

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126784.6 - phase: 0
(135 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
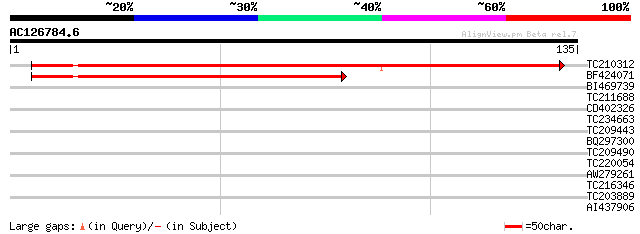
Score E
Sequences producing significant alignments: (bits) Value
TC210312 187 1e-48
BF424071 119 6e-28
BI469739 weakly similar to GP|2194119|gb| F20P5.5 gene product {... 28 1.7
TC211688 27 2.2
CD402326 similar to PIR|T17815|T178 proline-rich protein A316R -... 27 2.2
TC234663 27 2.2
TC209443 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (64%) 27 3.8
BQ297300 homologue to SP|O49856|FTRC Ferredoxin-thioredoxin redu... 26 5.0
TC209490 UP|Q8GSN9 (Q8GSN9) LRR receptor-like kinase (Fragment),... 26 6.5
TC220054 homologue to UP|Q9FVD4 (Q9FVD4) Ser/Thr specific protei... 25 8.5
AW279261 similar to GP|1181599|dbj subunit of photosystem I {Cuc... 25 8.5
TC216346 similar to UP|Q7XYY0 (Q7XYY0) AKIN gamma, partial (95%) 25 8.5
TC203889 homologue to GB|BAA84771.1|6006283|AB015861 photosystem... 25 8.5
AI437906 25 8.5
>TC210312
Length = 662
Score = 187 bits (476), Expect = 1e-48
Identities = 88/130 (67%), Positives = 104/130 (79%), Gaps = 3/130 (2%)
Frame = +2
Query: 6 EAKIDLKRDGIEDLSGRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGR 65
+A IDLKRDG +DLSGRVH LPCC+KHDGP VS YFKPK GV ++GLPLQ++HFRGR
Sbjct: 53 KATIDLKRDG-KDLSGRVHQLPCCVKHDGPASVSHYFKPKHKGVGEDEGLPLQEAHFRGR 229
Query: 66 LLEGTTLPLPHNYSGFVLGKKN---SVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRA 122
LL+GTTLPLPH YSGFVL K + S E+S+SW+T TF DITYWNHD PS ND+ RA
Sbjct: 230 LLQGTTLPLPHGYSGFVLSKSSPPTSDENSHSWDTKATFQDITYWNHDYSPSQNDELFRA 409
Query: 123 FHWIPVAEAV 132
FHW+ VA+A+
Sbjct: 410 FHWLTVAKAL 439
>BF424071
Length = 376
Score = 119 bits (297), Expect = 6e-28
Identities = 55/75 (73%), Positives = 64/75 (85%)
Frame = +3
Query: 6 EAKIDLKRDGIEDLSGRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGEDGLPLQQSHFRGR 65
+A IDLKRDG +DLSGRVH LPCC+KHDGP VS YFKPK GV ++GLPL+++HFRGR
Sbjct: 72 KATIDLKRDG-KDLSGRVHQLPCCVKHDGPAFVSHYFKPKHKGVGEDEGLPLREAHFRGR 248
Query: 66 LLEGTTLPLPHNYSG 80
LL+GTTLPLPH YSG
Sbjct: 249 LLQGTTLPLPHGYSG 293
>BI469739 weakly similar to GP|2194119|gb| F20P5.5 gene product {Arabidopsis
thaliana}, partial (25%)
Length = 424
Score = 27.7 bits (60), Expect = 1.7
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = -2
Query: 100 TFNDITYWNHDSVPSNNDDFSRAFHWIPVAEAV 132
T D T+W SN D S+ FHWI ++ ++
Sbjct: 240 TLEDCTHW---VAISNIDSLSQTFHWIRISHSL 151
>TC211688
Length = 1020
Score = 27.3 bits (59), Expect = 2.2
Identities = 14/36 (38%), Positives = 24/36 (65%)
Frame = -3
Query: 85 KKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDFS 120
K NSV S++ + T+ +F++ ++NH S SN + FS
Sbjct: 685 KSNSVPSTSIFITTKSFSETVFFNH-SAKSNPEIFS 581
>CD402326 similar to PIR|T17815|T178 proline-rich protein A316R - Chlorella
virus PBCV-1, partial (12%)
Length = 658
Score = 27.3 bits (59), Expect = 2.2
Identities = 20/50 (40%), Positives = 25/50 (50%), Gaps = 7/50 (14%)
Frame = +2
Query: 72 LPLPHNYSGFVLGKKNSVESSNSWETSVTFND-------ITYWNHDSVPS 114
L L N+SGFVLG ++SS E FND + +N DSV S
Sbjct: 317 LELSENFSGFVLGL--FIDSSEEIEDVSLFNDSFSSNFTVFNFNFDSVTS 460
>TC234663
Length = 773
Score = 27.3 bits (59), Expect = 2.2
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = +3
Query: 96 ETSVTFNDITYWNHDSVPSNNDDFSR 121
ET VT + +NH S+PSNN SR
Sbjct: 9 ETVVTRECVILFNH*SIPSNNSGSSR 86
>TC209443 similar to UP|Q6NLX7 (Q6NLX7) At4g36660, partial (64%)
Length = 1078
Score = 26.6 bits (57), Expect = 3.8
Identities = 11/30 (36%), Positives = 14/30 (46%)
Frame = +3
Query: 17 EDLSGRVHLLPCCIKHDGPTEVSQYFKPKP 46
E+ + H LPCC H GP +Q P
Sbjct: 204 EEATNSGH*LPCCSWHSGPCSPAQSLSAGP 293
>BQ297300 homologue to SP|O49856|FTRC Ferredoxin-thioredoxin reductase
catalytic chain chloroplast precursor (EC 1.18.-.-)
(FTR-C)., partial (88%)
Length = 426
Score = 26.2 bits (56), Expect = 5.0
Identities = 8/13 (61%), Positives = 11/13 (84%)
Frame = -3
Query: 75 PHNYSGFVLGKKN 87
PHN++GF+LG N
Sbjct: 160 PHNFNGFILGSNN 122
>TC209490 UP|Q8GSN9 (Q8GSN9) LRR receptor-like kinase (Fragment), complete
Length = 3298
Score = 25.8 bits (55), Expect = 6.5
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = -1
Query: 82 VLGKKNSVESSNSWETSVTFNDITYW 107
VLG S S+NSWE+S I+YW
Sbjct: 763 VLGTSASSTSTNSWESS----PISYW 698
>TC220054 homologue to UP|Q9FVD4 (Q9FVD4) Ser/Thr specific protein
phosphatase 2A B regulatory subunit beta isoform,
partial (25%)
Length = 559
Score = 25.4 bits (54), Expect = 8.5
Identities = 12/41 (29%), Positives = 20/41 (48%)
Frame = -1
Query: 67 LEGTTLPLPHNYSGFVLGKKNSVESSNSWETSVTFNDITYW 107
+E LPLPH + FV K + E + ++ + I +W
Sbjct: 202 VEPPELPLPHRHRSFVT-KSTATEHATESANPMSASKIAFW 83
>AW279261 similar to GP|1181599|dbj subunit of photosystem I {Cucumis
sativus}, partial (52%)
Length = 346
Score = 25.4 bits (54), Expect = 8.5
Identities = 10/20 (50%), Positives = 13/20 (65%)
Frame = +1
Query: 73 PLPHNYSGFVLGKKNSVESS 92
PLPHN F+L +KN + S
Sbjct: 253 PLPHNL*NFILQQKNPIHYS 312
>TC216346 similar to UP|Q7XYY0 (Q7XYY0) AKIN gamma, partial (95%)
Length = 1766
Score = 25.4 bits (54), Expect = 8.5
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = -3
Query: 15 GIEDLSGRVHLLPCC 29
GIE LSG + LPCC
Sbjct: 1260 GIEHLSGLCYQLPCC 1216
>TC203889 homologue to GB|BAA84771.1|6006283|AB015861 photosystem I subunit
PSI-L {Arabidopsis thaliana;} , partial (35%)
Length = 439
Score = 25.4 bits (54), Expect = 8.5
Identities = 11/30 (36%), Positives = 18/30 (59%)
Frame = +2
Query: 65 RLLEGTTLPLPHNYSGFVLGKKNSVESSNS 94
R + GT PLPHN F+L +++ ++ S
Sbjct: 278 RAVFGTRQPLPHNLWDFILQRRSPYATAPS 367
>AI437906
Length = 413
Score = 25.4 bits (54), Expect = 8.5
Identities = 13/34 (38%), Positives = 19/34 (55%)
Frame = +2
Query: 90 ESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAF 123
ES S +T+ TF+ YWNH + N+ S +F
Sbjct: 137 ESLLSQQTATTFSSQHYWNHHHL---NNSLSLSF 229
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.136 0.429
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,256,244
Number of Sequences: 63676
Number of extensions: 105806
Number of successful extensions: 486
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 483
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 485
length of query: 135
length of database: 12,639,632
effective HSP length: 87
effective length of query: 48
effective length of database: 7,099,820
effective search space: 340791360
effective search space used: 340791360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 53 (25.0 bits)
Medicago: description of AC126784.6


