

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.13 - phase: 0
(320 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
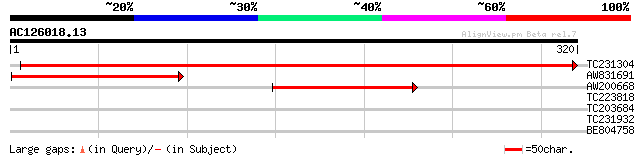
TC231304 UP|CYF_SOYBN (P49161) Apocytochrome f precursor , complete 589 e-169
AW831691 similar to SP|P49161|CYF_S Apocytochrome f precursor. [... 136 1e-32
AW200668 similar to SP|P49161|CYF_S Apocytochrome f precursor. [... 97 7e-21
TC223818 homologue to UP|Q9AYM4 (Q9AYM4) VuP5CS protein, partial... 28 5.5
TC203684 weakly similar to UP|Q7Q8B1 (Q7Q8B1) EbiP5126 (Fragment... 27 9.5
TC231932 similar to GB|AAS75312.1|45680886|AY568529 multidomain ... 27 9.5
BE804758 27 9.5
>TC231304 UP|CYF_SOYBN (P49161) Apocytochrome f precursor , complete
Length = 964
Score = 589 bits (1518), Expect = e-169
Identities = 296/314 (94%), Positives = 303/314 (96%)
Frame = +2
Query: 7 FSWIKEEITRSISVLLMIYIITRAPISNAYPIFAQQGYENPREATGRIVCANCHLANKPV 66
F+ IKE ITRSIS+ +MIYII RAPISNAYPIFAQQGYENPREATGRIVCANCHLANKPV
Sbjct: 20 FACIKEGITRSISISIMIYIIIRAPISNAYPIFAQQGYENPREATGRIVCANCHLANKPV 199
Query: 67 DIEVPQAVLPDTVFEAVVRIPYDMQVKQVLANGKKGALNVGAVLILPEGFELAPPDRISP 126
DIEVPQAVLPDTVFEAVVRIPYDMQVKQVLANGKKGALNVGAVLILPEGFELAPPDRISP
Sbjct: 200 DIEVPQAVLPDTVFEAVVRIPYDMQVKQVLANGKKGALNVGAVLILPEGFELAPPDRISP 379
Query: 127 EIKEKIGNLSFQSYRPTKKNILVVGPVPGKKYSEITFPILSPDPATKRDVHFLKYPIYVG 186
EIKEKIGNLSFQ+YRPTKKNILVVGPVPG+KY EITFPILSPDP TKRDVHFLKYPIYVG
Sbjct: 380 EIKEKIGNLSFQNYRPTKKNILVVGPVPGQKYKEITFPILSPDPTTKRDVHFLKYPIYVG 559
Query: 187 GNRGRGQIYPDGSKSNNNVYNATATGIVNKIIRKEKGGYEITIVDASDGREVIDIIPPGP 246
GNRGRGQIY DGSKSNNNVYNATA G+V KIIRKEKGGYEITIVDA DGREVIDIIPPGP
Sbjct: 560 GNRGRGQIYLDGSKSNNNVYNATAAGMVKKIIRKEKGGYEITIVDALDGREVIDIIPPGP 739
Query: 247 ELLVSEGESMKLDQPLTSNPNVGGFGQGDAEIVLQDPLRVQGLLFFLASIILAQIFLVLK 306
ELLVSEGES+KLDQPLTSNPNVGGFGQGDAEIVLQDPLRVQGLLFF ASIILAQIFLVLK
Sbjct: 740 ELLVSEGESIKLDQPLTSNPNVGGFGQGDAEIVLQDPLRVQGLLFFFASIILAQIFLVLK 919
Query: 307 KKQFEKVQLSEMNF 320
KKQFEKVQLSEMNF
Sbjct: 920 KKQFEKVQLSEMNF 961
>AW831691 similar to SP|P49161|CYF_S Apocytochrome f precursor. [Soybean]
{Glycine max}, partial (30%)
Length = 292
Score = 136 bits (342), Expect = 1e-32
Identities = 66/97 (68%), Positives = 80/97 (82%)
Frame = +2
Query: 2 QTRNAFSWIKEEITRSISVLLMIYIITRAPISNAYPIFAQQGYENPREATGRIVCANCHL 61
QT N FS+IKE IT+SIS+ +MIYII RAPISNAYPIF QQ Y+NPRE T RI+ ANCH+
Sbjct: 2 QTXNTFSYIKEGITQSISISIMIYIIIRAPISNAYPIFTQQNYKNPRETTSRIMYANCHV 181
Query: 62 ANKPVDIEVPQAVLPDTVFEAVVRIPYDMQVKQVLAN 98
ANKPVDI+V AVLPD +FE R+ YD++VKQV+A+
Sbjct: 182ANKPVDIKVRTAVLPDAIFETSGRMRYDVRVKQVVAS 292
>AW200668 similar to SP|P49161|CYF_S Apocytochrome f precursor. [Soybean]
{Glycine max}, partial (19%)
Length = 250
Score = 97.4 bits (241), Expect = 7e-21
Identities = 50/83 (60%), Positives = 58/83 (69%), Gaps = 1/83 (1%)
Frame = +2
Query: 149 VVG-PVPGKKYSEITFPILSPDPATKRDVHFLKYPIYVGGNRGRGQIYPDGSKSNNNVYN 207
VVG PV +KY EIT PILSPDP T RDVHFLKYPIYV N G +Y SKSNNNVYN
Sbjct: 2 VVGRPVTAQKYKEITSPILSPDPTTNRDVHFLKYPIYVRVNSGSRHLYLSASKSNNNVYN 181
Query: 208 ATATGIVNKIIRKEKGGYEITIV 230
AT G+V +++ G Y+ TI+
Sbjct: 182 ATTCGMVQHLLQY*TG*YDTTIL 250
>TC223818 homologue to UP|Q9AYM4 (Q9AYM4) VuP5CS protein, partial (16%)
Length = 772
Score = 28.1 bits (61), Expect = 5.5
Identities = 14/51 (27%), Positives = 23/51 (44%)
Frame = -1
Query: 18 ISVLLMIYIITRAPISNAYPIFAQQGYENPREATGRIVCANCHLANKPVDI 68
+SV+ + I P + YP QQ + R ++ C N +L KP +
Sbjct: 355 VSVIDPLISINHMPTPSQYPSVCQQPLNSYRTSSMNSTCTNSNLCAKPKSV 203
>TC203684 weakly similar to UP|Q7Q8B1 (Q7Q8B1) EbiP5126 (Fragment), partial
(49%)
Length = 1062
Score = 27.3 bits (59), Expect = 9.5
Identities = 13/34 (38%), Positives = 19/34 (55%)
Frame = +3
Query: 140 YRPTKKNILVVGPVPGKKYSEITFPILSPDPATK 173
Y+P K++ +V P+P I PIL P+P K
Sbjct: 555 YKPIPKSVPIVKPIP------IIKPILKPNPIVK 638
>TC231932 similar to GB|AAS75312.1|45680886|AY568529 multidomain cyclophilin
type peptidyl-prolyl cis-trans isomerase {Arabidopsis
thaliana;} , partial (13%)
Length = 329
Score = 27.3 bits (59), Expect = 9.5
Identities = 15/57 (26%), Positives = 26/57 (45%), Gaps = 1/57 (1%)
Frame = -3
Query: 5 NAFSWIKEEITRSISVLLMIYIITRAPISNAYPIFAQQGYENPRE-ATGRIVCANCH 60
N++SW+ IT + L I + + P +PI +++ E T + NCH
Sbjct: 285 NSYSWMPFFITEFFNFLQYIMPLHKLPKDYMFPIKMMISFQSDNEMGTECVTVCNCH 115
>BE804758
Length = 457
Score = 27.3 bits (59), Expect = 9.5
Identities = 15/55 (27%), Positives = 23/55 (41%)
Frame = -2
Query: 18 ISVLLMIYIITRAPISNAYPIFAQQGYENPREATGRIVCANCHLANKPVDIEVPQ 72
+S L+ II AP+ A + P+ ATG C + P ++ PQ
Sbjct: 276 LSELMSTPIILDAPLDRAPSATERPTARKPKTATGEPKCTSAVFQAAPTPVDTPQ 112
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.140 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,168,420
Number of Sequences: 63676
Number of extensions: 153050
Number of successful extensions: 669
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 661
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 669
length of query: 320
length of database: 12,639,632
effective HSP length: 97
effective length of query: 223
effective length of database: 6,463,060
effective search space: 1441262380
effective search space used: 1441262380
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126018.13


