

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126780.2 - phase: 0 /pseudo
(297 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
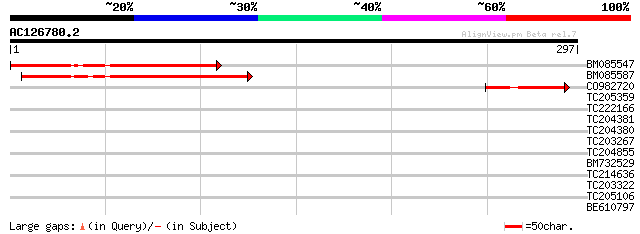
BM085547 170 6e-43
BM085587 101 4e-22
CO982720 58 6e-09
TC205359 homologue to GB|AAM64579.1|21592630|AY087018 silencing ... 37 0.008
TC222166 30 1.3
TC204381 similar to UP|CRTC_RICCO (P93508) Calreticulin precurso... 29 2.2
TC204380 homologue to UP|Q40567 (Q40567) Tobacco calretulin (Fra... 29 2.2
TC203267 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 29 2.9
TC204855 UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2, complete 28 5.0
BM732529 28 6.5
TC214636 similar to UP|Q39856 (Q39856) Epoxide hydrolase, partia... 28 6.5
TC203322 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 28 6.5
TC205106 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (33%) 27 8.5
BE610797 27 8.5
>BM085547
Length = 422
Score = 170 bits (431), Expect = 6e-43
Identities = 89/111 (80%), Positives = 100/111 (89%)
Frame = +1
Query: 1 MADTYSIESKVVIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIR 60
M DT+SIES+++IREFDEDRDVKVVGKLE+NC EI TKKG SIFTNMM GDPLSRIR
Sbjct: 103 MVDTFSIESRLLIREFDEDRDVKVVGKLEKNC-EIG--TKKGVSIFTNMM--GDPLSRIR 267
Query: 61 FYPLHVMLVAEMVESKELVGVVKGCIKSVQTPSGSLFKMGCILGLRVSPIH 111
FYPLHVMLVAE++ESKELVGVV+GCIKS++TPS SL K+GCILGLRVSP H
Sbjct: 268 FYPLHVMLVAELLESKELVGVVRGCIKSMRTPSESLLKIGCILGLRVSPTH 420
>BM085587
Length = 382
Score = 101 bits (252), Expect = 4e-22
Identities = 57/121 (47%), Positives = 80/121 (66%)
Frame = +1
Query: 7 IESKVVIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHV 66
+E V+++E++EDR V KLER C E+ + K S+ T++M GDP+ RIR + LH
Sbjct: 31 MEPLVLVKEYEEDRHKVAVEKLERLC-EVGQSGKP--SLVTDLM--GDPICRIRHFQLHA 195
Query: 67 MLVAEMVESKELVGVVKGCIKSVQTPSGSLFKMGCILGLRVSPIHRRKGVGLKLVTSIEE 126
MLVAE E E+VGV++GC+K+V + ++ ILGLRVSP HRR G+G KLV + E
Sbjct: 196 MLVAEYGEE*EVVGVIRGCVKTVTRGNSVYVELAYILGLRVSPRHRRFGIGTKLVEHL*E 375
Query: 127 W 127
W
Sbjct: 376 W 378
>CO982720
Length = 431
Score = 57.8 bits (138), Expect = 6e-09
Identities = 28/44 (63%), Positives = 32/44 (72%)
Frame = -2
Query: 250 SWIIFSLWNTCEACDNNLQVKTKLFQPLRFLHATLNHAKDKICP 293
SWIIFS+WNT EA ++ K QPLRFLH TLNHA+DKI P
Sbjct: 430 SWIIFSIWNTXEA----YRLXLKKXQPLRFLHTTLNHARDKIFP 311
>TC205359 homologue to GB|AAM64579.1|21592630|AY087018 silencing group B
protein {Arabidopsis thaliana;} , partial (92%)
Length = 830
Score = 37.4 bits (85), Expect = 0.008
Identities = 22/59 (37%), Positives = 31/59 (52%), Gaps = 1/59 (1%)
Frame = +1
Query: 100 GCILGLRVSPIHRRKGVGLKLVTSIEEWM-LTNGADYAFLATEKNNNASKNLFTNKCNY 157
G I L V HR+ G+ KL+T+ + M GA+Y L K+N A+ NL+T Y
Sbjct: 283 GHITSLAVLRTHRKLGLATKLMTAAQNAMEQVFGAEYVSLHVRKSNRAAFNLYTETLGY 459
>TC222166
Length = 791
Score = 30.0 bits (66), Expect = 1.3
Identities = 13/31 (41%), Positives = 22/31 (70%), Gaps = 3/31 (9%)
Frame = -3
Query: 30 RNCTEIN---GTTKKGFSIFTNMMSNGDPLS 57
RNC+ ++ G++ KGFS FTN +S+ D ++
Sbjct: 285 RNCSNLSCFVGSSLKGFSTFTNSLSDADSVN 193
>TC204381 similar to UP|CRTC_RICCO (P93508) Calreticulin precursor, partial
(81%)
Length = 1408
Score = 29.3 bits (64), Expect = 2.2
Identities = 18/47 (38%), Positives = 25/47 (52%)
Frame = +1
Query: 175 TNHISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSL 221
TNH+ KKDV + DQ YT IL+ Y + +D + K+ SL
Sbjct: 253 TNHLIKKDVPCE---TDQLTHVYTFILRPDATYSILIDNVEKQTGSL 384
>TC204380 homologue to UP|Q40567 (Q40567) Tobacco calretulin (Fragment),
partial (52%)
Length = 938
Score = 29.3 bits (64), Expect = 2.2
Identities = 18/47 (38%), Positives = 25/47 (52%)
Frame = +1
Query: 175 TNHISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSL 221
TNH+ KKDV + DQ YT IL+ Y + +D + K+ SL
Sbjct: 568 TNHLIKKDVPCE---TDQLTHVYTFILRPDATYSILIDNVEKQTGSL 699
>TC203267 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (61%)
Length = 1338
Score = 28.9 bits (63), Expect = 2.9
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = -2
Query: 69 VAEMVESKELVGVVKGCIKSVQTPSG 94
V+E+VE VGVV GC++ Q+P G
Sbjct: 737 VSEVVEPYLDVGVVVGCLRETQSPRG 660
>TC204855 UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2, complete
Length = 1307
Score = 28.1 bits (61), Expect = 5.0
Identities = 13/26 (50%), Positives = 17/26 (65%)
Frame = -3
Query: 69 VAEMVESKELVGVVKGCIKSVQTPSG 94
V+E+VE VGVV GC+ +Q P G
Sbjct: 711 VSEVVEPYLDVGVVVGCLGEIQNPRG 634
>BM732529
Length = 428
Score = 27.7 bits (60), Expect = 6.5
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = -3
Query: 29 ERNCTEINGTTKKGFSIFTNMMSN 52
ERN + N +K FS+F NM SN
Sbjct: 81 ERNFSSANTHSKSNFSVFCNMYSN 10
>TC214636 similar to UP|Q39856 (Q39856) Epoxide hydrolase, partial (58%)
Length = 778
Score = 27.7 bits (60), Expect = 6.5
Identities = 18/79 (22%), Positives = 33/79 (40%)
Frame = -1
Query: 104 GLRVSPIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSL 163
G VSP RR G + + E + + +NN + F+ C + FTS
Sbjct: 262 GASVSP*PRRSGATAR*PRELRERIWWRHEYQSSGKPWRNNTTGPSPFSAMCIFMPFTST 83
Query: 164 IIFLHPPTSFPTNHISKKD 182
++ + P S ++K++
Sbjct: 82 VLCITPSISLTPPSLAKRE 26
>TC203322 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (74%)
Length = 1664
Score = 27.7 bits (60), Expect = 6.5
Identities = 13/26 (50%), Positives = 17/26 (65%)
Frame = -2
Query: 69 VAEMVESKELVGVVKGCIKSVQTPSG 94
V+E+VE VGVV GC+ Q+P G
Sbjct: 856 VSEVVEPYLDVGVVVGCLPETQSPRG 779
>TC205106 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (33%)
Length = 1796
Score = 27.3 bits (59), Expect = 8.5
Identities = 26/117 (22%), Positives = 47/117 (39%), Gaps = 4/117 (3%)
Frame = +3
Query: 174 PTNHISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSLG----TWVSYYK 229
PTN+ + + +DK ++ + +K K + + + L + +SLG TW
Sbjct: 1140 PTNYPNCSPILLDKFPVESSKENEDLSVKAKSRFSISLRS-LSQPMSLGEIARTW----- 1301
Query: 230 DEGFKLNIEDIITHKSTTSSSWIIFSLWNTCEACDNNLQVKTKLFQPLRFLHATLNH 286
D ++ I + S S + W C +L +F PL+F +NH
Sbjct: 1302 DVCARIVISEHAQQSGGGSFS-SKYGTWENCLTT**SLPFHANIFFPLKFAKTCVNH 1469
>BE610797
Length = 391
Score = 27.3 bits (59), Expect = 8.5
Identities = 14/30 (46%), Positives = 19/30 (62%), Gaps = 1/30 (3%)
Frame = +3
Query: 88 SVQTPSGSLFKM-GCILGLRVSPIHRRKGV 116
+++TP GSLF M CIL P+H+R V
Sbjct: 186 TIETPKGSLFLMHQCILPRPPPPLHQRHRV 275
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,117,795
Number of Sequences: 63676
Number of extensions: 208963
Number of successful extensions: 1190
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1182
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1186
length of query: 297
length of database: 12,639,632
effective HSP length: 97
effective length of query: 200
effective length of database: 6,463,060
effective search space: 1292612000
effective search space used: 1292612000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126780.2


