

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126016.2 - phase: 0
(327 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
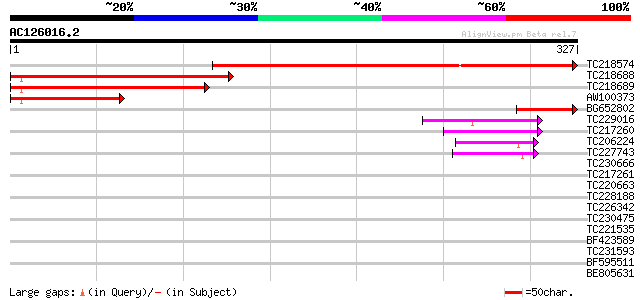
Sequences producing significant alignments: (bits) Value
TC218574 similar to PIR|S53400|S53400 RING finger protein YLR323... 401 e-112
TC218688 212 2e-55
TC218689 186 1e-47
AW100373 84 7e-17
BG652802 76 2e-14
TC229016 similar to UP|HES1_MOUSE (P35428) Transcription factor ... 46 3e-05
TC217260 42 4e-04
TC206224 similar to PIR|E86326|E86326 protein F18O14.3 [imported... 42 4e-04
TC227743 similar to PIR|E86326|E86326 protein F18O14.3 [imported... 41 6e-04
TC230666 similar to UP|O82767 (O82767) PRT1 protein, partial (11%) 40 0.001
TC217261 40 0.001
TC220663 similar to UP|Q9LG20 (Q9LG20) F14J16.17, partial (9%) 39 0.004
TC228188 similar to GB|AAP31929.1|30387527|BT006585 At3g58030 {A... 38 0.005
TC226342 homologue to UP|NPM2_MOUSE (Q80W85) Nucleoplasmin 2, pa... 38 0.007
TC230475 38 0.007
TC221535 weakly similar to GB|AAO39941.1|28372918|BT003713 At3g0... 37 0.009
BF423589 similar to PIR|T51853|T518 RING-H2 finger protein RHF1a... 37 0.012
TC231593 weakly similar to UP|Q86F30 (Q86F30) Clone ZZD1160 mRNA... 35 0.046
BF595511 35 0.060
BE805631 34 0.079
>TC218574 similar to PIR|S53400|S53400 RING finger protein YLR323c - yeast
(Saccharomyces cerevisiae) {Saccharomyces cerevisiae;} ,
partial (14%)
Length = 919
Score = 401 bits (1030), Expect = e-112
Identities = 191/211 (90%), Positives = 201/211 (94%), Gaps = 1/211 (0%)
Frame = +3
Query: 118 ARAIRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPL 177
+RAIRERALKQA ESLKGKS SS++ KLYKG+N+Y D+KAGFRREQTIASEKAGGSHGPL
Sbjct: 6 SRAIRERALKQAEESLKGKSPSSKNEKLYKGMNSYKDYKAGFRREQTIASEKAGGSHGPL 185
Query: 178 RASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKAR 237
RASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEW+EAEKAR
Sbjct: 186 RASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWEEAEKAR 365
Query: 238 KMRLATGEDA-EEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHH 296
KMRLA GEDA EEEGA+L DEDD ED+LPFACFICRN FVDPV TKCKHYFCEHCALKHH
Sbjct: 366 KMRLAAGEDADEEEGANLTDEDD-EDSLPFACFICRNSFVDPVVTKCKHYFCEHCALKHH 542
Query: 297 AKNKKCFVCNQPTLGIFNVAHEIRKKMAEDK 327
AKNKKCFVCNQPTLGIFNVAHEIR+KMAEDK
Sbjct: 543 AKNKKCFVCNQPTLGIFNVAHEIRRKMAEDK 635
>TC218688
Length = 488
Score = 212 bits (539), Expect = 2e-55
Identities = 108/131 (82%), Positives = 118/131 (89%), Gaps = 2/131 (1%)
Frame = +3
Query: 1 MEDSE--AKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNT 58
MEDS+ AKS ENQQTEQVCSFFRKPVN+KN+RKRTI NEDNE DSNNE SL+H+QKK
Sbjct: 96 MEDSDQPAKSAENQQTEQVCSFFRKPVNKKNIRKRTIVNEDNEEDSNNETSLLHIQKKTL 275
Query: 59 KADNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDA 118
K DNKL+FSTGSSKSSASA+P+EE KP F FESSKEIQVQHDSKATATLETET+FS+DA
Sbjct: 276 KPDNKLYFSTGSSKSSASAEPSEEPGKPVFQFESSKEIQVQHDSKATATLETETEFSKDA 455
Query: 119 RAIRERALKQA 129
RAIRERALKQA
Sbjct: 456 RAIRERALKQA 488
>TC218689
Length = 547
Score = 186 bits (472), Expect = 1e-47
Identities = 95/117 (81%), Positives = 104/117 (88%), Gaps = 2/117 (1%)
Frame = +2
Query: 1 MEDSE--AKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNT 58
MEDS+ AKS ENQQTEQVCSFFRKPVN+KN+RKRTIE EDNE DSNNE SL+H+QKK
Sbjct: 197 MEDSDQPAKSAENQQTEQVCSFFRKPVNKKNLRKRTIEIEDNEEDSNNETSLLHIQKKTL 376
Query: 59 KADNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFS 115
KADNKL+FSTGSSKSSASA+P+EES K F FESSKEIQVQHDSKATA LETET+FS
Sbjct: 377 KADNKLYFSTGSSKSSASAEPSEESGKTVFQFESSKEIQVQHDSKATAILETETEFS 547
>AW100373
Length = 426
Score = 84.3 bits (207), Expect = 7e-17
Identities = 45/69 (65%), Positives = 52/69 (75%), Gaps = 3/69 (4%)
Frame = +3
Query: 1 MEDSE--AKSTENQQTEQVCSFFRKPVNRK-NMRKRTIENEDNENDSNNEESLMHVQKKN 57
MEDS+ AKS ENQQTEQVCSFFRKP N+K N T+ NED E DSN+ SL+H+QKK
Sbjct: 219 MEDSDQPAKSAENQQTEQVCSFFRKPGNKKEN*GNPTLVNEDIEEDSNHGTSLLHIQKKT 398
Query: 58 TKADNKLFF 66
K DNKL+F
Sbjct: 399 LKPDNKLYF 425
>BG652802
Length = 402
Score = 75.9 bits (185), Expect = 2e-14
Identities = 33/35 (94%), Positives = 35/35 (99%)
Frame = +1
Query: 293 LKHHAKNKKCFVCNQPTLGIFNVAHEIRKKMAEDK 327
L+HHAKNKKCFVCNQPTLGIFNVAHEIR+KMAEDK
Sbjct: 118 LQHHAKNKKCFVCNQPTLGIFNVAHEIRRKMAEDK 222
>TC229016 similar to UP|HES1_MOUSE (P35428) Transcription factor HES-1 (Hairy
and enhancer of split 1), partial (6%)
Length = 769
Score = 45.8 bits (107), Expect = 3e-05
Identities = 23/73 (31%), Positives = 33/73 (44%), Gaps = 4/73 (5%)
Frame = +1
Query: 239 MRLATGEDAEEEGASLNDEDDDEDALP----FACFICRNPFVDPVSTKCKHYFCEHCALK 294
+ LA + E A E E P F C IC +P V+ +ST+C H FC++C
Sbjct: 316 VNLANNSSSASENAKKTPEPPKEPEAPKEPVFNCPICMSPLVEEMSTRCGHIFCKNCIRA 495
Query: 295 HHAKNKKCFVCNQ 307
+ KC C +
Sbjct: 496 AISAQAKCPTCRK 534
>TC217260
Length = 1114
Score = 42.0 bits (97), Expect = 4e-04
Identities = 19/57 (33%), Positives = 26/57 (45%)
Frame = +3
Query: 251 GASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
G S+ + L F C IC P V +ST+C H FC+ C + KC C +
Sbjct: 456 GGSVKKTPEPPKELVFNCPICMGPMVHEMSTRCGHIFCKDCIKAAISAQGKCPTCRK 626
>TC206224 similar to PIR|E86326|E86326 protein F18O14.3 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(43%)
Length = 1080
Score = 42.0 bits (97), Expect = 4e-04
Identities = 21/51 (41%), Positives = 27/51 (52%), Gaps = 3/51 (5%)
Frame = +1
Query: 258 DDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCA---LKHHAKNKKCFVC 305
+ DA F C IC + DPV T C H FC C L HH+ +++C VC
Sbjct: 286 NSSNDAGDFECNICFDLAQDPVITLCGHLFCWPCLYRWLHHHSHSQECPVC 438
>TC227743 similar to PIR|E86326|E86326 protein F18O14.3 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(73%)
Length = 1212
Score = 41.2 bits (95), Expect = 6e-04
Identities = 19/53 (35%), Positives = 29/53 (53%), Gaps = 3/53 (5%)
Frame = +2
Query: 256 DEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALK---HHAKNKKCFVC 305
+ +++ DA F C IC DP+ T C H FC C K H+++++C VC
Sbjct: 182 NNNNNNDAANFECNICFELAQDPIITLCGHLFCWPCLYKWLHFHSQSRECPVC 340
>TC230666 similar to UP|O82767 (O82767) PRT1 protein, partial (11%)
Length = 1102
Score = 40.4 bits (93), Expect = 0.001
Identities = 29/161 (18%), Positives = 58/161 (36%), Gaps = 12/161 (7%)
Frame = +2
Query: 160 RREQTIASEKAGGSHGPLRASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMH---D 216
R QT+ EK G + P + PD C+ + G+ G S ++ +
Sbjct: 116 RESQTLEEEKKSGFYSP-------------QFDPDTCESQAKFGHSGIPSSSSILNLTSN 256
Query: 217 RGDYKSGWQMEKEWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALPF---------A 267
+ + +E+ A + + + + + + G + ++ L
Sbjct: 257 SCNVGTSECLEQSGSTANEGDEGTIFSNREPDIIGTPAKGKKMPQEELSVQRKLSVADVT 436
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQP 308
C +C+ PV C H +C+ C + + +C VC P
Sbjct: 437 CTMCKQLLFHPVVLNCGHVYCQTCVINIDDEMLRCKVCQSP 559
>TC217261
Length = 1186
Score = 40.0 bits (92), Expect = 0.001
Identities = 22/69 (31%), Positives = 30/69 (42%)
Frame = +3
Query: 239 MRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAK 298
+ L G + E+ E E L F C IC P V +ST+C H FC+ C +
Sbjct: 447 INLEGGSSSMEQSFKKPPEPPKE--LVFNCPICMGPMVHEMSTRCGHIFCKDCIKAAISA 620
Query: 299 NKKCFVCNQ 307
KC C +
Sbjct: 621 QGKCPTCRK 647
>TC220663 similar to UP|Q9LG20 (Q9LG20) F14J16.17, partial (9%)
Length = 749
Score = 38.5 bits (88), Expect = 0.004
Identities = 27/107 (25%), Positives = 43/107 (39%), Gaps = 7/107 (6%)
Frame = +3
Query: 208 GDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDAEEEG-------ASLNDEDDD 260
G S F H + +Y++G + + +G +EE G LN
Sbjct: 300 GGSESFSHSQSNYQAG----------SSSANLSSVSGRHSEESGMWECTMCTLLNKR--- 440
Query: 261 EDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
L C +C STKC + C+ C L+++ K +KC C+Q
Sbjct: 441 ---LAPICELCGTQQPKDFSTKCNTWSCKFCTLENNVKLEKCSACDQ 572
>TC228188 similar to GB|AAP31929.1|30387527|BT006585 At3g58030 {Arabidopsis
thaliana;} , partial (27%)
Length = 877
Score = 38.1 bits (87), Expect = 0.005
Identities = 26/82 (31%), Positives = 34/82 (40%), Gaps = 3/82 (3%)
Frame = +1
Query: 227 EKEWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHY 286
E+ +E KA ED + ++ D F C IC + DPV T C H
Sbjct: 118 EERMEEVPKACDNINGVAEDETSQKKEDIEKGSGNDGDFFDCNICLDLARDPVVTCCGHL 297
Query: 287 FCEHCA---LKHHAKNKKCFVC 305
FC C L H+ K+C VC
Sbjct: 298 FCWPCLYRWLHLHSDAKECPVC 363
>TC226342 homologue to UP|NPM2_MOUSE (Q80W85) Nucleoplasmin 2, partial (8%)
Length = 1326
Score = 37.7 bits (86), Expect = 0.007
Identities = 13/38 (34%), Positives = 19/38 (49%)
Frame = +3
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVC 305
C IC+ P+ CKH FCE C + + + C +C
Sbjct: 828 CAICQEKMQAPILLSCKHMFCEECVSEWFERERTCPLC 941
>TC230475
Length = 528
Score = 37.7 bits (86), Expect = 0.007
Identities = 13/38 (34%), Positives = 20/38 (52%)
Frame = +3
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVC 305
C IC+ P+ +CKH FCE C + + + C +C
Sbjct: 297 CAICQEKMHAPILLRCKHIFCEDCVSEWFERERTCPLC 410
>TC221535 weakly similar to GB|AAO39941.1|28372918|BT003713 At3g07200
{Arabidopsis thaliana;} , partial (32%)
Length = 658
Score = 37.4 bits (85), Expect = 0.009
Identities = 15/44 (34%), Positives = 22/44 (49%)
Frame = +2
Query: 266 FACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQPT 309
F C IC + + ST+C H FC++C + KC C + T
Sbjct: 173 FNCPICMSALEEETSTRCAHIFCKNCIRAALSAQAKCPTCRKVT 304
>BF423589 similar to PIR|T51853|T518 RING-H2 finger protein RHF1a [imported]
- Arabidopsis thaliana (fragment), partial (21%)
Length = 413
Score = 37.0 bits (84), Expect = 0.012
Identities = 21/64 (32%), Positives = 33/64 (50%), Gaps = 3/64 (4%)
Frame = +3
Query: 247 AEEEGASLNDEDDDEDALPFACFICRNPFV--DPVS-TKCKHYFCEHCALKHHAKNKKCF 303
A A+ +D +D +C IC PF DP + T CKH + HC ++ ++K+C
Sbjct: 132 APSSSAAFDDSSED------SCSICLEPFSVHDPSTVTCCKHEYHLHCIIEWSQRSKECP 293
Query: 304 VCNQ 307
+C Q
Sbjct: 294 ICWQ 305
>TC231593 weakly similar to UP|Q86F30 (Q86F30) Clone ZZD1160 mRNA sequence,
partial (23%)
Length = 770
Score = 35.0 bits (79), Expect = 0.046
Identities = 11/21 (52%), Positives = 16/21 (75%)
Frame = +3
Query: 195 ICKDYKETGYCGYGDSCKFMH 215
+C + TG+C YGDSCK++H
Sbjct: 171 LCFHFVNTGFCRYGDSCKYLH 233
>BF595511
Length = 420
Score = 34.7 bits (78), Expect = 0.060
Identities = 20/63 (31%), Positives = 34/63 (53%), Gaps = 4/63 (6%)
Frame = +1
Query: 248 EEEGASLND---EDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALK-HHAKNKKCF 303
+ EG+S+ + E+ +E C IC++ + V TKC H FC C K ++++KC
Sbjct: 25 QNEGSSVTEKLQEELEEYRDIIKCSICQDRAKEVVITKCYHLFCYSCIQKVAGSRHRKCP 204
Query: 304 VCN 306
C+
Sbjct: 205 QCS 213
>BE805631
Length = 196
Score = 34.3 bits (77), Expect = 0.079
Identities = 12/31 (38%), Positives = 20/31 (63%)
Frame = +2
Query: 191 YQPDICKDYKETGYCGYGDSCKFMHDRGDYK 221
Y+ ++CK+++ G C +GD C F H R + K
Sbjct: 2 YKTEMCKNWELRGACKFGDKCCFAHGRHELK 94
Score = 30.0 bits (66), Expect = 1.5
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = +2
Query: 191 YQPDICKDYKETGYCGYGDSCKFMH 215
Y+ C Y +GYC YG C+++H
Sbjct: 119 YKTKRC*QYHHSGYCPYGLRCQYLH 193
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.127 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,457,076
Number of Sequences: 63676
Number of extensions: 213483
Number of successful extensions: 1473
Number of sequences better than 10.0: 142
Number of HSP's better than 10.0 without gapping: 1354
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1454
length of query: 327
length of database: 12,639,632
effective HSP length: 98
effective length of query: 229
effective length of database: 6,399,384
effective search space: 1465458936
effective search space used: 1465458936
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126016.2


