

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.36 + phase: 0
(475 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
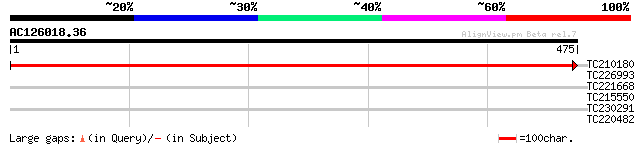
TC210180 UP|RBL_SOYBN (P27066) Ribulose bisphosphate carboxylase... 947 0.0
TC226993 homologue to UP|GMD1_ARATH (Q9SNY3) GDP-mannose 4,6 deh... 30 3.1
TC221668 homologue to UP|Q6ZZ86 (Q6ZZ86) LIN1 protein, partial (... 29 4.0
TC215550 similar to UP|Q8W423 (Q8W423) 26S proteasome regulatory... 28 6.8
TC230291 28 8.9
TC220482 similar to GB|AAS99727.1|46518491|BT012583 At2g27090 {A... 28 8.9
>TC210180 UP|RBL_SOYBN (P27066) Ribulose bisphosphate carboxylase large chain
precursor (RuBisCO large subunit) , complete
Length = 1556
Score = 947 bits (2447), Expect = 0.0
Identities = 453/475 (95%), Positives = 471/475 (98%)
Frame = +2
Query: 1 MSPQTETKATVGFKAGVKDYRLTYYTPDYETKDTDILAAFRVSPQPGVPAEEAGAAVAAE 60
MSPQTETKA+VGFKAGVKDY+LTYYTPDYETKDTDILAAFRV+PQPGVP EEAGAAVAAE
Sbjct: 17 MSPQTETKASVGFKAGVKDYKLTYYTPDYETKDTDILAAFRVTPQPGVPPEEAGAAVAAE 196
Query: 61 SSTGTWTTVWTDGLTSLDRYKGRCYHIEPVAGEESQFIAYVAYPLDLFEEGSVTNMFTSI 120
SSTGTWTTVWTDGLTSLDRYKGRCY +EPVAGEE+Q+IAYVAYPLDLFEEGSVTNMFTSI
Sbjct: 197 SSTGTWTTVWTDGLTSLDRYKGRCYGLEPVAGEENQYIAYVAYPLDLFEEGSVTNMFTSI 376
Query: 121 VGNVFGFKALRALRLEDLRIPVAYVKTFQGPPHGIQVERDKLNKYGRPLLGCTIKPKLGL 180
VGNVFGFKALRALRLEDLRIP +Y+KTFQGPPHGIQVERDKLNKYGRPLLGCTIKPKLGL
Sbjct: 377 VGNVFGFKALRALRLEDLRIPTSYIKTFQGPPHGIQVERDKLNKYGRPLLGCTIKPKLGL 556
Query: 181 SAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAIYKAQAETGEIKGHYL 240
SAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAI+K+QAETGEIKGHYL
Sbjct: 557 SAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAIFKSQAETGEIKGHYL 736
Query: 241 NATAGTCEDMMKRAVFARELGVPIVMHDYLTGGFTANTTLAHYCRDNGLLLHIHRAMHAV 300
NATAGTCE+MMKRAVFARELGVPIVMHDYLTGGFTANT+LAHYCRDNGLLLHIHRAMHAV
Sbjct: 737 NATAGTCEEMMKRAVFARELGVPIVMHDYLTGGFTANTSLAHYCRDNGLLLHIHRAMHAV 916
Query: 301 IDRQKNHGMHFRVLAKALRMSGGDHIHAGTVVGKLEGERDITLGFVDLLRDDFVEKDRSR 360
IDRQKNHGMHFRVLAKALR+SGGDH+HAGTVVGKLEGER+ITLGFVDLLRDDFVEKDRSR
Sbjct: 917 IDRQKNHGMHFRVLAKALRLSGGDHVHAGTVVGKLEGEREITLGFVDLLRDDFVEKDRSR 1096
Query: 361 GIFFTQDWVSLPGVLPVASGGIHVWHMPALTEIFGDDSVLQFGGGTLGHPWGNAPGAVAN 420
GI+FTQDWVSLPGVLPVASGGIHVWHMPALTEIFGDDSVLQFGGGTLGHPWGNAPGAVAN
Sbjct: 1097GIYFTQDWVSLPGVLPVASGGIHVWHMPALTEIFGDDSVLQFGGGTLGHPWGNAPGAVAN 1276
Query: 421 RVALEACVQARNEGRDLAREGNEIIREATKWSPELAAACEVWKEIKFEFPAMDTI 475
RVALEACVQARNEGRDLAREGNEIIREA+KWSPELAAACEVWKEIKFEF AMDT+
Sbjct: 1277RVALEACVQARNEGRDLAREGNEIIREASKWSPELAAACEVWKEIKFEFQAMDTL 1441
>TC226993 homologue to UP|GMD1_ARATH (Q9SNY3) GDP-mannose 4,6 dehydratase
(GDP-D-mannose dehydratase) (GMD) , partial (94%)
Length = 1388
Score = 29.6 bits (65), Expect = 3.1
Identities = 28/77 (36%), Positives = 39/77 (50%), Gaps = 7/77 (9%)
Frame = -3
Query: 321 SGGDHIHAGTVVGKLEGERDITLG--FVDLLRDDFVEKDRSRGIFFTQDWVSLPGVLPVA 378
+GGD + G VVG LEG+ D+ L VDL+ +D VE G V GV+ +
Sbjct: 462 AGGDDV--GGVVGYLEGDGDVRLSGEVVDLVGEDGVEPAAEGG------GVGEIGVVELH 307
Query: 379 SG-----GIHVWHMPAL 390
+G GIH+ + AL
Sbjct: 306 AGLVGVVGIHIDVVDAL 256
>TC221668 homologue to UP|Q6ZZ86 (Q6ZZ86) LIN1 protein, partial (18%)
Length = 813
Score = 29.3 bits (64), Expect = 4.0
Identities = 19/59 (32%), Positives = 24/59 (40%)
Frame = -3
Query: 169 LLGCTIKPKLGLSAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAIYK 227
+L C KP LG S+ NY A C K+ + WR + C AIYK
Sbjct: 652 ILSCDPKPALGQSSSNYSSATLGCFPA---HVKEKLCHLHSIVLLWRLQMQICI*AIYK 485
>TC215550 similar to UP|Q8W423 (Q8W423) 26S proteasome regulatory particle
non-ATPase subunit12, partial (66%)
Length = 787
Score = 28.5 bits (62), Expect = 6.8
Identities = 14/56 (25%), Positives = 22/56 (39%)
Frame = -1
Query: 192 CLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAIYKAQAETGEIKGHYLNATAGTC 247
CL G D+ N NS W LFC ++ + + + + + AG C
Sbjct: 163 CLMQGFSRAADESNSNSV----WNSAILFCTSSLNRVETQYRVLLRRWWXXXAGIC 8
>TC230291
Length = 677
Score = 28.1 bits (61), Expect = 8.9
Identities = 11/30 (36%), Positives = 17/30 (56%), Gaps = 4/30 (13%)
Frame = +1
Query: 444 IIREATKWSPELAA----ACEVWKEIKFEF 469
++ T WSPE C+VW E+KF++
Sbjct: 424 VVMTVT*WSPEAKRDHMPCCQVWNEVKFKW 513
>TC220482 similar to GB|AAS99727.1|46518491|BT012583 At2g27090 {Arabidopsis
thaliana;} , partial (13%)
Length = 550
Score = 28.1 bits (61), Expect = 8.9
Identities = 14/52 (26%), Positives = 22/52 (41%)
Frame = +2
Query: 424 LEACVQARNEGRDLAREGNEIIREATKWSPELAAACEVWKEIKFEFPAMDTI 475
L CV + + R ++R + P + A CE+W E E P D +
Sbjct: 227 LHKCVSLKQKPGKKKRPQRPLLR---MYGPPIYATCEIWLEKLGELPVQDVV 373
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,555,946
Number of Sequences: 63676
Number of extensions: 271778
Number of successful extensions: 1083
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1077
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1082
length of query: 475
length of database: 12,639,632
effective HSP length: 101
effective length of query: 374
effective length of database: 6,208,356
effective search space: 2321925144
effective search space used: 2321925144
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126018.36


