

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126782.2 - phase: 0 /pseudo
(912 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
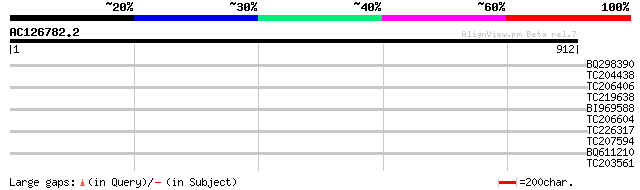
Score E
Sequences producing significant alignments: (bits) Value
BQ298390 34 0.33
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 33 0.43
TC206406 weakly similar to UP|Q9LRY5 (Q9LRY5) Gb|AAF16548.1 (AT3... 33 0.57
TC219638 33 0.57
BI969588 33 0.74
TC206604 similar to UP|Q9LSD4 (Q9LSD4) Gb|AAF23830.1, partial (37%) 31 2.8
TC226317 similar to UP|Q9LSM1 (Q9LSM1) Arabidopsis thaliana geno... 30 3.7
TC207594 similar to GB|AAF79266.1|8778257|AC023279 F12K21.21 {Ar... 30 4.8
BQ611210 similar to GP|4519673|dbj| WREBP-2 {Nicotiana tabacum},... 29 8.2
TC203561 similar to UP|Q6ZI86 (Q6ZI86) Dehydrogenase-like protei... 29 8.2
>BQ298390
Length = 424
Score = 33.9 bits (76), Expect = 0.33
Identities = 17/42 (40%), Positives = 26/42 (61%)
Frame = +1
Query: 138 IVIFKKCVKLMELFFVCLVLTHLLKMVNLNEKYGPLTILFEL 179
IV F + + L+E FF+C LTHL+ + L+E++ L F L
Sbjct: 154 IVQFLQELVLLEFFFICHQLTHLVSSLELSEQHAELVQSFVL 279
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 33.5 bits (75), Expect = 0.43
Identities = 61/223 (27%), Positives = 89/223 (39%), Gaps = 3/223 (1%)
Frame = +2
Query: 435 GT*YLDHLMLMLFGPCGFLDIKKNLTVLLRGIKPDL*VMVQVNRLALTMEKLLAQL*NQL 494
G+*+LD LM P G K VL +PDL + + T+ KL L +
Sbjct: 3293 GS*FLDPRELM*LAPSGSSRTKPMKKVL*PETRPDLLLKATLRLKV*TLMKLSPLLLDLS 3472
Query: 495 LFAQFSALLHPDLGKFISWMSRMLFFMVN*KKQCTCTNLWDLKILKTPIMYAFFANHSMG 554
+ L WM R F M * K+ ++ DL I IMY SM
Sbjct: 3473 PSDCYLV*LASSNSSCTRWM*RARF*MDT*MKKPMWSSQRDL*IQLIQIMYTGSRRLSMD 3652
Query: 555 LNKLLELGVSDLPTTCHLLVFLKANVITLFSCTRKTLIW---HIFYSRLMILFSPRPQMI 611
+KL ELG+ ++ L + +T S + K L H + LM L +M
Sbjct: 3653 *SKLQELGMKG*QSSL-LSKGIGREELTRLSLSNKMLKT***HRY--MLMTLCLEGCRMR 3823
Query: 612 CDNLSSHYSVLNSL*KIWDP*VTSWELQSLVTNKGCFSHKRSM 654
C ++ S+ LN * + + *+ W+ + SHK SM
Sbjct: 3824 CFDILSNRCNLNLR*VLLES*LIFWDSK*SRWKTPYSSHKASM 3952
>TC206406 weakly similar to UP|Q9LRY5 (Q9LRY5) Gb|AAF16548.1
(AT3g24740/K7P8_3), partial (14%)
Length = 881
Score = 33.1 bits (74), Expect = 0.57
Identities = 15/34 (44%), Positives = 23/34 (67%)
Frame = +3
Query: 146 KLMELFFVCLVLTHLLKMVNLNEKYGPLTILFEL 179
KL+ELF++C LTHL+ + L+E++ L F L
Sbjct: 483 KLLELFWICHRLTHLVSSLELSEQHAELVQSFVL 584
>TC219638
Length = 968
Score = 33.1 bits (74), Expect = 0.57
Identities = 20/90 (22%), Positives = 44/90 (48%)
Frame = +1
Query: 100 CHINPKFIQFLKN*EHTSTHNFKEKLKQFNVTTAVNMSIVIFKKCVKLMELFFVCLVLTH 159
C N F+Q H+S + ++ + ++ ++++ C ++ +L + +L H
Sbjct: 706 CRSNILFLQTCSLLSHSSQTSMQKPFQWRKLSLMETWNLLVL--CFRVFQL--IHKILVH 873
Query: 160 LLKMVNLNEKYGPLTILFELY*FTRHYHLH 189
L+KM++ ++ PL+ + T HY LH
Sbjct: 874 LIKMMDSIQEQNPLSTKL*MIIMTLHYELH 963
>BI969588
Length = 441
Score = 32.7 bits (73), Expect = 0.74
Identities = 23/59 (38%), Positives = 34/59 (56%), Gaps = 1/59 (1%)
Frame = -2
Query: 235 YVFLSFLHQKYTNFNLVP-QNVCFLRIRQIIEAINV*IYCQIKLLFVVMSYLMKILSHM 292
Y+F F+ KY FN+ P +N+C R II AI * Y QI++ V + +I+SH+
Sbjct: 290 YLFAQFIRNKYLPFNVCPKKNLC---PRPIIYAIYK*EYSQIEIF*V---SIFEIISHI 132
>TC206604 similar to UP|Q9LSD4 (Q9LSD4) Gb|AAF23830.1, partial (37%)
Length = 1091
Score = 30.8 bits (68), Expect = 2.8
Identities = 14/43 (32%), Positives = 25/43 (57%)
Frame = +3
Query: 240 FLHQKYTNFNLVPQNVCFLRIRQIIEAINV*IYCQIKLLFVVM 282
FL Q + L+PQ+VC L + Q+ E N + +++ LF+ +
Sbjct: 705 FLLQSFLL*GLIPQSVCTLALPQVAETSNFKLSLKLQCLFICL 833
>TC226317 similar to UP|Q9LSM1 (Q9LSM1) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K21L13, partial (15%)
Length = 876
Score = 30.4 bits (67), Expect = 3.7
Identities = 19/59 (32%), Positives = 28/59 (47%)
Frame = -2
Query: 525 KKQCTCTNLWDLKILKTPIMYAFFANHSMGLNKLLELGVSDLPTTCHLLVFLKANVITL 583
+ C CT LK+L +M +H +G+N LL L + LP LL K ++ L
Sbjct: 299 RSSCCCTPSQSLKLLLLLLMLLLMKSHILGMNHLLHLRM-HLPLHLKLLRQPKGTLLLL 126
>TC207594 similar to GB|AAF79266.1|8778257|AC023279 F12K21.21 {Arabidopsis
thaliana;} , partial (55%)
Length = 1137
Score = 30.0 bits (66), Expect = 4.8
Identities = 20/74 (27%), Positives = 36/74 (48%)
Frame = +1
Query: 262 QIIEAINV*IYCQIKLLFVVMSYLMKILSHMQSCTFLNLIHTLS*TMSCPLILYNI*WTK 321
Q+ + V +C LL +++S L + +H+ S + T + + C LI+Y++
Sbjct: 838 QLGRRMRVSYHCSFGLLMIMLSDLA-LHAHLHS-----IPETKANVIFCTLIIYSVPQLN 999
Query: 322 PKLAHLILNWLISQ 335
+ H ILNW Q
Sbjct: 1000ENIHHQILNWTALQ 1041
>BQ611210 similar to GP|4519673|dbj| WREBP-2 {Nicotiana tabacum}, partial
(8%)
Length = 305
Score = 29.3 bits (64), Expect = 8.2
Identities = 21/80 (26%), Positives = 35/80 (43%), Gaps = 4/80 (5%)
Frame = +3
Query: 105 KFIQFLKN*EHTSTHNFKEKLKQFNVTT----AVNMSIVIFKKCVKLMELFFVCLVLTHL 160
+ + LK K +K+ N+ T A +M + FKK + L++LFFV +
Sbjct: 48 RIVTMLKMVGDMQEETMKMMMKKQNMKTESVMASSMELTFFKKYISLLQLFFVLSLFYLA 227
Query: 161 LKMVNLNEKYGPLTILFELY 180
L+M + L L +Y
Sbjct: 228 LEMTSCQTVCWLLQALGRIY 287
>TC203561 similar to UP|Q6ZI86 (Q6ZI86) Dehydrogenase-like protein, partial
(33%)
Length = 680
Score = 29.3 bits (64), Expect = 8.2
Identities = 15/42 (35%), Positives = 26/42 (61%), Gaps = 2/42 (4%)
Frame = -3
Query: 291 HMQSCTFLNLIHT--LS*TMSCPLILYNI*WTKPKLAHLILN 330
H++ C + I T +S* +C ++ N+*+ +PKL+H I N
Sbjct: 135 HIKFCILSDKIQTH*VS*CHACSMLTKNL*YFQPKLSHSITN 10
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.346 0.153 0.515
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 49,934,305
Number of Sequences: 63676
Number of extensions: 868809
Number of successful extensions: 9983
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 7406
Number of HSP's successfully gapped in prelim test: 362
Number of HSP's that attempted gapping in prelim test: 2294
Number of HSP's gapped (non-prelim): 8184
length of query: 912
length of database: 12,639,632
effective HSP length: 106
effective length of query: 806
effective length of database: 5,889,976
effective search space: 4747320656
effective search space used: 4747320656
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC126782.2


