

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
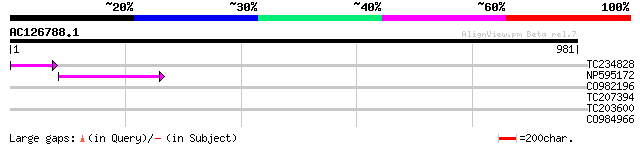
Query= AC126788.1 - phase: 0 /pseudo
(981 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC234828 57 4e-08
NP595172 polyprotein [Glycine max] 48 2e-05
CO982196 31 2.3
TC207394 similar to UP|Q6YY75 (Q6YY75) Serine/threonine-protein ... 30 5.2
TC203600 similar to UP|O64455 (O64455) Ca2+/H+ exchanger, partia... 30 6.8
CO984966 29 8.9
>TC234828
Length = 857
Score = 57.0 bits (136), Expect = 4e-08
Identities = 35/83 (42%), Positives = 43/83 (51%)
Frame = +1
Query: 1 KTKL*LAGYQW*TSFLKFFRMRFQMYHQKGRLSSPSIWCQERSRCRWHLIVCRHPNLLS* 60
K L L Y+W +F KFFRM F H + + +S WC R +CR LI C N *
Sbjct: 607 KRTLKLVVYRWCQNFQKFFRMMFVNCHLREKWNSLLTWCLGRIQCRLRLIECLRWN*QR* 786
Query: 61 RNS*RTCSTRSL*DQVFHRGERL 83
R+ RT +L Q HRGERL
Sbjct: 787 RHKYRTF*ANNLFVQAHHRGERL 855
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 48.1 bits (113), Expect = 2e-05
Identities = 57/184 (30%), Positives = 85/184 (45%)
Frame = +2
Query: 85 CW*RRKMVV*GYALTTVS*IKLQ*RIDIRFRELTT*WISWLVQRFLVKLT*GQVTIRSR* 144
C RRKM V A *++ *RI + ++ ++++ *GQ TI+
Sbjct: 1913 CLLRRKMEVGDSAQIIEL*MQSL*RIAFQCQQWMNC*MNYMELNIFQNWI*GQDTIKFWY 2092
Query: 145 KMRTCRRLPLAHDMVITNIR*CLSVLLMRLVCSWSI*TEFFMLTWINLWLYSLMTF*SIR 204
+R R+L L H M I N *C VLLM S + *T++F L NL+L LMT * I
Sbjct: 2093 SLRIERKLLLEHIMAIMNG**CHLVLLMHQPHSNA**TKYFSLLSENLFLCFLMTS*YIV 2272
Query: 205 ELRKSMQNI*GLYCKY*KRRSCMPNCQSASFG*VK*VFLAILFLVVVLQWILQKLMQYHN 264
L + + + L + * S + +C + K + A +L W +Q+ QY
Sbjct: 2273 LLGRIISSTWNLCYRP*SNISYLLDCPNVHLEIQKLITWATRYLG*EFPWRIQRYKQY*I 2452
Query: 265 GRLR 268
G L+
Sbjct: 2453 GLLQ 2464
>CO982196
Length = 812
Score = 31.2 bits (69), Expect = 2.3
Identities = 30/105 (28%), Positives = 48/105 (45%)
Frame = +2
Query: 578 CFGGLV*SVMLLNLCMHV*LARSLRLNIRSLQGC*HHWMYRSGNGIVFLWIL*QVYIILR 637
CFG * +L + +HV +RL LQGC ++ +++ G++ LWI + Y L+
Sbjct: 455 CFGK-E*RKLLGIMLLHVKSVGEIRLVHYPLQGCYNYCLFQLRCGLISLWISLEAYQKLK 631
Query: 638 EDMILYGLWSID*RSRRTLFR*ISVFRWLSWRRFIFGTL*SCMVF 682
I W I * S *+++ + + * CMVF
Sbjct: 632 GRTIY*W*W-IG*LSMLIFLH*VTLIQLRK*LSYSLKNW*DCMVF 763
>TC207394 similar to UP|Q6YY75 (Q6YY75) Serine/threonine-protein kinase
Nek4-like, partial (12%)
Length = 902
Score = 30.0 bits (66), Expect = 5.2
Identities = 14/29 (48%), Positives = 20/29 (68%)
Frame = +2
Query: 586 VMLLNLCMHV*LARSLRLNIRSLQGC*HH 614
+ML NLC + L+ S ++N +SL GC* H
Sbjct: 560 LMLWNLCSNYVLSYSSKINSKSLLGC*DH 646
>TC203600 similar to UP|O64455 (O64455) Ca2+/H+ exchanger, partial (97%)
Length = 2134
Score = 29.6 bits (65), Expect = 6.8
Identities = 15/48 (31%), Positives = 23/48 (47%)
Frame = +3
Query: 867 YLRESGQWLIGWVCPLIFRICMTCSTCRSCGSMYRILRILFLGMTCKL 914
+ R S W++ W C + + C+ C CG Y I F+G C+L
Sbjct: 975 FRRTSCDWIMEWHCLVGWNDCIHCCVV*ICGG-YN*GCIRFMGFVCEL 1115
>CO984966
Length = 852
Score = 29.3 bits (64), Expect = 8.9
Identities = 8/22 (36%), Positives = 18/22 (81%)
Frame = -2
Query: 171 LMRLVCSWSI*TEFFMLTWINL 192
++ L+C+W +*++FF++T N+
Sbjct: 602 ILSLICTWKV*SQFFLITSFNM 537
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.373 0.166 0.653
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,618,395
Number of Sequences: 63676
Number of extensions: 754390
Number of successful extensions: 10394
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 4179
Number of HSP's successfully gapped in prelim test: 364
Number of HSP's that attempted gapping in prelim test: 5971
Number of HSP's gapped (non-prelim): 5026
length of query: 981
length of database: 12,639,632
effective HSP length: 106
effective length of query: 875
effective length of database: 5,889,976
effective search space: 5153729000
effective search space used: 5153729000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (22.0 bits)
S2: 64 (29.3 bits)
Medicago: description of AC126788.1


