

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.1 + phase: 0 /pseudo
(596 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
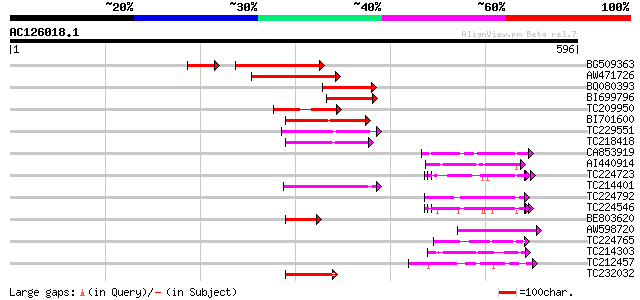
Score E
Sequences producing significant alignments: (bits) Value
BG509363 similar to GP|14331118|e ABC1 protein {Nicotiana plumba... 177 3e-50
AW471726 similar to PIR|T45888|T45 ABC transporter-like protein ... 135 5e-32
BQ080393 90 3e-18
BI699796 similar to PIR|G85167|G85 ABC transporter like protein ... 88 9e-18
TC209950 similar to UP|Q7PC84 (Q7PC84) PDR11 ABC transporter, pa... 74 2e-13
BI701600 71 2e-12
TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, p... 66 4e-11
TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partia... 60 3e-09
CA853919 homologue to PIR|T11622|T11 extensin class 1 precursor ... 56 4e-08
AI440914 53 3e-07
TC224723 hydroxyproline-rich glycoprotein 51 2e-06
TC214401 weakly similar to UP|Q949G3 (Q949G3) ABC1 protein, part... 50 2e-06
TC224792 homologue to UP|Q39865 (Q39865) Hydroxyproline-rich gly... 50 3e-06
TC224546 similar to UP|Q41707 (Q41707) Extensin class 1 protein ... 49 5e-06
BE803620 homologue to GP|13605839|gb| At2g01320/F10A8.20 {Arabid... 49 6e-06
AW598720 weakly similar to GP|21322711|e pherophorin-dz1 protein... 48 1e-05
TC224765 similar to UP|Q09082 (Q09082) Extensin (Class I), parti... 47 2e-05
TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, pa... 47 3e-05
TC212457 UP|Q39871 (Q39871) Maturation polypeptide, complete 46 4e-05
TC232032 weakly similar to UP|Q6K376 (Q6K376) ABC transporter-li... 46 5e-05
>BG509363 similar to GP|14331118|e ABC1 protein {Nicotiana plumbaginifolia},
partial (8%)
Length = 455
Score = 177 bits (450), Expect(2) = 3e-50
Identities = 87/94 (92%), Positives = 91/94 (96%)
Frame = +1
Query: 238 SGRADSVTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSG 297
SGR DSV ESSHGKKKGMVLPFEPHSITFD+++YSVDMP EMKEQGV+EDRLVLLKGVSG
Sbjct: 172 SGRGDSVVESSHGKKKGMVLPFEPHSITFDEVIYSVDMPQEMKEQGVQEDRLVLLKGVSG 351
Query: 298 AFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI 331
AFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI
Sbjct: 352 AFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI 453
Score = 39.7 bits (91), Expect(2) = 3e-50
Identities = 20/33 (60%), Positives = 24/33 (72%)
Frame = +3
Query: 188 GIGWICLSFQRGVWCGSRCTRPI**A*CNNN*R 220
G GWIC++FQR V+C +R PI**A NNN R
Sbjct: 18 GNGWICVAFQRDVFCCTRDPWPI**ATSNNNRR 116
>AW471726 similar to PIR|T45888|T45 ABC transporter-like protein -
Arabidopsis thaliana, partial (6%)
Length = 425
Score = 135 bits (340), Expect = 5e-32
Identities = 66/93 (70%), Positives = 77/93 (81%)
Frame = +3
Query: 255 MVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAG 314
MVLPFEP SI F D+ Y VD+P EMK+ G E RL LL ++GAFRPG+LTALMGVSGAG
Sbjct: 144 MVLPFEPLSIAFKDVQYFVDIPPEMKKHGSDEKRLQLLCDITGAFRPGILTALMGVSGAG 323
Query: 315 KTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETF 347
KTTLMDVL+GRKTGG I+GDI++ GYPK Q+TF
Sbjct: 324 KTTLMDVLSGRKTGGIIEGDIRIGGYPKVQKTF 422
>BQ080393
Length = 347
Score = 89.7 bits (221), Expect = 3e-18
Identities = 39/56 (69%), Positives = 47/56 (83%)
Frame = +2
Query: 330 YIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSN 385
YI+GDI++SG+PK QETFAR+SGYCE DIHSP VT+ ESLLYSA+LRLP V +
Sbjct: 2 YIEGDIRISGFPKNQETFARVSGYCEQTDIHSPQVTIRESLLYSAYLRLPKEVSKD 169
>BI699796 similar to PIR|G85167|G85 ABC transporter like protein [imported] -
Arabidopsis thaliana, partial (13%)
Length = 421
Score = 88.2 bits (217), Expect = 9e-18
Identities = 36/53 (67%), Positives = 47/53 (87%)
Frame = +1
Query: 334 DIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNT 386
+I++ GYPK QETFAR+SGYCE NDIHSP++TV ES+++SAWLRLPS +D+ T
Sbjct: 7 EIRIGGYPKVQETFARVSGYCEQNDIHSPNITVEESVMFSAWLRLPSQIDAKT 165
>TC209950 similar to UP|Q7PC84 (Q7PC84) PDR11 ABC transporter, partial (4%)
Length = 769
Score = 73.9 bits (180), Expect = 2e-13
Identities = 39/71 (54%), Positives = 48/71 (66%)
Frame = +2
Query: 278 EMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKV 337
EM+ Q + +D+L LL+ VSGAFRPG+LT TL+DV AGRK GGYI+G I +
Sbjct: 383 EMRSQ*INKDQLQLLQDVSGAFRPGILT-----------TLVDVFAGRKIGGYIEGSITM 529
Query: 338 SGYPKKQETFA 348
GYPK Q TFA
Sbjct: 530 *GYPKNQTTFA 562
>BI701600
Length = 426
Score = 70.9 bits (172), Expect = 2e-12
Identities = 38/89 (42%), Positives = 58/89 (64%)
Frame = +3
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARI 350
+LKGV+G +PG + A++G SG+GK+TL+ LAGR G + G I ++ K + R
Sbjct: 72 ILKGVTGIAQPGEILAVLGPSGSGKSTLLHALAGRLHGPGLTGTI-LANSSKLTKPVLRR 248
Query: 351 SGYCEHNDIHSPHVTVHESLLYSAWLRLP 379
+G+ +DI PH+TV E+L++ A LRLP
Sbjct: 249 TGFVTQDDILYPHLTVRETLVFCAMLRLP 335
>TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (34%)
Length = 1436
Score = 66.2 bits (160), Expect = 4e-11
Identities = 43/107 (40%), Positives = 62/107 (57%), Gaps = 1/107 (0%)
Frame = +3
Query: 286 EDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYID-GDIKVSGYPKKQ 344
++R ++L G++G +PG L A++G SG+GK+TL+D LAGR T G I ++G+ KQ
Sbjct: 60 KNRKLILHGLTGYAQPGRLLAIIGPSGSGKSTLLDALAGRLTSNIKQTGKILINGH--KQ 233
Query: 345 ETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNTTKVSE 391
E SGY +D +T E+L YSA L+ P NT V E
Sbjct: 234 ELAYGTSGYVTQDDAMLSCLTAGETLYYSAMLQFP-----NTMSVEE 359
>TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (49%)
Length = 1220
Score = 60.1 bits (144), Expect = 3e-09
Identities = 35/93 (37%), Positives = 55/93 (58%), Gaps = 1/93 (1%)
Frame = +1
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGR-KTGGYIDGDIKVSGYPKKQETFAR 349
+L+G++G PG TALMG SG+GK+TL+D L+ R ++ G I ++G +K +
Sbjct: 298 VLEGLTGYAEPGTFTALMGPSGSGKSTLLDALSSRLAANAFLSGTILLNG--RKAKLSFG 471
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGV 382
+ Y +D +TV E++ YSA LRLP +
Sbjct: 472 TAAYVTQDDNLIGTLTVRETISYSARLRLPDNM 570
>CA853919 homologue to PIR|T11622|T11 extensin class 1 precursor - cowpea,
partial (30%)
Length = 454
Score = 56.2 bits (134), Expect = 4e-08
Identities = 40/117 (34%), Positives = 51/117 (43%)
Frame = +3
Query: 434 LTVFHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVF 493
LT+ + P P P P P + H+ PS PY+ P P P P P Y +
Sbjct: 30 LTIINAPP-PPSPVPKPPYYYHSPPPPSPSPPPPYYYKSPP---PPSPSPPPPYY--YKS 191
Query: 494 PPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFF 550
PPP P P PSP+ PP PP S SP P + + H P PSP P ++
Sbjct: 192 PPPPPYYYKSPPPPSPSPPPPYYYHSPPPPSPSPPPPY-YYHSPPPPSPSPPPPYYY 359
Score = 37.0 bits (84), Expect = 0.025
Identities = 32/106 (30%), Positives = 41/106 (38%), Gaps = 11/106 (10%)
Frame = +3
Query: 437 FHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDP---FFF----PSYPFTPSPLYS 489
+H P P P P P + + PS PY+ P +++ P P P P Y
Sbjct: 87 YHSPP-PPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPPYYYKSPPPPSPSPPPPYYY 263
Query: 490 SFVFPP----PFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLF 531
PP P P P PSP+ PP PP S SP P +
Sbjct: 264 HSPPPPSPSPPPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPPXY 401
Score = 30.4 bits (67), Expect = 2.3
Identities = 26/90 (28%), Positives = 35/90 (38%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTR 503
P P P P + H+ PS PY+ P P +PSP PPP+ +
Sbjct: 231 PSPSPPPPYYYHSPPPPSPSPPPPYYYHSPP------PPSPSP-------PPPY---YYK 362
Query: 504 SPLPSPAARPPTAIFLPPITSSSPFPLFPH 533
SP P + PP + P S+ L H
Sbjct: 363 SPPPPSPSPPPXYYYQSPPPPSTIKVLLHH 452
Score = 30.0 bits (66), Expect = 3.0
Identities = 25/75 (33%), Positives = 32/75 (42%), Gaps = 15/75 (20%)
Frame = +3
Query: 487 LYSSFVFPPPFPMAVTRSPL-------PSPAARPPTAIFLPPITSSSPFPLF-------- 531
L + + PP P V + P PSP+ PP PP S SP P +
Sbjct: 24 LLLTIINAPPPPSPVPKPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPP 203
Query: 532 PHSHKLLPLFPSPSP 546
P+ +K P PSPSP
Sbjct: 204 PYYYKSPPP-PSPSP 245
>AI440914
Length = 344
Score = 53.1 bits (126), Expect = 3e-07
Identities = 39/111 (35%), Positives = 49/111 (44%), Gaps = 6/111 (5%)
Frame = +3
Query: 438 HFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPF 497
H P P PFP+ T ++ PS P PF P+ P TPSP S P PF
Sbjct: 21 HSPPSPATPSPFPSPT--TPVTPSPSHSPPSPATPSPFPSPTTPVTPSPPSPS--TPSPF 188
Query: 498 PMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLF------PHSHKLLPLFP 542
P + T PSP PP+ ++LPP SS P PH P++P
Sbjct: 189 P-SPTTPVTPSPVHSPPSPVYLPPPHSSPTPPAHNYGSPPPHHSPAQPVYP 338
Score = 34.3 bits (77), Expect = 0.16
Identities = 27/75 (36%), Positives = 36/75 (48%), Gaps = 1/75 (1%)
Frame = +3
Query: 483 TPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFP 542
TPSP +S PP P T SP PSP + P + ++P P + + P P
Sbjct: 6 TPSPSHS-----PPSP--ATPSPFPSPTTPVTPSPSHSPPSPATPSPFPSPTTPVTPSPP 164
Query: 543 SPS-PAVFFSPSCAV 556
SPS P+ F SP+ V
Sbjct: 165 SPSTPSPFPSPTTPV 209
>TC224723 hydroxyproline-rich glycoprotein
Length = 2345
Score = 50.8 bits (120), Expect = 2e-06
Identities = 38/107 (35%), Positives = 45/107 (41%), Gaps = 4/107 (3%)
Frame = +1
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP----PFPM 499
P P P P + H+ PS PY+ P P P PSP Y PP P P
Sbjct: 892 PSPSPPPPYYYHSPPPPSPSPPPPYYYHSPP---PPSPSPPSPYYYKSPPPPSPSPPPPY 1062
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P PSP + PP PP +S P P + H + P PSPSP
Sbjct: 1063YYQSPPPPSPTSHPPYYYKSPPPPTSYPPPPY---HYVSPPPPSPSP 1194
Score = 46.6 bits (109), Expect = 3e-05
Identities = 40/116 (34%), Positives = 51/116 (43%), Gaps = 5/116 (4%)
Frame = +1
Query: 437 FHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPP 496
+H P P P P P + H+ PS SPY+ P P P P P Y PPP
Sbjct: 922 YHSPP-PPSPSPPPPYYYHSPPPPSPSPPSPYYYKSPP---PPSPSPPPPYYYQSP-PPP 1086
Query: 497 FPMA----VTRSPLPSPAARPPTAIFL-PPITSSSPFPLFPHSHKLLPLFPSPSPA 547
P + +SP P + PP ++ PP S SP P + H P PSP+PA
Sbjct: 1087 SPTSHPPYYYKSPPPPTSYPPPPYHYVSPPPPSPSPPPPY---HYTSPPPPSPAPA 1245
Score = 44.3 bits (103), Expect = 2e-04
Identities = 39/114 (34%), Positives = 50/114 (43%), Gaps = 1/114 (0%)
Frame = +1
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPM 499
PS +P PPP+ +Y S P + P+++ S P PSP PP P
Sbjct: 745 PSPSP-PPPY----------YYHSPPPPKEHVHPPYYYHSPPPPPSP-------SPPPPY 870
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPS-PSPAVFFSP 552
P PSP+ PP PP S SP P + + H P PS PSP + SP
Sbjct: 871 YYKSPPPPSPSPPPPYYYHSPPPPSPSPPPPY-YYHSPPPPSPSPPSPYYYKSP 1029
Score = 41.6 bits (96), Expect = 0.001
Identities = 36/107 (33%), Positives = 42/107 (38%), Gaps = 4/107 (3%)
Frame = +1
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP----PFPM 499
P P P P + H+ PY+ P P P P P Y PP P P
Sbjct: 745 PSPSPPPPYYYHSPPPPKEHVHPPYYYHSPPP--PPSPSPPPPYYYKSPPPPSPSPPPPY 918
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P PSP+ PP PP SP P P+ +K P PSPSP
Sbjct: 919 YYHSPPPPSPSPPPPYYYHSPP--PPSPSPPSPYYYKSPPP-PSPSP 1050
Score = 39.3 bits (90), Expect = 0.005
Identities = 30/84 (35%), Positives = 35/84 (40%), Gaps = 3/84 (3%)
Frame = -3
Query: 467 PYFTLLDPFFFPSYPFTPSPLYSSFVFPP---PFPMAVTRSPLPSPAARPPTAIFLPPIT 523
PY+ P P P P P Y PP P P +SP P + PP PP
Sbjct: 567 PYYYKSPP---PPSPSPPPPYYYQSPPPPSPTPHPPYYYKSPPPPTSHPPPYHYVSPPPP 397
Query: 524 SSSPFPLFPHSHKLLPLFPSPSPA 547
S SP P + H P PSP+PA
Sbjct: 396 SPSPPPPY---HYTSPPPPSPAPA 334
Score = 36.2 bits (82), Expect = 0.042
Identities = 27/86 (31%), Positives = 37/86 (42%), Gaps = 9/86 (10%)
Frame = +1
Query: 474 PFFFPSYPF-TPSPLYSSFVFPPPFPMAVTRSPL--------PSPAARPPTAIFLPPITS 524
P+++ S P +PSP + PP P P PSP+ PP PP S
Sbjct: 718 PYYYKSPPPPSPSPPPPYYYHSPPPPKEHVHPPYYYHSPPPPPSPSPPPPYYYKSPPPPS 897
Query: 525 SSPFPLFPHSHKLLPLFPSPSPAVFF 550
SP P + + H P PSP P ++
Sbjct: 898 PSPPPPY-YYHSPPPPSPSPPPPYYY 972
Score = 35.0 bits (79), Expect = 0.094
Identities = 29/98 (29%), Positives = 39/98 (39%), Gaps = 10/98 (10%)
Frame = +1
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPF------ 497
P P P P + + P+ PY+ P P + P P + +V PPP
Sbjct: 1036 PSPSPPPPYYYQSPPPPSPTSHPPYYYKSPP---PPTSYPPPPYH--YVSPPPPSPSPPP 1200
Query: 498 PMAVTRSPLPSPAARPPTAIFLPP----ITSSSPFPLF 531
P T P PSPA P PP I +S P P++
Sbjct: 1201 PYHYTSPPPPSPAPAPKYIYKSPPPPVYIYASPPPPIY 1314
Score = 33.5 bits (75), Expect = 0.27
Identities = 22/64 (34%), Positives = 26/64 (40%), Gaps = 12/64 (18%)
Frame = -3
Query: 495 PPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLF------------PHSHKLLPLFP 542
PP P P PSP+ PP PP S +P P + P H + P P
Sbjct: 576 PPPPYYYKSPPPPSPSPPPPYYYQSPPPPSPTPHPPYYYKSPPPPTSHPPPYHYVSPPPP 397
Query: 543 SPSP 546
SPSP
Sbjct: 396 SPSP 385
Score = 32.7 bits (73), Expect = 0.47
Identities = 28/97 (28%), Positives = 37/97 (37%), Gaps = 9/97 (9%)
Frame = -3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMA--- 500
P P P P + + P+ PY+ + P P + P Y PPP P
Sbjct: 540 PSPSPPPPYYYQSPPPPSPTPHPPYY-----YKSPPPPTSHPPPYHYVSPPPPSPSPPPP 376
Query: 501 --VTRSPLPSPAARPPTAIFLPP----ITSSSPFPLF 531
T P PSPA P PP I +S P P++
Sbjct: 375 YHYTSPPPPSPAPAPKYIYKSPPPPVYIYASPPPPIY 265
>TC214401 weakly similar to UP|Q949G3 (Q949G3) ABC1 protein, partial (18%)
Length = 1177
Score = 50.4 bits (119), Expect = 2e-06
Identities = 35/104 (33%), Positives = 54/104 (51%), Gaps = 1/104 (0%)
Frame = +1
Query: 289 LVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGY-IDGDIKVSGYPKKQETF 347
L +L+ VSG +P +T L+G G+GKTTL+ LAG+ G + +G+ ++
Sbjct: 757 LRILQNVSGIIKPRRMTLLLGPPGSGKTTLLLALAGKLDKDLNHSGRVTYNGHGLEEFVP 936
Query: 348 ARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNTTKVSE 391
R S Y D H +TV E+L +SA + GV N ++E
Sbjct: 937 QRTSAYISQYDNHIGEMTVRETLAFSARCQ---GVGQNYEMLAE 1059
>TC224792 homologue to UP|Q39865 (Q39865) Hydroxyproline-rich glycoprotein
(Fragment), partial (86%)
Length = 688
Score = 50.1 bits (118), Expect = 3e-06
Identities = 37/111 (33%), Positives = 48/111 (42%), Gaps = 1/111 (0%)
Frame = +2
Query: 437 FHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPF-TPSPLYSSFVFPP 495
+H+PS P P P P + PS P P+++ S P +PSP + P
Sbjct: 137 YHYPSPPPPPSPSPPPPYYYKSPPPPSPSPP-----PPYYYHSPPPPSPSPPPPYYYHSP 301
Query: 496 PFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P P P PSP PP PP +S P P + H + P PSPSP
Sbjct: 302 PPPYYYQSPPPPSPTPHPPYYYKSPPPPTSHPPPPY---HYVSPPPPSPSP 445
Score = 35.4 bits (80), Expect = 0.072
Identities = 34/109 (31%), Positives = 44/109 (40%), Gaps = 14/109 (12%)
Frame = +2
Query: 437 FHFPSFAPLPPPFPAHLLHTSLS--HYPSGFSPYFTLLDPFFFPSYP---FTPSPLYSSF 491
+H P P P P P + H+ +Y S P T P+++ S P P P Y
Sbjct: 242 YHSPP-PPSPSPPPPYYYHSPPPPYYYQSPPPPSPTPHPPYYYKSPPPPTSHPPPPYHYV 418
Query: 492 VFPPPFPMA-----VTRSPLPSPAARPPTAIFLPP----ITSSSPFPLF 531
PPP P T P PSPA P PP I +S P P++
Sbjct: 419 SPPPPSPSPPPPYHYTSPPPPSPAPAPKYIYKSPPPPVYIYASPPPPIY 565
Score = 35.0 bits (79), Expect = 0.094
Identities = 32/109 (29%), Positives = 41/109 (37%), Gaps = 4/109 (3%)
Frame = +2
Query: 442 FAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAV 501
+ P+ PP+ HYPS P P +PSP PPP+
Sbjct: 116 YPPVSPPY----------HYPS--------------PPPPPSPSP-------PPPY--YY 196
Query: 502 TRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLF----PSPSP 546
P PSP+ PP PP S SP P + + P + P PSP
Sbjct: 197 KSPPPPSPSPPPPYYYHSPPPPSPSPPPPYYYHSPPPPYYYQSPPPPSP 343
Score = 32.3 bits (72), Expect = 0.61
Identities = 21/65 (32%), Positives = 29/65 (44%)
Frame = +2
Query: 486 PLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPS 545
P+ + +P P P PSP+ PP PP S SP P + + H P PSP
Sbjct: 122 PVSPPYHYPSP-------PPPPSPSPPPPYYYKSPPPPSPSPPPPY-YYHSPPPPSPSPP 277
Query: 546 PAVFF 550
P ++
Sbjct: 278 PPYYY 292
>TC224546 similar to UP|Q41707 (Q41707) Extensin class 1 protein precursor,
partial (67%)
Length = 1240
Score = 49.3 bits (116), Expect = 5e-06
Identities = 39/114 (34%), Positives = 47/114 (41%), Gaps = 3/114 (2%)
Frame = +3
Query: 437 FHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP- 495
+H P P P P P + H+ PS PY+ P P P P P Y PP
Sbjct: 630 YHSPP-PPSPSPPPPYYYHSPPPPSPSPPPPYYYKSPP---PPSPSPPPPYYYQSPPPPS 797
Query: 496 --PFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPA 547
P P +SP P + PP PP S SP P + H P PSP+PA
Sbjct: 798 PTPHPPYYYKSPPPPTSHPPPYHYVSPPPPSPSPPPPY---HYTSPPPPSPAPA 950
Score = 45.8 bits (107), Expect = 5e-05
Identities = 37/114 (32%), Positives = 47/114 (40%), Gaps = 7/114 (6%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTR 503
P P P P + H+ PS PY+ P P P P P Y + PPP P V +
Sbjct: 72 PSPIPKPPYYYHSPPPPSPSPPPPYYYKSPP---PPSPSPPPPYY--YKSPPP-PSPVPK 233
Query: 504 SPL-------PSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFF 550
P PSP+ PP PP S P P + + H P PSP P ++
Sbjct: 234 PPYYYHSPPPPSPSPPPPYYYKSPPPPSPVPKPPY-YYHSPPPPSPSPPPPYYY 392
Score = 44.7 bits (104), Expect = 1e-04
Identities = 39/114 (34%), Positives = 46/114 (40%), Gaps = 4/114 (3%)
Frame = +3
Query: 437 FHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP- 495
+H P P P P P + + PS PY+ P P P P Y PP
Sbjct: 102 YHSPP-PPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPP---PPSPVPKPPYYYHSPPPPS 269
Query: 496 ---PFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P P P PSP +PP PP S SP P P+ +K P PSPSP
Sbjct: 270 PSPPPPYYYKSPPPPSPVPKPPYYYHSPPPPSPSPPP--PYYYKSPPP-PSPSP 422
Score = 44.3 bits (103), Expect = 2e-04
Identities = 37/107 (34%), Positives = 43/107 (39%), Gaps = 4/107 (3%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP----PFPM 499
P P P P + H+ PY+ P P P P P Y PP P P
Sbjct: 456 PSPSPPPPYYYHSPPPPKEHVHPPYYYHSPP---PPSPSPPPPYYYKSPPPPSPSPPPPY 626
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P PSP+ PP PP S SP P P+ +K P PSPSP
Sbjct: 627 YYHSPPPPSPSPPPPYYYHSPPPPSPSPPP--PYYYKSPPP-PSPSP 758
Score = 44.3 bits (103), Expect = 2e-04
Identities = 37/107 (34%), Positives = 42/107 (38%), Gaps = 4/107 (3%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP----PFPM 499
P P P P + H+ PS PY+ P P P P P Y PP P P
Sbjct: 312 PSPVPKPPYYYHSPPPPSPSPPPPYYYKSPP---PPSPSPPPPYYYKSPPPPSPSPPPPY 482
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P P PP PP S SP P P+ +K P PSPSP
Sbjct: 483 YYHSPPPPKEHVHPPYYYHSPPPPSPSPPP--PYYYKSPPP-PSPSP 614
Score = 43.9 bits (102), Expect = 2e-04
Identities = 39/126 (30%), Positives = 47/126 (36%), Gaps = 16/126 (12%)
Frame = +3
Query: 437 FHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP- 495
+H P P P P P + + PS PY+ P P P P P Y PP
Sbjct: 534 YHSPP-PPSPSPPPPYYYKSPPPPSPSPPPPYYYHSPP---PPSPSPPPPYYYHSPPPPS 701
Query: 496 ---PFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLF------------PHSHKLLPL 540
P P P PSP+ PP PP S +P P + P H + P
Sbjct: 702 PSPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSPTPHPPYYYKSPPPPTSHPPPYHYVSPP 881
Query: 541 FPSPSP 546
PSPSP
Sbjct: 882 PPSPSP 899
Score = 43.9 bits (102), Expect = 2e-04
Identities = 38/120 (31%), Positives = 48/120 (39%), Gaps = 6/120 (5%)
Frame = +3
Query: 437 FHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPP 496
+H P P P P P + + PS PY+ P P P P P Y + PPP
Sbjct: 342 YHSPP-PPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPP---PPSPSPPPPYY--YHSPPP 503
Query: 497 F------PMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFF 550
P P PSP+ PP PP S SP P + + H P PSP P ++
Sbjct: 504 PKEHVHPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPPPY-YYHSPPPPSPSPPPPYYY 680
Score = 42.4 bits (98), Expect = 6e-04
Identities = 35/116 (30%), Positives = 47/116 (40%), Gaps = 9/116 (7%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFT---------LLDPFFFPSYPFTPSPLYSSFVFP 494
P P P P + + PS PY+ + P+++ S P PSP
Sbjct: 408 PSPSPPPPYYYKSPPPPSPSPPPPYYYHSPPPPKEHVHPPYYYHSPP-PPSP-------S 563
Query: 495 PPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFF 550
PP P P PSP+ PP PP S SP P + + H P PSP P ++
Sbjct: 564 PPPPYYYKSPPPPSPSPPPPYYYHSPPPPSPSPPPPY-YYHSPPPPSPSPPPPYYY 728
Score = 42.0 bits (97), Expect = 8e-04
Identities = 37/116 (31%), Positives = 45/116 (37%), Gaps = 9/116 (7%)
Frame = +3
Query: 440 PSFAPLPPP-----FPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFP 494
P + PPP P + H+ PS PY+ P P P P P Y P
Sbjct: 477 PYYYHSPPPPKEHVHPPYYYHSPPPPSPSPPPPYYYKSPP---PPSPSPPPPYYYHSPPP 647
Query: 495 P----PFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P P P P PSP+ PP PP S SP P + + P PSP+P
Sbjct: 648 PSPSPPPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYQS---PPPPSPTP 806
Score = 41.6 bits (96), Expect = 0.001
Identities = 36/103 (34%), Positives = 43/103 (40%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTR 503
P P P P + H+ PS PY+ P P P P Y + PPP
Sbjct: 216 PSPVPKPPYYYHSPPPPSPSPPPPYYYKSPP---PPSPVPKPPYY--YHSPPP------- 359
Query: 504 SPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
PSP+ PP PP S SP P P+ +K P PSPSP
Sbjct: 360 ---PSPSPPPPYYYKSPPPPSPSPPP--PYYYKSPPP-PSPSP 470
Score = 38.9 bits (89), Expect = 0.007
Identities = 35/117 (29%), Positives = 44/117 (36%), Gaps = 3/117 (2%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTR 503
P P P P + + P PY+ P P P P P Y + PPP
Sbjct: 24 PSPVPKPPYYYQSPPPPSPIPKPPYYYHSPP---PPSPSPPPPYY--YKSPPP------- 167
Query: 504 SPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFFS---PSCAVP 557
PSP+ PP PP S P P + + H P PSP P ++ P VP
Sbjct: 168 ---PSPSPPPPYYYKSPPPPSPVPKPPY-YYHSPPPPSPSPPPPYYYKSPPPPSPVP 326
Score = 34.7 bits (78), Expect = 0.12
Identities = 24/58 (41%), Positives = 28/58 (47%), Gaps = 5/58 (8%)
Frame = +3
Query: 494 PPPFPMA-----VTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
PPP P+ P PSP +PP PP S SP P P+ +K P PSPSP
Sbjct: 18 PPPSPVPKPPYYYQSPPPPSPIPKPPYYYHSPPPPSPSPPP--PYYYKSPPP-PSPSP 182
Score = 32.7 bits (73), Expect = 0.47
Identities = 28/97 (28%), Positives = 37/97 (37%), Gaps = 9/97 (9%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMA--- 500
P P P P + + P+ PY+ + P P + P Y PPP P
Sbjct: 744 PSPSPPPPYYYQSPPPPSPTPHPPYY-----YKSPPPPTSHPPPYHYVSPPPPSPSPPPP 908
Query: 501 --VTRSPLPSPAARPPTAIFLPP----ITSSSPFPLF 531
T P PSPA P PP I +S P P++
Sbjct: 909 YHYTSPPPPSPAPAPKYIYKSPPPPVYIYASPPPPIY 1019
Score = 31.6 bits (70), Expect = 1.0
Identities = 25/73 (34%), Positives = 32/73 (43%), Gaps = 7/73 (9%)
Frame = +3
Query: 481 PFTPSPLYSSFVFPPPFPMAVTRSPL-------PSPAARPPTAIFLPPITSSSPFPLFPH 533
P +P P + PP P + + P PSP+ PP PP S SP P P+
Sbjct: 21 PPSPVPKPPYYYQSPPPPSPIPKPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPP--PY 194
Query: 534 SHKLLPLFPSPSP 546
+K P PSP P
Sbjct: 195 YYKSPPP-PSPVP 230
Score = 31.2 bits (69), Expect = 1.4
Identities = 17/46 (36%), Positives = 22/46 (46%)
Frame = +3
Query: 505 PLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFF 550
P PSP +PP PP S P P + + H P PSP P ++
Sbjct: 18 PPPSPVPKPPYYYQSPPPPSPIPKPPY-YYHSPPPPSPSPPPPYYY 152
>BE803620 homologue to GP|13605839|gb| At2g01320/F10A8.20 {Arabidopsis
thaliana}, partial (8%)
Length = 413
Score = 48.9 bits (115), Expect = 6e-06
Identities = 24/37 (64%), Positives = 30/37 (80%)
Frame = +3
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKT 327
LLK VSG +PG L A+MG SG+GKTTL++VLAG+ T
Sbjct: 294 LLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLT 404
>AW598720 weakly similar to GP|21322711|e pherophorin-dz1 protein {Volvox
carteri f. nagariensis}, partial (12%)
Length = 411
Score = 48.1 bits (113), Expect = 1e-05
Identities = 32/91 (35%), Positives = 41/91 (44%), Gaps = 2/91 (2%)
Frame = +2
Query: 471 LLDPFFFPSYPFTPSPLYSSFVFPPP-FPMAVTRSPLPSPAARPPTAIFLP-PITSSSPF 528
L + FFFP PF P PL + PPP P PL +PP+ +P P SP
Sbjct: 20 LAEDFFFPPNPFFPPPLVPNPFQPPPLIPNPFQPPPLIPNPLQPPSPPLIPNPFQPPSPP 199
Query: 529 PLFPHSHKLLPLFPSPSPAVFFSPSCAVPLV 559
PL P+ + P P+ P F P PL+
Sbjct: 200 PLIPNPFQPPPSPPTLFPNPFQPPPSPPPLI 292
Score = 38.5 bits (88), Expect = 0.009
Identities = 38/122 (31%), Positives = 48/122 (39%), Gaps = 12/122 (9%)
Frame = +2
Query: 437 FHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSP--------LY 488
F FP PPP + P+ F P + +P PS P P+P L
Sbjct: 32 FFFPPNPFFPPPLVPNPFQPP-PLIPNPFQPPPLIPNPLQPPSPPLIPNPFQPPSPPPLI 208
Query: 489 SSFVFPPPFPMAVTRSPL---PSPAARPPTAIFLPPITSSSPFPLFPHSHKLLP-LFPSP 544
+ PPP P + +P PSP P PP S FP FP ++P L PSP
Sbjct: 209 PNPFQPPPSPPTLFPNPFQPPPSPPPLIPNPFQPPPSPPPSLFPPFPPI--VIPGLTPSP 382
Query: 545 SP 546
P
Sbjct: 383 PP 388
Score = 33.1 bits (74), Expect = 0.36
Identities = 29/94 (30%), Positives = 39/94 (40%), Gaps = 10/94 (10%)
Frame = +2
Query: 446 PPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSP----LYSSFVFPPPFPMAV 501
PP P L S P+ F P P P+ PF P P L+ + PPP P +
Sbjct: 122 PPLIPNPLQPPSPPLIPNPFQPPSP---PPLIPN-PFQPPPSPPTLFPNPFQPPPSPPPL 289
Query: 502 TRSPLPSPAARPPT------AIFLPPITSSSPFP 529
+P P + PP+ I +P +T S P P
Sbjct: 290 IPNPFQPPPSPPPSLFPPFPPIVIPGLTPSPPPP 391
Score = 32.3 bits (72), Expect = 0.61
Identities = 30/84 (35%), Positives = 35/84 (40%), Gaps = 6/84 (7%)
Frame = +2
Query: 437 FHFPSFAPL------PPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSS 490
F PS PL PPP P L +P+ F P + P P+ PF P P
Sbjct: 179 FQPPSPPPLIPNPFQPPPSPPTL-------FPNPFQPPPS--PPPLIPN-PFQPPPSPPP 328
Query: 491 FVFPPPFPMAVTRSPLPSPAARPP 514
+FPP FP V PSP PP
Sbjct: 329 SLFPP-FPPIVIPGLTPSPPPPPP 397
>TC224765 similar to UP|Q09082 (Q09082) Extensin (Class I), partial (46%)
Length = 708
Score = 47.4 bits (111), Expect = 2e-05
Identities = 38/102 (37%), Positives = 45/102 (43%), Gaps = 1/102 (0%)
Frame = +3
Query: 446 PPPFPA-HLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRS 504
PPP P + H+ PS SPY+ P PSP+Y PPP P V +S
Sbjct: 15 PPPSPTPYYYHSPPPPSPSPPSPYYYKSPP--------PPSPVYIYKSPPPPSPTYVYKS 170
Query: 505 PLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P P P PP PP S P P P+ +K P PSP P
Sbjct: 171 P-PPPVKSPPYYYQSPPPPSPKPKP--PYYYKSPPP-PSPKP 284
Score = 40.4 bits (93), Expect = 0.002
Identities = 37/108 (34%), Positives = 44/108 (40%), Gaps = 5/108 (4%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVT- 502
P P P P + + P PY+ P P P P P Y PPP P+
Sbjct: 222 PSPKPKPPYYYKSPPPPSPKPKPPYYYQSPP---PPSP-VPKPPYYYKSPPPPSPVPKPP 389
Query: 503 ----RSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P PSP+ PP PP S SP P P+ +K P PSPSP
Sbjct: 390 YYYHSPPPPSPSPPPPYYYKSPPPPSPSPPP--PYYYKSPPP-PSPSP 524
Score = 40.4 bits (93), Expect = 0.002
Identities = 37/116 (31%), Positives = 49/116 (41%), Gaps = 15/116 (12%)
Frame = +3
Query: 446 PPPFPAHL-------LHTSLSHYPSGFSPYFTLLDPFFFPSYPFT---PSPLYSSFVFPP 495
PPP P ++ + + +Y S P P+++ S P P P Y PP
Sbjct: 138 PPPSPTYVYKSPPPPVKSPPYYYQSPPPPSPKPKPPYYYKSPPPPSPKPKPPYYYQSPPP 317
Query: 496 PFPMA-----VTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P P+ P PSP +PP PP S SP P P+ +K P PSPSP
Sbjct: 318 PSPVPKPPYYYKSPPPPSPVPKPPYYYHSPPPPSPSPPP--PYYYKSPPP-PSPSP 476
Score = 35.4 bits (80), Expect = 0.072
Identities = 31/110 (28%), Positives = 39/110 (35%), Gaps = 4/110 (3%)
Frame = +3
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPP----PFPM 499
P P P P + + P PY+ P P P P Y PP P P
Sbjct: 270 PSPKPKPPYYYQSPPPPSPVPKPPYYYKSPP---PPSPVPKPPYYYHSPPPPSPSPPPPY 440
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVF 549
P PSP+ PP PP S SP P + + SP P ++
Sbjct: 441 YYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPY--------YYHSPPPPIY 566
>TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, partial (18%)
Length = 1014
Score = 46.6 bits (109), Expect = 3e-05
Identities = 39/108 (36%), Positives = 46/108 (42%)
Frame = +2
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPM 499
P+++P PPP P+ H P P + L P P P P+YS PPP P
Sbjct: 44 PTYSPPPPP-PS----PPPPHSPPPPPPVYPYLSPPPPPPVHSPPPPVYSP-PPPPPSPP 205
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPA 547
SP P P PP PP SP P PHS P P PSP+
Sbjct: 206 PCIESPPPPP---PP-----PPCEEHSPPPPSPHS---APYHPPPSPS 316
Score = 38.9 bits (89), Expect = 0.007
Identities = 33/111 (29%), Positives = 45/111 (39%), Gaps = 3/111 (2%)
Frame = +2
Query: 439 FPSFAPLPPPFPAHLLHTSL-SHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPF 497
+P +P PPP P H + S P SP P P P P PPP
Sbjct: 116 YPYLSPPPPP-PVHSPPPPVYSPPPPPPSP-----PPCIESPPPPPPPPPCEEHSPPPPS 277
Query: 498 PMAVTRSPLPSPAARPPTAIF--LPPITSSSPFPLFPHSHKLLPLFPSPSP 546
P + P PSP+ PP + PP + P P++ ++ P P P+P
Sbjct: 278 PHSAPYHPPPSPSPPPPPIQYNSPPPPSPPPPTPVYHYNSPPPPSSPPPAP 430
Score = 37.7 bits (86), Expect = 0.015
Identities = 35/113 (30%), Positives = 46/113 (39%), Gaps = 6/113 (5%)
Frame = +2
Query: 444 PLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTR 503
P PPP P H P SP+ S P+ P P S PPP P+
Sbjct: 227 PPPPPPPCE------EHSPPPPSPH----------SAPYHPPPSPS----PPPPPIQYNS 346
Query: 504 SPLPSPAARPPTAIF----LPPITSSSPFPLF--PHSHKLLPLFPSPSPAVFF 550
P PSP PPT ++ PP +S P P++ P + + SP P F+
Sbjct: 347 PPPPSPP--PPTPVYHYNSPPPPSSPPPAPVYEGPLPPVIGVSYASPPPPPFY 499
Score = 32.3 bits (72), Expect = 0.61
Identities = 20/60 (33%), Positives = 29/60 (48%), Gaps = 1/60 (1%)
Frame = +2
Query: 494 PPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPH-SHKLLPLFPSPSPAVFFSP 552
PPP P+ +P P+ + PP PP + P P++P+ S P SP P V+ P
Sbjct: 5 PPPPPVYSPPAPPPTYSPPPPPPSPPPPHSPPPPPPVYPYLSPPPPPPVHSPPPPVYSPP 184
>TC212457 UP|Q39871 (Q39871) Maturation polypeptide, complete
Length = 1741
Score = 46.2 bits (108), Expect = 4e-05
Identities = 49/147 (33%), Positives = 60/147 (40%), Gaps = 12/147 (8%)
Frame = -1
Query: 420 HAMLSAELTPLMTSLTVFH-----FPSFAPLPPPFPA--HLL--HTSLSHYPSGFSPYFT 470
H +L E L S FH FP +PLP F A H L + S +PS PY +
Sbjct: 1288 HTLLYLEALALPFSSLQFHLHHSHFPWCSPLPVQFSASPHPLSQRSCPSLHPSSPKPYSS 1109
Query: 471 LLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRSPLP---SPAARPPTAIFLPPITSSSP 527
S P +P +S PPP P SP P P PP+ LPP SSP
Sbjct: 1108 --------SLPRSPRSCFSR---PPPSPRLCP*SPPPYSSPPPRSPPSFSLLPP---SSP 971
Query: 528 FPLFPHSHKLLPLFPSPSPAVFFSPSC 554
P P + SP P++ F SC
Sbjct: 970 LPYSPSPRR------SPKPSL*FPLSC 908
>TC232032 weakly similar to UP|Q6K376 (Q6K376) ABC transporter-like protein,
partial (11%)
Length = 799
Score = 45.8 bits (107), Expect = 5e-05
Identities = 26/55 (47%), Positives = 38/55 (68%), Gaps = 1/55 (1%)
Frame = +2
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGR-KTGGYIDGDIKVSGYPKKQ 344
+L+G++G +PG L A+MG SG GK+TL+D LAGR + G+I ++G KKQ
Sbjct: 620 ILEGLTGYAKPGQLLAIMGPSGCGKSTLLDTLAGRLGSNTRQTGEILING--KKQ 778
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.331 0.145 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,662,567
Number of Sequences: 63676
Number of extensions: 716777
Number of successful extensions: 11944
Number of sequences better than 10.0: 774
Number of HSP's better than 10.0 without gapping: 8624
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 10154
length of query: 596
length of database: 12,639,632
effective HSP length: 103
effective length of query: 493
effective length of database: 6,081,004
effective search space: 2997934972
effective search space used: 2997934972
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC126018.1


