

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126786.7 - phase: 0 /pseudo
(521 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
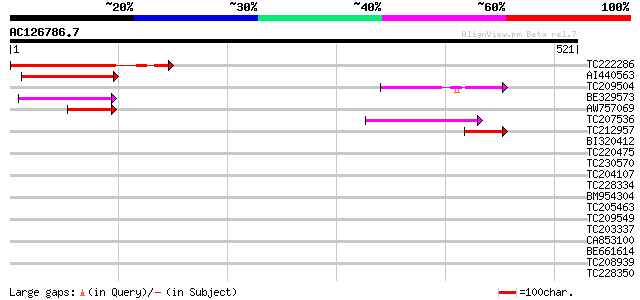
TC222286 similar to UP|ML10_ARATH (Q9FKY5) MLO-like protein 10 (... 137 9e-33
AI440563 similar to PIR|T02582|T025 H. vulgare Mlo protein homol... 97 2e-20
TC209504 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (A... 91 2e-18
BE329573 67 2e-11
AW757069 similar to GP|15982791|gb AT5g53760/MGN6_12 {Arabidopsi... 52 5e-07
TC207536 similar to UP|MLO1_ARATH (O49621) MLO-like protein 1 (A... 48 1e-05
TC212957 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (A... 42 9e-04
BI320412 similar to GP|14091574|gb| membrane protein Mlo2 {Arabi... 35 0.11
TC220475 similar to UP|ML11_ARATH (Q9FI00) MLO-like protein 11 (... 34 0.18
TC230570 similar to UP|MLO6_ARATH (Q94KB7) MLO-like protein 6 (A... 33 0.31
TC204107 similar to UP|RS10_ORYSA (Q9AYP4) 40S ribosomal protein... 31 1.2
TC228334 similar to UP|Q9SW80 (Q9SW80) Bel1-like homeodomain 2 (... 28 1.5
BM954304 30 2.0
TC205463 UP|Q39804 (Q39804) BiP isoform B, complete 30 2.6
TC209549 weakly similar to UP|Q94CH1 (Q94CH1) Seven transmembran... 30 3.4
TC203337 similar to UP|Q9S728 (Q9S728) En/Spm-like transposon pr... 30 3.4
CA853100 weakly similar to PIR|C96700|C967 protein F12A21.15 [im... 29 4.4
BE661614 29 4.4
TC208939 UP|Q39830 (Q39830) BiP isoform A, complete 29 4.4
TC228350 similar to UP|Q75IA4 (Q75IA4) Expressed protein, partia... 29 5.8
>TC222286 similar to UP|ML10_ARATH (Q9FKY5) MLO-like protein 10 (AtMlo10),
partial (25%)
Length = 731
Score = 137 bits (346), Expect = 9e-33
Identities = 73/150 (48%), Positives = 95/150 (62%)
Frame = +3
Query: 1 MGGGGGGGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEK 60
M G SR LD TPTWAVAAVC + +++SI LEK +HK + ++KK ALLEALEK
Sbjct: 141 MAAGESSSSSRDLDQTPTWAVAAVCTVFILVSIALEKSLHKVGTWLGQKKKKALLEALEK 320
Query: 61 IKAELMVLGFISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAERTADHPGEPALEPQGTE 120
+KAELM+LGFISLLLTFGQ+YI ++CIP K ++ MLPC P +
Sbjct: 321 VKAELMILGFISLLLTFGQSYIVRICIPEKLADNMLPC-----------------PYKYK 449
Query: 121 HEPTPNIPAGEGESKGEHHRRLLSYERRFL 150
+ + +S+ EH R+LLSYERR+L
Sbjct: 450 EDKKAS------DSEEEHRRKLLSYERRYL 521
>AI440563 similar to PIR|T02582|T025 H. vulgare Mlo protein homolog
[imported] - Arabidopsis thaliana, partial (15%)
Length = 376
Score = 96.7 bits (239), Expect = 2e-20
Identities = 45/89 (50%), Positives = 64/89 (71%)
Frame = +1
Query: 12 QLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGFI 71
+L+ TPTWAVA VC +++ ISIL+E ++ + +++ K AL EALEK+K ELM+LGFI
Sbjct: 109 KLEATPTWAVAVVCFVMLAISILIEHILEELGKWLKKKHKKALHEALEKVKGELMLLGFI 288
Query: 72 SLLLTFGQNYISKVCIPVKYSNTMLPCQP 100
SLLL Q++IS +CIP ++T PC P
Sbjct: 289 SLLLVMFQDHISNICIPKSVASTWHPCDP 375
>TC209504 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (AtMlo8),
partial (24%)
Length = 735
Score = 90.5 bits (223), Expect = 2e-18
Identities = 55/126 (43%), Positives = 76/126 (59%), Gaps = 9/126 (7%)
Frame = +1
Query: 341 RKTCSSAGDTSCTSL*QVFLV*MASISSLSPPLCPLSECI*ADLFLVDMV*IWVGILLL* 400
RKTC D+SC+ L Q+FLV ASISS S P C + EC+ ++FLVDMV W L
Sbjct: 58 RKTCCCPRDSSCSRLRQIFLVWSASISSSSYPFCFVPECVPNNIFLVDMVFFWAEKL--- 228
Query: 401 R*QSYDC*--SCS-------WGGCSICLQLYHTSSLCTRHTDGINDEAVNI*RANIKGIA 451
+ C* +CS WG S LQL+H S +C+ ++DG+ +E +NI*R NI+G+
Sbjct: 229 ----FPC*LQACSSESSFRAWGAMS--LQLHHPSIICSCYSDGLKNEKINI*RTNIEGVE 390
Query: 452 KLA*KC 457
++A C
Sbjct: 391 EMAHGC 408
>BE329573
Length = 400
Score = 67.0 bits (162), Expect = 2e-11
Identities = 34/91 (37%), Positives = 54/91 (58%), Gaps = 1/91 (1%)
Frame = +3
Query: 9 DSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVL 68
+ R L TPT+AVA V ++V +S L + + K + K+ +LL AL+KIK ELM+
Sbjct: 3 EGRSLAETPTYAVATVITVLVSLSFLFQGTLKKLVKWLDRTKRKSLLSALDKIKEELMLF 182
Query: 69 GFISLLLTFGQNYISKVCIPVK-YSNTMLPC 98
G +SLL+ +++K+C+ S+T PC
Sbjct: 183GLLSLLMGHWIIFVAKICVKSSVLSSTFFPC 275
>AW757069 similar to GP|15982791|gb AT5g53760/MGN6_12 {Arabidopsis
thaliana}, partial (22%)
Length = 443
Score = 52.4 bits (124), Expect = 5e-07
Identities = 27/46 (58%), Positives = 33/46 (71%), Gaps = 1/46 (2%)
Frame = +2
Query: 54 LLEALEKIKAELMVLGFISLLLTFGQNYISKVCIPVK-YSNTMLPC 98
LL ALEK+K ELM+LGFISLLLT I+ +CIP K Y++ PC
Sbjct: 14 LLAALEKMKEELMLLGFISLLLTATSRMIANICIPSKFYNSAFAPC 151
>TC207536 similar to UP|MLO1_ARATH (O49621) MLO-like protein 1 (AtMlo1) (MLO
protein homolog 1) (AtMLO-H1), partial (28%)
Length = 874
Score = 47.8 bits (112), Expect = 1e-05
Identities = 32/107 (29%), Positives = 50/107 (45%)
Frame = +2
Query: 328 ASSNYN*NGTRHTRKTCSSAGDTSCTSL*QVFLV*MASISSLSPPLCPLSECI*ADLFLV 387
A + N +R + K CS SC ++ FLV A L L P +C+* +F +
Sbjct: 35 AGARNNPTSSRSS*KACSHRR*FSCAAIR*TFLVSSAPCCPLFDSLYPFPKCL*DSIFFL 214
Query: 388 DMV*IWVGILLL*R*QSYDC*SCSWGGCSICLQLYHTSSLCTRHTDG 434
DM IW+ +L + +C WG S + L H +++C + DG
Sbjct: 215 DMGHIWI*LLYNGTSSIHCSKACYWGIYSGTM*LQHPATVCNCYADG 355
>TC212957 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (AtMlo8),
partial (15%)
Length = 780
Score = 41.6 bits (96), Expect = 9e-04
Identities = 20/39 (51%), Positives = 27/39 (68%)
Frame = +2
Query: 419 LQLYHTSSLCTRHTDGINDEAVNI*RANIKGIAKLA*KC 457
LQLYH S +C+ ++DG DE NI*RAN +G ++A C
Sbjct: 134 LQLYHPSIICSCNSDGFKDENSNI*RANKQGSEEMAHGC 250
>BI320412 similar to GP|14091574|gb| membrane protein Mlo2 {Arabidopsis
thaliana}, partial (12%)
Length = 213
Score = 34.7 bits (78), Expect = 0.11
Identities = 25/71 (35%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Frame = +3
Query: 364 ASISSLSPPLCPLSECI*ADLFLVDMV*IWVGILLL*R*QSYDC*SCSWGGC-SICLQLY 422
+S SL C +SEC +FL++ +*I +LL R C GGC + +QL
Sbjct: 3 SSPPSLFDSSCSVSECFSTRIFLLEYL*ILRKVLLP-RNNRRQCHKTCNGGCHTSSVQLC 179
Query: 423 HTSSLCTRHTD 433
+S+C+ HTD
Sbjct: 180 DIASICSSHTD 212
>TC220475 similar to UP|ML11_ARATH (Q9FI00) MLO-like protein 11 (AtMlo11),
partial (24%)
Length = 749
Score = 33.9 bits (76), Expect = 0.18
Identities = 25/64 (39%), Positives = 35/64 (54%), Gaps = 1/64 (1%)
Frame = +2
Query: 372 PLCPLSECI*ADLFLVDMV*IWVGILLL*R*QSYDC*SCSWGGC-SICLQLYHTSSLCTR 430
P L ECI*A L+ +V + + LL * S + G C ++ LQL H ++LCT
Sbjct: 8 PFHSLPECI*AGFILLVLVAVRILFLLY*ESPS-PLYKANIGVCRTVSLQLQHLATLCTC 184
Query: 431 HTDG 434
+TDG
Sbjct: 185 YTDG 196
>TC230570 similar to UP|MLO6_ARATH (Q94KB7) MLO-like protein 6 (AtMlo6),
partial (24%)
Length = 957
Score = 33.1 bits (74), Expect = 0.31
Identities = 29/87 (33%), Positives = 42/87 (47%)
Frame = +2
Query: 368 SLSPPLCPLSECI*ADLFLVDMV*IWVGILLL*R*QSYDC*SCSWGGCSICLQLYHTSSL 427
SL C LS+C A + +W IL + + G S LQL ++SSL
Sbjct: 62 SLPYQFCALSKCFSAGFLRMVCASVWD*ILFPFAHGGRGDQNHNGGPYSNPLQLRYSSSL 241
Query: 428 CTRHTDGINDEAVNI*RANIKGIAKLA 454
+ TDG N+EA +I R + +A+LA
Sbjct: 242 RSSDTDGFNNEANDIQR*SSGSVAELA 322
>TC204107 similar to UP|RS10_ORYSA (Q9AYP4) 40S ribosomal protein S10,
partial (93%)
Length = 873
Score = 31.2 bits (69), Expect = 1.2
Identities = 20/51 (39%), Positives = 23/51 (44%), Gaps = 3/51 (5%)
Frame = +1
Query: 78 GQNYISKVCIPVKYSNTML---PCQPLAERTADHPGEPALEPQGTEHEPTP 125
G+ +K KYS + P PL R DHPGE EPQG P P
Sbjct: 4 GRQVRNKAIAVWKYSQNFVSQFPLPPLPPRNHDHPGE---EPQGDLQIPLP 147
>TC228334 similar to UP|Q9SW80 (Q9SW80) Bel1-like homeodomain 2
(AT4g36870/C7A10_490), partial (9%)
Length = 910
Score = 28.5 bits (62), Expect(2) = 1.5
Identities = 14/35 (40%), Positives = 20/35 (57%)
Frame = -3
Query: 4 GGGGGDSRQLDVTPTWAVAAVCAIIVIISILLEKL 38
GGGGG + +DV V VCA + ++ +LL L
Sbjct: 185 GGGGGGAVAVDVVAADGVVGVCAWLPLMLLLLFSL 81
Score = 20.8 bits (42), Expect(2) = 1.5
Identities = 7/7 (100%), Positives = 7/7 (100%)
Frame = -2
Query: 2 GGGGGGG 8
GGGGGGG
Sbjct: 339 GGGGGGG 319
>BM954304
Length = 421
Score = 30.4 bits (67), Expect = 2.0
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = +2
Query: 72 SLLLTFGQNYISKVCIPVKYSNTMLPC 98
SLLLT + I+ +CIP T++PC
Sbjct: 5 SLLLTVSEKSIANICIPKGAGETLIPC 85
>TC205463 UP|Q39804 (Q39804) BiP isoform B, complete
Length = 2383
Score = 30.0 bits (66), Expect = 2.6
Identities = 17/53 (32%), Positives = 25/53 (47%), Gaps = 2/53 (3%)
Frame = -2
Query: 109 PGEPALEP--QGTEHEPTPNIPAGEGESKGEHHRRLLSYERRFLGGGGGGPGC 159
P +L+P GT H TP G K +HH++++ + R G G GC
Sbjct: 1137 P*PSSLDPWSSGTSHCSTPQTELG*VAQKNQHHQKVIQFPR------GPGAGC 997
>TC209549 weakly similar to UP|Q94CH1 (Q94CH1) Seven transmembrane protein
Mlo4, partial (11%)
Length = 556
Score = 29.6 bits (65), Expect = 3.4
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = +2
Query: 419 LQLYHTSSLCTRHTDGI 435
+QLY+ SSLC HTDG+
Sbjct: 125 VQLYNVSSLCYHHTDGV 175
>TC203337 similar to UP|Q9S728 (Q9S728) En/Spm-like transposon protein
(Protodermal factor 1), partial (46%)
Length = 1410
Score = 29.6 bits (65), Expect = 3.4
Identities = 20/63 (31%), Positives = 27/63 (42%), Gaps = 11/63 (17%)
Frame = +2
Query: 112 PALEPQGTEHEPTPNIPAGEGES-----------KGEHHRRLLSYERRFLGGGGGGPGCK 160
P+ P G ++PTP+ P+G G S G H +LL + GGGG
Sbjct: 281 PSTPPSGGSYDPTPSPPSGGGGSY*SNLHQCPPVGGSHVSQLLHHS----SGGGGSYNPT 448
Query: 161 PWT 163
P T
Sbjct: 449 PST 457
>CA853100 weakly similar to PIR|C96700|C967 protein F12A21.15 [imported] -
Arabidopsis thaliana, partial (10%)
Length = 588
Score = 29.3 bits (64), Expect = 4.4
Identities = 16/35 (45%), Positives = 22/35 (62%), Gaps = 3/35 (8%)
Frame = +1
Query: 96 LPCQ-PLAERTADHPGEPALEPQGTE--HEPTPNI 127
+PC+ P+ E+ +HPG P +E QGT H P NI
Sbjct: 463 VPCEMPIEEQWPNHPG-PGVEQQGTSYGHHPPLNI 564
>BE661614
Length = 581
Score = 29.3 bits (64), Expect = 4.4
Identities = 18/67 (26%), Positives = 24/67 (34%), Gaps = 4/67 (5%)
Frame = +2
Query: 100 PLAERTADHPGEPALEPQGTEHEPTPNIPAGEGESKGEHHRRLLSYERRFLGG----GGG 155
P + P PA G +P PA G R +LS + + G G
Sbjct: 83 PSLPKPPSSPPTPAASSAGAS---SPAFPANTGNGDARRRRDILSADSHAVLGPLLRGAS 253
Query: 156 GPGCKPW 162
P C+PW
Sbjct: 254 VPSCRPW 274
>TC208939 UP|Q39830 (Q39830) BiP isoform A, complete
Length = 2452
Score = 29.3 bits (64), Expect = 4.4
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = -2
Query: 118 GTEHEPTPNIPAGEGESKGEHHRRLLSYERRFLGGGGGGPGC 159
GT H TP G K +HH++++ + R G G GC
Sbjct: 1077 GTSHCSTPQTELG*VAQKNQHHQKVIQFPR------GPGAGC 970
>TC228350 similar to UP|Q75IA4 (Q75IA4) Expressed protein, partial (65%)
Length = 1410
Score = 28.9 bits (63), Expect = 5.8
Identities = 25/103 (24%), Positives = 35/103 (33%), Gaps = 11/103 (10%)
Frame = +2
Query: 98 CQPLAERTADHPGEPALEPQGTEHEPTPNIPAGEGESKGEHHRRLLSYERRFLGGGG--- 154
C P A P P L P + +P + + R+ + FLG
Sbjct: 335 CSPPASSAFTSPCPPPLTPTSSSPARSPIFASSRNIFR---RMRVTTLRSSFLGTAPPTS 505
Query: 155 --------GGPGCKPWTGYISYIFSFSSWLSSM*YSVPLQ*HL 189
G C W G S ++ S WL +S PL+ HL
Sbjct: 506 SCRLS*FLGQSSCPSWLGRFSELYEESYWL----FSTPLRVHL 622
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.354 0.158 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,876,250
Number of Sequences: 63676
Number of extensions: 417269
Number of successful extensions: 8763
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 4797
Number of HSP's successfully gapped in prelim test: 139
Number of HSP's that attempted gapping in prelim test: 1417
Number of HSP's gapped (non-prelim): 7246
length of query: 521
length of database: 12,639,632
effective HSP length: 102
effective length of query: 419
effective length of database: 6,144,680
effective search space: 2574620920
effective search space used: 2574620920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126786.7


