

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.19 - phase: 0
(686 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
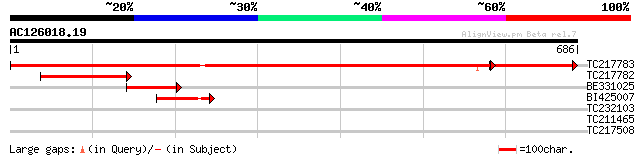
TC217783 UP|Q8HVY5 (Q8HVY5) RpoB, partial (71%) 1083 0.0
TC217782 UP|RPOC_SOYBN (Q8HVY4) DNA-directed RNA polymerase beta... 216 3e-56
BE331025 similar to EGAD|152251|162 RF548 (E. coli rpoC) {Nicoti... 126 3e-29
BI425007 similar to SP|Q9BBS8|RPOC_ DNA-directed RNA polymerase ... 86 5e-17
TC232103 weakly similar to UP|O22989 (O22989) Cellulose synthase... 31 2.1
TC211465 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp prote... 31 2.1
TC217508 similar to UP|Q6W2J1 (Q6W2J1) VDAC3.1, complete 29 6.0
>TC217783 UP|Q8HVY5 (Q8HVY5) RpoB, partial (71%)
Length = 4459
Score = 1083 bits (2802), Expect(2) = 0.0
Identities = 536/590 (90%), Positives = 552/590 (92%), Gaps = 3/590 (0%)
Frame = +2
Query: 1 MIDQYKHQHLRIGSVSPEQISAWAKKILPNGEIVGEVTKPYTLHYKTNKPEKDGLFCERI 60
MIDQYKHQ LRIG VSP+QISAWAKKILPNGEIVGEVTKPYT HY+TNKPEKDGLFCERI
Sbjct: 2327 MIDQYKHQQLRIGLVSPQQISAWAKKILPNGEIVGEVTKPYTFHYRTNKPEKDGLFCERI 2506
Query: 61 FGPIKSGICACGNYRVIGDKKDQPKFCEQCGVEFVDSRVRRYQMGYIKLACPVTHVWYLK 120
FGPIKSGICACGNYRVI DKKD PKFCEQCGVEF+DSR+RRYQMGYIKLAC VTHVWYLK
Sbjct: 2507 FGPIKSGICACGNYRVIRDKKDDPKFCEQCGVEFIDSRIRRYQMGYIKLACLVTHVWYLK 2686
Query: 121 RLPSYIASLLDKPLKELEGLVYCDFSFARPVVKKPTFLRLRGSFEYEIQSWKYSIPLFFA 180
RLPSYIA+LLDK LKELE LVYCDFSFARP+ KKPTFLRLRG FEYEIQSWKYSIPLFF
Sbjct: 2687 RLPSYIANLLDKSLKELESLVYCDFSFARPIAKKPTFLRLRGLFEYEIQSWKYSIPLFFT 2866
Query: 181 TQGFDTFRNREISSGAGAIREQLVDLDLRIIMDSSLVEWKELGEEGSADNENENEWEDRK 240
TQGFDTFRNREIS+GAGAIREQL DLDLR I+D S EWKELGEEGS NE WEDRK
Sbjct: 2867 TQGFDTFRNREISTGAGAIREQLADLDLRTIIDYSFAEWKELGEEGSTGNE----WEDRK 3034
Query: 241 VGRRKNFLVRRMELVKHFIRTNIEPEWMVLSLLPVLPPELRPIIQIDGGKLMSSDINELY 300
VGRRKNFLVRR+EL KHF+RTNIEPEWMVL LLPVLPPELRPIIQIDGGKLMSSDINELY
Sbjct: 3035 VGRRKNFLVRRIELAKHFLRTNIEPEWMVLCLLPVLPPELRPIIQIDGGKLMSSDINELY 3214
Query: 301 RRVIYRNNTLIDLLTTSRSTPGELVMCQEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSF 360
RRVIYRNNTLIDLLTTSRSTPGELVMCQEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSF
Sbjct: 3215 RRVIYRNNTLIDLLTTSRSTPGELVMCQEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSF 3394
Query: 361 SDIIEGKEGRFRETLLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFLIRGLIR 420
SDIIEGKEGRFRETLLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFLIRGLI
Sbjct: 3395 SDIIEGKEGRFRETLLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFLIRGLIP 3574
Query: 421 KHFASNIGVAKSKIREKEPIVWEILQEVMRGHPVLLNRAPTLHRLGIQAFQPILVEGRAI 480
KHFASNIG+AKSKIREKEPIVWEILQE M+GHPVLLNRAPTLHRLGIQAFQPILVEGRAI
Sbjct: 3575 KHFASNIGIAKSKIREKEPIVWEILQEGMQGHPVLLNRAPTLHRLGIQAFQPILVEGRAI 3754
Query: 481 CLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHTNLLSPAIGDPISVPTQDML 540
CLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHTNLLSPAIGDPISVPTQDML
Sbjct: 3755 CLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHTNLLSPAIGDPISVPTQDML 3934
Query: 541 IGLYVLTSGNRRGICANRYNPFNC---RNSKNEKISNNNSKYMKKKEPFF 587
IGLY+LTSGNRRGI +NRYNP NC RN KNE+I +NN KY KKK F
Sbjct: 3935 IGLYILTSGNRRGIYSNRYNPRNCGNFRNLKNERIRDNNYKYTKKKRTLF 4084
Score = 170 bits (431), Expect(2) = 0.0
Identities = 87/108 (80%), Positives = 92/108 (84%), Gaps = 2/108 (1%)
Frame = +3
Query: 581 KKKEPFFCNSYDAIGAYRQKRINLDSPFWLRWRIDQCIMSSREVPIEVHYESFGT-YYEI 639
KKKEPFFCNSYDAIGAY+QKRIN DSP WLRWR+DQ I+SSREVPIEVHYES GT +I
Sbjct: 4065 KKKEPFFCNSYDAIGAYQQKRINFDSPLWLRWRLDQRIISSREVPIEVHYESLGTLIMKI 4244
Query: 640 Y-GHYLVIRSIKKEIRCIYIRTTVGHISFYREIEEAIQGFSRAYSYGI 686
Y G V+RS KKEIR IYIRT VGHISFYREIEEAIQGF RAYSY I
Sbjct: 4245 Y*GII*VVRSTKKEIRSIYIRTNVGHISFYREIEEAIQGFCRAYSYDI 4388
>TC217782 UP|RPOC_SOYBN (Q8HVY4) DNA-directed RNA polymerase beta' chain
(PEP) (Plastid-encoded RNA polymerase beta' subunit)
(RNA polymerase beta' subunit) , partial (16%)
Length = 871
Score = 216 bits (550), Expect = 3e-56
Identities = 99/110 (90%), Positives = 102/110 (92%)
Frame = +2
Query: 38 TKPYTLHYKTNKPEKDGLFCERIFGPIKSGICACGNYRVIGDKKDQPKFCEQCGVEFVDS 97
TKPYT HYKTNKPEKDGLFCERIFGPIKSGICACGNYRVI DKKD PKFCEQCGVEF+DS
Sbjct: 2 TKPYTFHYKTNKPEKDGLFCERIFGPIKSGICACGNYRVIRDKKDDPKFCEQCGVEFIDS 181
Query: 98 RVRRYQMGYIKLACPVTHVWYLKRLPSYIASLLDKPLKELEGLVYCDFSF 147
R+RRYQMGYIKLAC VTHVWYLKRLPSYIA+LLDK LKELE LVYCD F
Sbjct: 182 RIRRYQMGYIKLACLVTHVWYLKRLPSYIANLLDKSLKELESLVYCDV*F 331
>BE331025 similar to EGAD|152251|162 RF548 (E. coli rpoC) {Nicotiana
tabacum}, partial (12%)
Length = 462
Score = 126 bits (317), Expect = 3e-29
Identities = 61/67 (91%), Positives = 63/67 (93%)
Frame = -1
Query: 142 YCDFSFARPVVKKPTFLRLRGSFEYEIQSWKYSIPLFFATQGFDTFRNREISSGAGAIRE 201
Y +FSFARPVVKKPTFLRLRGSFEYEIQSWK+SIPLFF TQGFD FRNREISSGAGAIRE
Sbjct: 201 YLNFSFARPVVKKPTFLRLRGSFEYEIQSWKHSIPLFFTTQGFDIFRNREISSGAGAIRE 22
Query: 202 QLVDLDL 208
QL DLDL
Sbjct: 21 QLADLDL 1
>BI425007 similar to SP|Q9BBS8|RPOC_ DNA-directed RNA polymerase beta' chain
(EC 2.7.7.6). {Lotus japonicus}, partial (9%)
Length = 428
Score = 85.9 bits (211), Expect = 5e-17
Identities = 51/70 (72%), Positives = 54/70 (76%)
Frame = -2
Query: 178 FFATQGFDTFRNREISSGAGAIREQLVDLDLRIIMDSSLVEWKELGEEGSADNENENEWE 237
FF TQGFD F NR ISSGA AIREQLVDLDLRI+M SSL+E KELGE S NEN+ E
Sbjct: 232 FFTTQGFDIF*NR*ISSGACAIREQLVDLDLRILMGSSLIE*KELGE**SP--YNENK*E 59
Query: 238 DRKVGRRKNF 247
DRKVGR K F
Sbjct: 58 DRKVGRTKIF 29
>TC232103 weakly similar to UP|O22989 (O22989) Cellulose synthase isolog,
partial (10%)
Length = 536
Score = 30.8 bits (68), Expect = 2.1
Identities = 16/57 (28%), Positives = 27/57 (47%)
Frame = +3
Query: 506 SLEAQAEARLLMFSHTNLLSPAIGDPISVPTQDMLIGLYVLTSGNRRGICANRYNPF 562
SLEA +E + + +G ++D+L GL + T G R +C+ +PF
Sbjct: 42 SLEAASEVSSCEYEYGTAWGKQVGWMYGSTSEDLLTGLKIHTKGWRSEVCSPELSPF 212
>TC211465 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (15%)
Length = 593
Score = 30.8 bits (68), Expect = 2.1
Identities = 19/54 (35%), Positives = 26/54 (47%)
Frame = +3
Query: 179 FATQGFDTFRNREISSGAGAIREQLVDLDLRIIMDSSLVEWKELGEEGSADNEN 232
F GFD ++ AGAI+E + L ++M SLV W +G D EN
Sbjct: 105 FCISGFDIVIGYLCTASAGAIQESWLGFWLSLLM--SLV*WLSIGMVSKRDQEN 260
>TC217508 similar to UP|Q6W2J1 (Q6W2J1) VDAC3.1, complete
Length = 1448
Score = 29.3 bits (64), Expect = 6.0
Identities = 12/28 (42%), Positives = 18/28 (63%)
Frame = +2
Query: 107 IKLACPVTHVWYLKRLPSYIASLLDKPL 134
+ LAC + +W L+ LPSY +L KP+
Sbjct: 1016 LNLACLLNLIWRLQLLPSYHFTLFPKPM 1099
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.141 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,135,552
Number of Sequences: 63676
Number of extensions: 474529
Number of successful extensions: 2116
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 2097
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2110
length of query: 686
length of database: 12,639,632
effective HSP length: 104
effective length of query: 582
effective length of database: 6,017,328
effective search space: 3502084896
effective search space used: 3502084896
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC126018.19


