

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
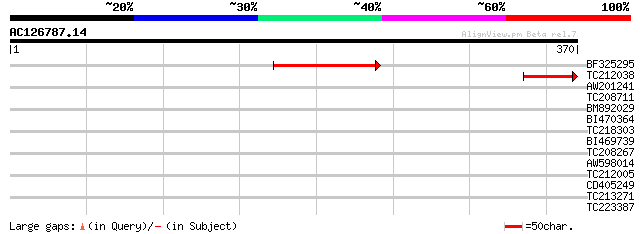
Query= AC126787.14 - phase: 0
(370 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF325295 115 3e-26
TC212038 65 5e-11
AW201241 weakly similar to GP|2194119|gb|A F20P5.5 gene product ... 33 0.16
TC208711 weakly similar to UP|O22716 (O22716) F8A5.30 protein, p... 32 0.60
BM892029 32 0.60
BI470364 weakly similar to GP|3600054|gb|A F2P3.4 gene product {... 30 1.7
TC218303 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, p... 30 2.3
BI469739 weakly similar to GP|2194119|gb| F20P5.5 gene product {... 30 2.3
TC208267 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, p... 29 3.0
AW598014 similar to GP|28190689|gb| sucrose-phosphatase {Lycoper... 29 3.9
TC212005 weakly similar to UP|Q9SEZ9 (Q9SEZ9) At2g40160/T7M7.25 ... 29 3.9
CD405249 28 6.6
TC213271 28 6.6
TC223387 28 8.6
>BF325295
Length = 219
Score = 115 bits (288), Expect = 3e-26
Identities = 51/70 (72%), Positives = 58/70 (82%)
Frame = +3
Query: 173 LFKRHSDYHIDIDEMGLKLDFIWAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHIT 232
LFKRHSDYH +DE+G+KLDF+WAPY NLT LV FK+ +YPD+LVMGSGLWHMLH T
Sbjct: 9 LFKRHSDYHTVVDEIGMKLDFMWAPYVTNLTSLVAGFKRNRVYPDLLVMGSGLWHMLHFT 188
Query: 233 NASDYGGLLG 242
NASDYG LG
Sbjct: 189 NASDYGFSLG 218
>TC212038
Length = 375
Score = 65.1 bits (157), Expect = 5e-11
Identities = 29/35 (82%), Positives = 31/35 (87%)
Frame = +3
Query: 336 GIKCTDDGMHYDGAVYEAELHIMFNALLIESHQKL 370
GIKCTDDGMHYDG VYEA + IM NALLIESHQ+L
Sbjct: 6 GIKCTDDGMHYDGVVYEAGVQIMLNALLIESHQRL 110
>AW201241 weakly similar to GP|2194119|gb|A F20P5.5 gene product {Arabidopsis
thaliana}, partial (11%)
Length = 367
Score = 33.5 bits (75), Expect = 0.16
Identities = 24/79 (30%), Positives = 33/79 (41%)
Frame = +2
Query: 120 SECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSD 179
SEC R + + L+ V VGDS AR SLL L+ + + V K
Sbjct: 11 SECHLPRFEPNTFLQLIRNKHVAFVGDSMARNQIESLLCLLATASTPKRVH---HKGSRR 181
Query: 180 YHIDIDEMGLKLDFIWAPY 198
+H D L L W+P+
Sbjct: 182 WHFDSHNASLSL--YWSPF 232
>TC208711 weakly similar to UP|O22716 (O22716) F8A5.30 protein, partial (9%)
Length = 954
Score = 31.6 bits (70), Expect = 0.60
Identities = 27/94 (28%), Positives = 36/94 (37%), Gaps = 6/94 (6%)
Frame = +2
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVR 170
W W ECE ++L+ G + VGDS R SLL L+ E V
Sbjct: 428 WRWKP----DECELPFFNATQFLNLVRGKKMAFVGDSVGRNQMQSLLCLLSHVSEPEDVS 595
Query: 171 S------FLFKRHSDYHIDIDEMGLKLDFIWAPY 198
FKR+ YH + L +W+PY
Sbjct: 596 HKYSSDVVYFKRYF-YH----DYNFTLGNLWSPY 682
>BM892029
Length = 421
Score = 31.6 bits (70), Expect = 0.60
Identities = 17/52 (32%), Positives = 24/52 (45%)
Frame = +2
Query: 194 IWAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLLR 245
IW P P + +KK DVL G+ + + HI N S G + +LR
Sbjct: 263 IWLPIPPQTVFNFFKDEKKRPQWDVLSNGNAVQEVAHIANGSHPGNCISVLR 418
>BI470364 weakly similar to GP|3600054|gb|A F2P3.4 gene product {Arabidopsis
thaliana}, partial (16%)
Length = 421
Score = 30.0 bits (66), Expect = 1.7
Identities = 33/128 (25%), Positives = 52/128 (39%), Gaps = 20/128 (15%)
Frame = +3
Query: 121 ECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSL---VLDSERV--ESVRSFLFK 175
EC R + + L++ V VGDS R SLL + V+ RV E R +L
Sbjct: 3 ECHLPRFEPNIFLQLISNKHVAFVGDSVCRNHIESLLCMLATVIKPNRVRHEGSRRWLIP 182
Query: 176 RHSDYHIDIDEMGLKLDFIWAPY-----------PN----NLTDLVMRFKKKHLYPDVLV 220
H+ L F W+P+ P+ +L + +R++K D++V
Sbjct: 183 SHNAI----------LSFYWSPFLVQGVQRQIKGPHYNTIHLDRVNIRWEKDLDEMDMIV 332
Query: 221 MGSGLWHM 228
+ G W M
Sbjct: 333 LSFGHWFM 356
>TC218303 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, partial (32%)
Length = 1230
Score = 29.6 bits (65), Expect = 2.3
Identities = 19/60 (31%), Positives = 25/60 (41%), Gaps = 1/60 (1%)
Frame = +2
Query: 192 DFIWAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLLR-NSVTS 250
+F W P +R + D+L G + M HI N D G + LLR NS S
Sbjct: 197 NFFWLPVSPKRVFEFLRDENSRSEWDILSNGGVVQEMAHIANGRDTGNCVSLLRVNSANS 376
>BI469739 weakly similar to GP|2194119|gb| F20P5.5 gene product {Arabidopsis
thaliana}, partial (25%)
Length = 424
Score = 29.6 bits (65), Expect = 2.3
Identities = 21/78 (26%), Positives = 31/78 (38%), Gaps = 9/78 (11%)
Frame = +3
Query: 110 FWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQAR---------IFTLSLLSLV 160
+W W SEC R + + L+ + VGDS AR + T+S +LV
Sbjct: 201 YWRWQP----SECNLPRFEPLTFLQLVQNKHIAFVGDSLARNQLESLLCMLSTVSSPNLV 368
Query: 161 LDSERVESVRSFLFKRHS 178
S R + F H+
Sbjct: 369 YQSANDNKFRRWHFPSHN 422
>TC208267 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, partial (31%)
Length = 1049
Score = 29.3 bits (64), Expect = 3.0
Identities = 19/57 (33%), Positives = 24/57 (41%), Gaps = 1/57 (1%)
Frame = +2
Query: 195 WAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLLR-NSVTS 250
W P P +R + D+L G + M HI N D G + LLR NS S
Sbjct: 239 WLPVPPKRVFDFLRDENSRNEWDILSNGGVVQEMAHIANGRDTGNCVSLLRVNSANS 409
>AW598014 similar to GP|28190689|gb| sucrose-phosphatase {Lycopersicon
esculentum}, partial (19%)
Length = 235
Score = 28.9 bits (63), Expect = 3.9
Identities = 17/32 (53%), Positives = 22/32 (68%), Gaps = 1/32 (3%)
Frame = +1
Query: 116 LNSVSECEFQRLKRKDVMDLLNGSW-VMIVGD 146
L +S +FQ KRK++MD LNGS +MIV D
Sbjct: 67 LTRLSFGDFQWCKRKELMDRLNGSANLMIVSD 162
>TC212005 weakly similar to UP|Q9SEZ9 (Q9SEZ9) At2g40160/T7M7.25 (At2g40160
protein), partial (26%)
Length = 436
Score = 28.9 bits (63), Expect = 3.9
Identities = 21/88 (23%), Positives = 40/88 (44%), Gaps = 1/88 (1%)
Frame = +1
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVR 170
W W +C+ R +++ L ++ VGDS R +S++ LV +S +++
Sbjct: 178 WRWKP----HQCDLPRFNATALLERLRNKRMVFVGDSLNRGQWVSMVCLV-ESSVPPTLK 342
Query: 171 SFLFKRHSDYHI-DIDEMGLKLDFIWAP 197
S + +I +E ++F WAP
Sbjct: 343 SMRTIANGSLNIFKAEEYNATIEFYWAP 426
>CD405249
Length = 695
Score = 28.1 bits (61), Expect = 6.6
Identities = 12/28 (42%), Positives = 17/28 (59%), Gaps = 1/28 (3%)
Frame = +1
Query: 274 PHLFWLGMPTL-INSMLNTQEKRVKMND 300
PH +WL PT+ I NT++K+ K D
Sbjct: 364 PHSYWLSSPTIYIYHFFNTKKKKKKTRD 447
>TC213271
Length = 503
Score = 28.1 bits (61), Expect = 6.6
Identities = 9/31 (29%), Positives = 17/31 (54%)
Frame = -2
Query: 82 LLHEIAWEVMPSKVFDPLLRNNSLNYDKFWS 112
+L W + S+ FD + NS N+ ++W+
Sbjct: 100 ILSSCLWHIFISREFDRIAERNSFNFWRYWA 8
>TC223387
Length = 569
Score = 27.7 bits (60), Expect = 8.6
Identities = 18/57 (31%), Positives = 24/57 (41%), Gaps = 1/57 (1%)
Frame = +2
Query: 195 WAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLLR-NSVTS 250
W P P +R + D+L G + + HI N D G + LLR NS S
Sbjct: 272 WLPVPPKRVFHFLRDQNSRNEWDILSNGGLVQELAHIANGRDPGNCVSLLRVNSANS 442
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,652,329
Number of Sequences: 63676
Number of extensions: 259951
Number of successful extensions: 1164
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1160
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1164
length of query: 370
length of database: 12,639,632
effective HSP length: 99
effective length of query: 271
effective length of database: 6,335,708
effective search space: 1716976868
effective search space used: 1716976868
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126787.14


