

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126786.5 - phase: 0
(487 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
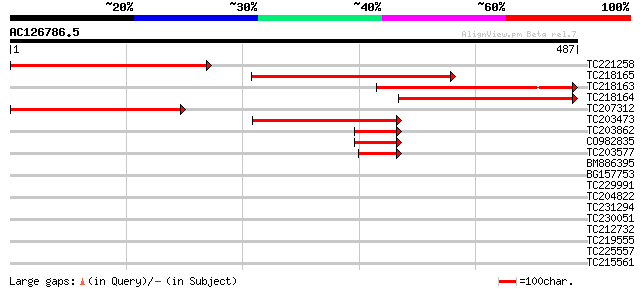
Score E
Sequences producing significant alignments: (bits) Value
TC221258 similar to PIR|T04482|T04482 ribophorin I homolog - bar... 323 8e-89
TC218165 similar to GB|AAM98094.1|22655006|AY139776 At1g76400/F1... 302 2e-82
TC218163 weakly similar to GB|AAM98094.1|22655006|AY139776 At1g7... 263 1e-70
TC218164 similar to GB|AAM98094.1|22655006|AY139776 At1g76400/F1... 234 8e-62
TC207312 weakly similar to PIR|T04482|T04482 ribophorin I homolo... 185 5e-47
TC203473 weakly similar to UP|Q9SME7 (Q9SME7) Ribophorin I (Frag... 168 6e-42
TC203862 58 1e-08
CO982835 57 2e-08
TC203577 51 1e-06
BM886395 35 0.099
BG157753 34 0.17
TC229991 33 0.22
TC204822 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, part... 33 0.37
TC231294 similar to UP|Q8W505 (Q8W505) Phosphoenolpyruvate carbo... 30 1.9
TC230051 weakly similar to UP|SYW_PROMA (Q7VBM9) Tryptophanyl-tR... 30 1.9
TC212732 UP|Q93XG4 (Q93XG4) Phytase, complete 29 4.1
TC219555 similar to UP|Q41042 (Q41042) Pisum sativum L. (clone n... 29 5.4
TC225557 similar to UP|Q9XFM6 (Q9XFM6) Progesterone-binding prot... 28 9.2
TC215561 similar to UP|Q9C5V4 (Q9C5V4) Calcium-binding protein a... 28 9.2
>TC221258 similar to PIR|T04482|T04482 ribophorin I homolog - barley
(fragment) {Hordeum vulgare;} , partial (44%)
Length = 634
Score = 323 bits (829), Expect = 8e-89
Identities = 155/173 (89%), Positives = 163/173 (93%)
Frame = +1
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 60
+D+L VFTH LQPFPEKI QADIQLLLFQESA YLSPY VKVQSL VKLP+ARIESYTKL
Sbjct: 115 LDVLAVFTHSLQPFPEKINQADIQLLLFQESAHYLSPYAVKVQSLTVKLPDARIESYTKL 294
Query: 61 ENTKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHY 120
EN KLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGN+QITEHY
Sbjct: 295 ENAKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNIQITEHY 474
Query: 121 NLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNI 173
N++H GAQSKGEFSRLDYQ RP++RGASAFRRL AKLPPRAHSVYYRDEIGNI
Sbjct: 475 NIIHAGAQSKGEFSRLDYQTRPFLRGASAFRRLVAKLPPRAHSVYYRDEIGNI 633
>TC218165 similar to GB|AAM98094.1|22655006|AY139776 At1g76400/F15M4_10
{Arabidopsis thaliana;} , partial (29%)
Length = 541
Score = 302 bits (774), Expect = 2e-82
Identities = 149/176 (84%), Positives = 159/176 (89%)
Frame = +1
Query: 208 GLPLQDFLFGVDGKRFLNISFGSPINELVIDTLVVKVVLPEGSKDISPSVPFPVKERHET 267
GLPL+DFLFG DGKRFLNISFG+PI+ELVIDTL VKVVLPEGSKDIS SVPFPVK+ ET
Sbjct: 10 GLPLRDFLFGSDGKRFLNISFGAPISELVIDTLFVKVVLPEGSKDISVSVPFPVKQSQET 189
Query: 268 KFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFNRLSMLREPLMLISGFFFLFLASIVY 327
K SHLDI GRPVVVLEKNN VPEHNEHFQVYYKFN LSMLREPLMLISGF FLF+A IVY
Sbjct: 190 KLSHLDIVGRPVVVLEKNNVVPEHNEHFQVYYKFNSLSMLREPLMLISGFLFLFVACIVY 369
Query: 328 MHADLSISKTSASYLAKIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGD 383
HAD+SISK+SASYLAK+QW+EVQATI QVH I+ RCLT HDKLEASL DLSRTGD
Sbjct: 370 THADISISKSSASYLAKLQWDEVQATIQQVHGIIGRCLTAHDKLEASLHDLSRTGD 537
>TC218163 weakly similar to GB|AAM98094.1|22655006|AY139776
At1g76400/F15M4_10 {Arabidopsis thaliana;} , partial
(28%)
Length = 733
Score = 263 bits (672), Expect = 1e-70
Identities = 132/172 (76%), Positives = 147/172 (84%)
Frame = +2
Query: 316 GFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVSRCLTTHDKLEASL 375
GF FL + Y HAD+SISK ASYLAK+QW+EVQATI QVH I+ RCLT HDKLEASL
Sbjct: 2 GFLFLXVXXXXYTHADISISKXXASYLAKLQWDEVQATIQQVHGIIGRCLTAHDKLEASL 181
Query: 376 RDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILPKVEEVIAKERDL 435
DLSRTGD Q CKATRKSVDSSLKEL+KELK P+ LQSSPQAAQILPKVEE++ KER+L
Sbjct: 182 HDLSRTGDTQACKATRKSVDSSLKELSKELKQPLAILQSSPQAAQILPKVEELVTKEREL 361
Query: 436 QEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDLMDLIDEI 487
Q+KL+ KHST+VD YEKK GREIENRIAS QQKITAL++EIDDLMDLIDEI
Sbjct: 362 QDKLLVKHSTIVDAYEKK-AGREIENRIASQQQKITALRREIDDLMDLIDEI 514
>TC218164 similar to GB|AAM98094.1|22655006|AY139776 At1g76400/F15M4_10
{Arabidopsis thaliana;} , partial (24%)
Length = 628
Score = 234 bits (596), Expect = 8e-62
Identities = 119/153 (77%), Positives = 133/153 (86%)
Frame = +1
Query: 335 SKTSASYLAKIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSV 394
++ SASYLAK+Q +EVQATI QVH I+SRCL HDKLE SL DLSRTGDIQ CKATRKSV
Sbjct: 4 TRPSASYLAKLQLDEVQATIQQVHGIISRCLXAHDKLEMSLHDLSRTGDIQACKATRKSV 183
Query: 395 DSSLKELTKELKSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKL 454
DS LKEL+KELK P+ LQSSPQAAQILPKVEE++ KER+LQ+KL+ KHSTVVD YEKK
Sbjct: 184 DSLLKELSKELKQPLAILQSSPQAAQILPKVEELVTKERELQDKLLVKHSTVVDGYEKKS 363
Query: 455 GGREIENRIASHQQKITALKQEIDDLMDLIDEI 487
GREIENRIAS Q KITAL++EIDDLMDLIDEI
Sbjct: 364 AGREIENRIASQQLKITALRREIDDLMDLIDEI 462
>TC207312 weakly similar to PIR|T04482|T04482 ribophorin I homolog - barley
(fragment) {Hordeum vulgare;} , partial (26%)
Length = 868
Score = 185 bits (469), Expect = 5e-47
Identities = 82/151 (54%), Positives = 116/151 (76%)
Frame = +2
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 60
+++L + TH L+PFP +I+Q++ QL+ F++SA LSPY VK Q+ +K P R+ES+T +
Sbjct: 416 LEVLYILTHSLEPFPVEISQSESQLVYFRDSAILLSPYHVKQQTTFIKTPSTRVESFTVM 595
Query: 61 ENTKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHY 120
+ TK G+ELKYGPYEN PP+SY P++IHFENN FAV +EL REIEISHWG++Q+TE Y
Sbjct: 596 DPTKRAGTELKYGPYENHPPYSYSPVLIHFENNNAFAVVEELEREIEISHWGSIQVTEQY 775
Query: 121 NLVHGGAQSKGEFSRLDYQARPYVRGASAFR 151
+L+H GAQ K FS+++YQ +P G S+F+
Sbjct: 776 SLIHAGAQHKSVFSKVEYQTKPXGTGVSSFK 868
>TC203473 weakly similar to UP|Q9SME7 (Q9SME7) Ribophorin I (Fragment),
partial (43%)
Length = 1052
Score = 168 bits (425), Expect = 6e-42
Identities = 82/129 (63%), Positives = 99/129 (76%), Gaps = 1/129 (0%)
Frame = +3
Query: 209 LPLQDFLF-GVDGKRFLNISFGSPINELVIDTLVVKVVLPEGSKDISPSVPFPVKERHET 267
LPLQDFLF DG+R+LN +FG P+ E V+D L+VKVVLPEGSKD + +PF VK+ E
Sbjct: 315 LPLQDFLFESPDGRRYLNFTFGCPLVETVVDKLIVKVVLPEGSKDPTVEIPFEVKQHLEI 494
Query: 268 KFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFNRLSMLREPLMLISGFFFLFLASIVY 327
K+S+LD+ GR VVVLEK NAVPEHN FQVYY FN + ML EPLML+S FF F+AS+ Y
Sbjct: 495 KYSYLDVVGRTVVVLEKRNAVPEHNAPFQVYYSFNPIFMLAEPLMLVSAFFLFFVASVAY 674
Query: 328 MHADLSISK 336
+H DLSI K
Sbjct: 675 LHIDLSIRK 701
>TC203862
Length = 1094
Score = 57.8 bits (138), Expect = 1e-08
Identities = 25/40 (62%), Positives = 31/40 (77%)
Frame = +2
Query: 297 VYYKFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISK 336
VYY+FN + ML EPLML+S FF F+AS+ Y+H DLSI K
Sbjct: 569 VYYRFNPIFMLAEPLMLVSAFFLFFVASVAYLHIDLSIRK 688
>CO982835
Length = 869
Score = 57.0 bits (136), Expect = 2e-08
Identities = 25/40 (62%), Positives = 30/40 (74%)
Frame = -1
Query: 297 VYYKFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISK 336
VYY FN + ML EPLML+S FF F+AS+ Y+H DLSI K
Sbjct: 521 VYYSFNPIFMLAEPLMLVSAFFLFFVASVAYLHIDLSIRK 402
>TC203577
Length = 442
Score = 50.8 bits (120), Expect = 1e-06
Identities = 22/37 (59%), Positives = 28/37 (75%)
Frame = +2
Query: 300 KFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISK 336
+FN + ML EPLML+S FF F+AS+ Y+H DLSI K
Sbjct: 2 RFNPIFMLAEPLMLVSAFFLFFVASVAYLHIDLSIRK 112
>BM886395
Length = 426
Score = 34.7 bits (78), Expect = 0.099
Identities = 16/22 (72%), Positives = 17/22 (76%)
Frame = +3
Query: 281 VLEKNNAVPEHNEHFQVYYKFN 302
VLEK NAVPEHN FQV+ K N
Sbjct: 3 VLEKRNAVPEHNAPFQVFTKPN 68
>BG157753
Length = 432
Score = 33.9 bits (76), Expect = 0.17
Identities = 15/19 (78%), Positives = 18/19 (93%)
Frame = +1
Query: 469 KITALKQEIDDLMDLIDEI 487
K+ AL++EIDDLMDLIDEI
Sbjct: 73 KLLALRREIDDLMDLIDEI 129
>TC229991
Length = 1011
Score = 33.5 bits (75), Expect = 0.22
Identities = 19/82 (23%), Positives = 39/82 (47%)
Frame = +3
Query: 377 DLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILPKVEEVIAKERDLQ 436
DL +G+++ D +++E + K + Q SPQ + V E++ E+D+
Sbjct: 204 DLYESGEVEKLVDAFLEGDFNIEEAIRFCKIGLLCTQDSPQLRPSMSSVLEMLLGEKDVN 383
Query: 437 EKLMAKHSTVVDCYEKKLGGRE 458
E+ + K + + E K G++
Sbjct: 384 EENVTKPGMIFEFVEAKSAGKQ 449
>TC204822 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (67%)
Length = 1232
Score = 32.7 bits (73), Expect = 0.37
Identities = 18/66 (27%), Positives = 30/66 (45%), Gaps = 8/66 (12%)
Frame = -3
Query: 254 SPSVPFPVKERHETKFSHLDIAGRP--------VVVLEKNNAVPEHNEHFQVYYKFNRLS 305
SP +PF RH+ + H V+ L + +PEH++H++VY KF
Sbjct: 840 SP*LPFRPNSRHQEQQRHHSFLSHQTL*QSQSHVIDLTPCHRLPEHSQHYRVY*KFASAG 661
Query: 306 MLREPL 311
+ P+
Sbjct: 660 YCQHPM 643
>TC231294 similar to UP|Q8W505 (Q8W505) Phosphoenolpyruvate carboxykinase,
partial (43%)
Length = 1123
Score = 30.4 bits (67), Expect = 1.9
Identities = 16/52 (30%), Positives = 27/52 (51%)
Frame = +1
Query: 353 TIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKE 404
TIH++HS+ + T L A+ + D + K +S+ +SL LT+E
Sbjct: 211 TIHELHSLQKKKSTPSTPLSATQGPFATVSDEERQKQQLQSISASLASLTRE 366
>TC230051 weakly similar to UP|SYW_PROMA (Q7VBM9) Tryptophanyl-tRNA
synthetase (Tryptophan--tRNA ligase) (TrpRS) , partial
(33%)
Length = 823
Score = 30.4 bits (67), Expect = 1.9
Identities = 16/59 (27%), Positives = 25/59 (42%)
Frame = -3
Query: 87 VIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVHGGAQSKGEFSRLDYQARPYVR 145
+IH P ++V E+ + W N + HY+ +HGG SR + P R
Sbjct: 272 IIHILKGNPILDGAKIVSEVN*TRWKN---SRHYSFLHGGRSGSL*RSRSESSTTPDAR 105
>TC212732 UP|Q93XG4 (Q93XG4) Phytase, complete
Length = 1644
Score = 29.3 bits (64), Expect = 4.1
Identities = 27/119 (22%), Positives = 48/119 (39%), Gaps = 12/119 (10%)
Frame = +1
Query: 235 LVIDTLVVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGR---------PVVVLEKN 285
L+ D + L G+ S FP+ HET D GR P++V+E N
Sbjct: 676 LIGDVTYANLYLTNGTGSDCYSCSFPLTPIHETYQPRWDYWGRFMQNLVSNVPIMVVEGN 855
Query: 286 NAVPEHNEHFQVYYKFNRLSMLREPLMLISGFFFLFLAS---IVYMHADLSISKTSASY 341
+ + + E+ +R + + S F++ F A + + A ++ KT+ Y
Sbjct: 856 HEIEKQAENRTFVAYSSRFAFPSQESGSSSTFYYSFNAGGIHFIMLGAYINYDKTAEQY 1032
>TC219555 similar to UP|Q41042 (Q41042) Pisum sativum L. (clone na-481-5),
partial (16%)
Length = 647
Score = 28.9 bits (63), Expect = 5.4
Identities = 17/47 (36%), Positives = 28/47 (59%), Gaps = 3/47 (6%)
Frame = -3
Query: 56 SYTKLENTKLQGSELKYGPYENL---PPFSYLPIVIHFENNQPFAVA 99
S+ K++ LQ S+L+ +L PPFS+ ++ F + QPF+VA
Sbjct: 639 SFLKVQ*LLLQVSQLQILKKNHLNYFPPFSFAQLLHSFVSQQPFSVA 499
>TC225557 similar to UP|Q9XFM6 (Q9XFM6) Progesterone-binding protein-like,
partial (67%)
Length = 965
Score = 28.1 bits (61), Expect = 9.2
Identities = 12/32 (37%), Positives = 19/32 (58%)
Frame = -3
Query: 327 YMHADLSISKTSASYLAKIQWEEVQATIHQVH 358
Y+ DLS+ KTS L ++++ +HQVH
Sbjct: 492 YLQLDLSLQKTSLLKLC*HPCQQIEHRVHQVH 397
>TC215561 similar to UP|Q9C5V4 (Q9C5V4) Calcium-binding protein annexin 5,
partial (12%)
Length = 592
Score = 28.1 bits (61), Expect = 9.2
Identities = 16/58 (27%), Positives = 32/58 (54%)
Frame = -1
Query: 324 SIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRT 381
SI++ + L +SKT+ ++L K++W+ V S+ S L+ D L+ ++ + T
Sbjct: 541 SILHRQSSLILSKTNINHLCKVEWKIVA-------SLTSI*LSMFDLLKQEMKIVKHT 389
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,404,251
Number of Sequences: 63676
Number of extensions: 261307
Number of successful extensions: 1329
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1323
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1327
length of query: 487
length of database: 12,639,632
effective HSP length: 101
effective length of query: 386
effective length of database: 6,208,356
effective search space: 2396425416
effective search space used: 2396425416
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126786.5


