 191 KANKKNQLNVVEMRLLHWMYW------TR*D*KCTNWRKSWDSTYGRK 66
Score = 24.3 bits (51), Expect(3) = 2e-07
Identities = 8/14 (57%), Positives = 11/14 (78%)
Frame = -2
Query: 151 DVSHRIQVGWLKWR 164
DV+HRI G +KW+
Sbjct: 334 DVNHRIHAGRMKWK 293
>TC208792 homologue to UP|Q9S834 (Q9S834) ATP-dependent Clp protease subunit
ClpP (NClpP1) (ATP-dependent Clp protease proteolytic
subunit ClpP5) , partial (14%)
Length = 739
Score = 50.4 bits (119), Expect = 8e-07
Identities = 24/48 (50%), Positives = 32/48 (66%)
Frame = -2
Query: 168 CVLCDKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVSVAEMR 215
C +KKVP KLKGK T++RS ++YGT+ W VK Q + +VAE R
Sbjct: 609 CETNNKKVPLKLKGKLQDTSIRSMMVYGTKEWVVKGQQ*RKHNVAETR 466
>TC213340
Length = 501
Score = 39.7 bits (91), Expect(2) = 3e-05
Identities = 23/60 (38%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Frame = -1
Query: 152 VSH-RIQVGWLKWRRASCVLCDKKVPFKLKGKFYRTAVRSALLYGTECWAVKSQHENQVS 210
+SH R ++ W K V+ KV LK +FY T + +LY +ECW K HE +VS
Sbjct: 450 MSHTRCKLMWKKSIGGLFVIEICKVLTNLKREFYCTVIEPTILYSSECWGSKG*HEKKVS 271
Score = 24.6 bits (52), Expect(2) = 3e-05
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = -3
Query: 142 VQNDGEIEADVSHRIQV 158
VQN+ EI DV+H++QV
Sbjct: 478 VQNNVEINKDVTHKMQV 428
>TC234358
Length = 432
Score = 43.1 bits (100), Expect = 1e-04
Identities = 29/69 (42%), Positives = 40/69 (57%)
Frame = -2
Query: 67 FADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTLEVKVGD 126
F DDVVL+ DSRE V + + W + +CL RSKT Y+ F K L+VK+
Sbjct: 317 FLDDVVLIEDSREVVYFKHKLWIYNFDTRSYCLRRSKTVYI-LQFQ*EEYK--LKVKI-R 150
Query: 127 HIIPQVTRF 135
+I+ QV+RF
Sbjct: 149 NILSQVSRF 123
>TC227138 similar to GB|AAO44062.1|28466907|BT004796 At1g51160 {Arabidopsis
thaliana;} , partial (97%)
Length = 1259
Score = 34.7 bits (78), Expect(2) = 0.002
Identities = 18/48 (37%), Positives = 30/48 (62%)
Frame = -3
Query: 127 HIIPQVTRFKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKK 174
H+I +VTRF +L+ + EI+ +++ RIQ+ W + V+CDKK
Sbjct: 1209 HVI-RVTRFTHLQFIIYIGREIQGNINCRIQIKW----SITSVICDKK 1081
Score = 23.5 bits (49), Expect(2) = 0.002
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = -2
Query: 114 RRSKSTLEVKVGDHII 129
RR EVK+GDHI+
Sbjct: 1258 RRINLXFEVKIGDHIL 1211
>AI495509
Length = 394
Score = 38.9 bits (89), Expect = 0.002
Identities = 13/27 (48%), Positives = 23/27 (85%)
Frame = +2
Query: 188 VRSALLYGTECWAVKSQHENQVSVAEM 214
++ +L+GTECW VKSQ +N+++VA++
Sbjct: 314 IQXVMLFGTECWTVKSQQDNKLNVADI 394
>AW755845
Length = 152
Score = 34.7 bits (78), Expect = 0.043
Identities = 14/30 (46%), Positives = 20/30 (66%)
Frame = -2
Query: 132 VTRFKYLESFVQNDGEIEADVSHRIQVGWL 161
VT F+YL S +Q GE + DV+ +I GW+
Sbjct: 94 VT*FRYLGSIIQRKGETKGDVNQKIHSGWM 5
>TC221201
Length = 880
Score = 33.5 bits (75), Expect = 0.095
Identities = 18/45 (40%), Positives = 24/45 (53%)
Frame = +3
Query: 97 FCLSRSKTEYMECNFSGRRSKSTLEVKVGDHIIPQVTRFKYLESF 141
FCL SKT Y F + S LEV + + II +F+YL+ F
Sbjct: 396 FCLGWSKTLYT*YKFQKTQLISVLEVPIDESIISIAVQFRYLQYF 530
>TC221733 similar to UP|Q94BV7 (Q94BV7) AT4g05020/T32N4_4, partial (12%)
Length = 502
Score = 31.6 bits (70), Expect = 0.36
Identities = 16/33 (48%), Positives = 23/33 (69%), Gaps = 1/33 (3%)
Frame = -2
Query: 220 MSGKTRHDRIRN-DIIRERERESGGSTYSRKVG 251
++GK R++++R + RERERE GGS RK G
Sbjct: 114 ITGKERYEKVRERERERERERERGGSCRERKRG 16
>BM307171
Length = 422
Score = 29.3 bits (64), Expect = 1.8
Identities = 11/24 (45%), Positives = 18/24 (74%)
Frame = -1
Query: 132 VTRFKYLESFVQNDGEIEADVSHR 155
VT+F+YL S +Q +GE + DV+ +
Sbjct: 299 VTKFRYLGSIIQRNGETKGDVNSK 228
>TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (58%)
Length = 1213
Score = 27.7 bits (60), Expect = 5.2
Identities = 16/37 (43%), Positives = 20/37 (53%)
Frame = -3
Query: 68 ADDVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKT 104
A DVV VG+ EE R E W + L YG C S++
Sbjct: 125 AGDVVRVGEGGEEEQRRREQWPEELH-YGRCGRESES 18
>TC227140 similar to GB|AAO44062.1|28466907|BT004796 At1g51160 {Arabidopsis
thaliana;} , partial (75%)
Length = 754
Score = 27.7 bits (60), Expect = 5.2
Identities = 16/40 (40%), Positives = 25/40 (62%)
Frame = -1
Query: 135 FKYLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKK 174
FK+L+ + EI+ +++ RIQ LKW S V+C+KK
Sbjct: 703 FKHLQLIIYIGREIQGNINCRIQ---LKWSITS-VICNKK 596
>TC209246
Length = 970
Score = 27.3 bits (59), Expect = 6.8
Identities = 21/63 (33%), Positives = 26/63 (40%), Gaps = 2/63 (3%)
Frame = +3
Query: 92 LEAYGFCLSRSKTEYM--ECNFSGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIE 149
L +C+ TE + C SG RS TLE K + V R K SFV D +
Sbjct: 351 LRKINYCVPALDTEIVCYVCRESGDRSCFTLEKKKANDYSNVVERVKGTHSFVVGDETLN 530
Query: 150 ADV 152
V
Sbjct: 531 VKV 539
>TC223128 similar to UP|WR27_ARATH (Q9FLX8) Probable WRKY transcription
factor 27 (WRKY DNA-binding protein 27), partial (21%)
Length = 821
Score = 26.9 bits (58), Expect = 8.9
Identities = 25/92 (27%), Positives = 38/92 (41%)
Frame = -1
Query: 96 GFCLSRSKTEYMECNFSGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQNDGEIEADVSHR 155
G+C+ S + + + SG SKS + V +P T LES V + +
Sbjct: 404 GYCI*MSPSSFPSVSISGSGSKSAVSVD-----LPPAT----LESVVA--------MKKK 276
Query: 156 IQVGWLKWRRASCVLCDKKVPFKLKGKFYRTA 187
+ GWL RR + V +P G F R +
Sbjct: 275 KKKGWLGSRRVTQVTVADWLPESYSGYFRRAS 180
>TC225079 similar to UP|Q8L8M0 (Q8L8M0) ARF GAP-like zinc finger-containing
protein ZIGA3, partial (48%)
Length = 1845
Score = 26.9 bits (58), Expect = 8.9
Identities = 12/38 (31%), Positives = 19/38 (49%)
Frame = -2
Query: 151 DVSHRIQVGWLKWRRASCVLCDKKVPFKLKGKFYRTAV 188
++SH I+ GW R C +C VP KF + ++
Sbjct: 1442 NLSHTIETGWWILNRTPCKMCWGHVPSLTLHKFLQLSL 1329
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.135 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,088,214
Number of Sequences: 63676
Number of extensions: 163530
Number of successful extensions: 824
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 821
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 821
length of query: 253
length of database: 12,639,632
effective HSP length: 95
effective length of query: 158
effective length of database: 6,590,412
effective search space: 1041285096
effective search space used: 1041285096
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126792.7


