

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.14 + phase: 0 /pseudo
(241 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
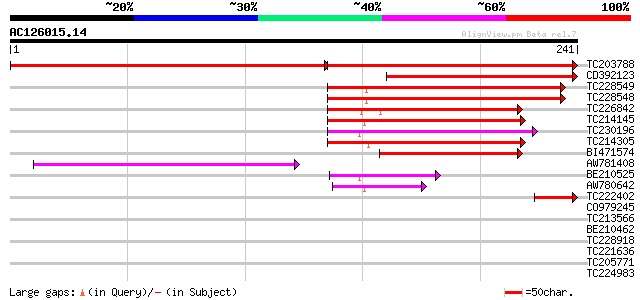
Sequences producing significant alignments: (bits) Value
TC203788 similar to GB|AAL91270.1|19699320|AY090367 AT3g12090/T2... 213 5e-97
CD392123 142 2e-34
TC228549 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {A... 115 2e-26
TC228548 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {A... 114 4e-26
TC226842 weakly similar to UP|Q84VG2 (Q84VG2) Senescence-associa... 100 6e-22
TC214145 similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated pro... 96 2e-20
TC230196 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associa... 92 2e-19
TC214305 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associa... 92 2e-19
BI471574 77 7e-15
AW781408 similar to GP|19699320|gb| AT3g12090/T21B14_110 {Arabid... 70 9e-13
BE210525 52 3e-07
AW780642 49 3e-06
TC222402 42 2e-04
CO979245 33 0.090
TC213566 weakly similar to UP|Q79FY9 (Q79FY9) PE-PGRS FAMILY PRO... 31 0.58
BE210462 similar to GP|4102692|gb|A late-embryogenesis abundant ... 31 0.58
TC228918 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (4%) 31 0.58
TC221636 similar to UP|Q79G09 (Q79G09) PE-PGRS FAMILY PROTEIN, p... 31 0.58
TC205771 similar to UP|Q9JM93 (Q9JM93) SRp25 nuclear protein (AD... 30 0.76
TC224983 similar to UP|Q8LUS1 (Q8LUS1) NADH dehydrogenase 1 (Fra... 30 1.3
>TC203788 similar to GB|AAL91270.1|19699320|AY090367 AT3g12090/T21B14_110
{Arabidopsis thaliana;} , partial (84%)
Length = 1321
Score = 213 bits (541), Expect(2) = 5e-97
Identities = 92/106 (86%), Positives = 97/106 (90%)
Frame = +1
Query: 136 TGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVL 195
+GCCKPPTAC YN+ MMTQD DCY+W+N P LLCYECDSCKAGVLEDIR NWHKLSVL
Sbjct: 655 SGCCKPPTACTYNVATTMMTQDPDCYRWNNAPNLLCYECDSCKAGVLEDIRGNWHKLSVL 834
Query: 196 TVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRPRWDYH 241
TVTML+LLIGIYSIGCCAFRN RRAETDYPYGENRMTKVRPRWDYH
Sbjct: 835 TVTMLVLLIGIYSIGCCAFRNTRRAETDYPYGENRMTKVRPRWDYH 972
Score = 159 bits (402), Expect(2) = 5e-97
Identities = 83/136 (61%), Positives = 97/136 (71%)
Frame = +3
Query: 1 MDGKEQYNLCKFSPNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRF 60
MDGKEQ+NL KFSPN + R C C + S HWC+ SC MCTMVVLG +V+ +SST F
Sbjct: 246 MDGKEQHNL*KFSPNPSFGYRICCACGVTSRLHWCMLSCGMCTMVVLGGHVVPHSSTDGF 425
Query: 61 DYIWFWCD**RWWCRSAW*IL**VSSYRLFTLVEEKNTRSSLLEYYKELYFRV*NL*QTC 120
DYIWFW D* W C SAW L VSS L T+VEE+N+ SSLLE+Y LYF V NL*+ C
Sbjct: 426 DYIWFWGD*QGWRCGSAWKGLQGVSSSGLLTMVEEENSGSSLLEHY*RLYFGVQNL*KAC 605
Query: 121 LLDPFGLHAE*YVSNT 136
LLDP L+A+ YVSNT
Sbjct: 606 LLDPS*LYAKGYVSNT 653
>CD392123
Length = 601
Score = 142 bits (357), Expect = 2e-34
Identities = 66/81 (81%), Positives = 68/81 (83%)
Frame = -2
Query: 161 YKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVLTVTMLILLIGIYSIGCCAFRNARRA 220
+ W P DSCKAGVLEDIR NWHKLSVLTVTML+LLIGIYSIGCCAFRN RRA
Sbjct: 423 FGWIKTPRAEFGTRDSCKAGVLEDIRGNWHKLSVLTVTMLVLLIGIYSIGCCAFRNTRRA 244
Query: 221 ETDYPYGENRMTKVRPRWDYH 241
ETDYPYGENRMTKVRPRWDYH
Sbjct: 243 ETDYPYGENRMTKVRPRWDYH 181
>TC228549 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Arabidopsis
thaliana;} , complete
Length = 1010
Score = 115 bits (287), Expect = 2e-26
Identities = 49/109 (44%), Positives = 73/109 (66%), Gaps = 8/109 (7%)
Frame = +2
Query: 136 TGCCKPPTACNYNME--------AVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKPPT C Y + + +M ++DC +WSN+ LCY CDSCKAGVL +++
Sbjct: 524 SGCCKPPTDCGYVYQNETVWIPGSGLMGTNADCTRWSNDQEQLCYACDSCKAGVLASLKK 703
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRP 236
+W K+SV+ + ++I+L+ +Y I A+RN R+ + D PYGE RMTK +P
Sbjct: 704 SWRKVSVINIVVMIILVIVYIIAYAAYRNNRKMDNDEPYGEARMTKAQP 850
>TC228548 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Arabidopsis
thaliana;} , partial (46%)
Length = 690
Score = 114 bits (285), Expect = 4e-26
Identities = 49/109 (44%), Positives = 72/109 (65%), Gaps = 8/109 (7%)
Frame = +1
Query: 136 TGCCKPPTACNYNME--------AVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKPPT C Y + + +M + DC +WSN+ LCY CDSCKAGVL +++
Sbjct: 49 SGCCKPPTDCGYVYQNETVWIPGSGLMGANPDCTRWSNDQEQLCYACDSCKAGVLASLKK 228
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRP 236
+W K+SV+ + ++I+L+ +Y I A+RN R+ + D PYGE RMTK +P
Sbjct: 229 SWRKVSVINIVVMIILVIVYIIAYAAYRNNRKMDNDEPYGEARMTKAQP 375
>TC226842 weakly similar to UP|Q84VG2 (Q84VG2) Senescence-associated
protein-like protein, partial (30%)
Length = 1116
Score = 100 bits (249), Expect = 6e-22
Identities = 41/90 (45%), Positives = 62/90 (68%), Gaps = 7/90 (7%)
Frame = +1
Query: 136 TGCCKPPTACNYN-MEAVMMTQ------DSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPPTAC YN + ++ T DSDCY W+N+ LCY C++CKAG+L ++R+
Sbjct: 511 SGCCKPPTACGYNYVNPILWTNPVNPMADSDCYLWNNDQNQLCYNCNACKAGLLGNLRKE 690
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNAR 218
W K +++ + +++LI +Y I C AFRNA+
Sbjct: 691 WRKANIILIVAVVVLIWVYVIACSAFRNAQ 780
>TC214145 similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated protein-like,
partial (87%)
Length = 1867
Score = 95.5 bits (236), Expect = 2e-20
Identities = 39/93 (41%), Positives = 60/93 (63%), Gaps = 9/93 (9%)
Frame = +1
Query: 136 TGCCKPPTACNYNM---------EAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIR 186
+GCCKP C ++ V + DC W+N+PT+LC+ C SCKAG+L++++
Sbjct: 1366 SGCCKPAEECQFSYVNPTTWTKPTNVTNQSNPDCDAWNNDPTVLCFNCQSCKAGLLQNLK 1545
Query: 187 RNWHKLSVLTVTMLILLIGIYSIGCCAFRNARR 219
+W +++V+ + L+ LI +YSIGCCAFRN RR
Sbjct: 1546 TDWKRVAVVNIVFLVFLIIVYSIGCCAFRNNRR 1644
>TC230196 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated
protein-like, partial (37%)
Length = 662
Score = 92.4 bits (228), Expect = 2e-19
Identities = 38/97 (39%), Positives = 58/97 (59%), Gaps = 8/97 (8%)
Frame = +2
Query: 136 TGCCKPPTACNY--------NMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKP CN+ N + + DC W N+ +LC+ C+SCKAG L++++
Sbjct: 101 SGCCKPSNDCNFAYQGPSVWNKTDGVNHSNPDCNAWDNDSNVLCFNCESCKAGFLQNLKT 280
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDY 224
+W K++++ V L+ LI +YS+GCCAFRN R Y
Sbjct: 281 DWKKVTIVNVIFLVFLIIVYSVGCCAFRNNMRGNWKY 391
>TC214305 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated
protein-like, partial (89%)
Length = 1003
Score = 92.0 bits (227), Expect = 2e-19
Identities = 40/93 (43%), Positives = 56/93 (60%), Gaps = 9/93 (9%)
Frame = +1
Query: 136 TGCCKPPTACNYNMEA---------VMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIR 186
+GCCKP C + E V + DC W+N T+LC+ C SCKAG L++ +
Sbjct: 574 SGCCKPAEECLFTYENSTSWTKPGNVTSYNNPDCDAWNNNQTVLCFNCQSCKAGFLQNFK 753
Query: 187 RNWHKLSVLTVTMLILLIGIYSIGCCAFRNARR 219
W +++V+ + L+LLI +YSIGCCAFRN RR
Sbjct: 754 TEWKRVAVVNIVFLVLLIIVYSIGCCAFRNNRR 852
>BI471574
Length = 422
Score = 77.0 bits (188), Expect = 7e-15
Identities = 28/61 (45%), Positives = 45/61 (72%)
Frame = +1
Query: 158 SDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVLTVTMLILLIGIYSIGCCAFRNA 217
SDCY W+N+ LCY C++CKAG+L ++R+ W K +++ + +++LI +Y I C AFRNA
Sbjct: 217 SDCYLWNNDQNQLCYNCNACKAGLLGNLRKEWRKANIILIVAVVVLIWVYVIACSAFRNA 396
Query: 218 R 218
+
Sbjct: 397 Q 399
>AW781408 similar to GP|19699320|gb| AT3g12090/T21B14_110 {Arabidopsis
thaliana}, partial (23%)
Length = 367
Score = 70.1 bits (170), Expect = 9e-13
Identities = 46/113 (40%), Positives = 59/113 (51%)
Frame = +1
Query: 11 KFSPNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRFDYIWFWCD** 70
KF PN T +R C CD S FH +FSC M + VLGD+ + S + W CD
Sbjct: 1 KFLPNPTFGDRLCCACDFASGFHRGMFSCGMGIVGVLGDHGVHYSCPLMVCRFWLCCDGT 180
Query: 71 RWWCRSAW*IL**VSSYRLFTLVEEKNTRSSLLEYYKELYFRV*NL*QTCLLD 123
W SA * V S LF + EE + SS+LE + EL+ V*+L + C LD
Sbjct: 181 GWGSGSAC*GFQIVLSRPLFAMAEE*D*GSSVLELHYELHMGV*HLCKACRLD 339
>BE210525
Length = 381
Score = 51.6 bits (122), Expect = 3e-07
Identities = 20/55 (36%), Positives = 33/55 (59%), Gaps = 8/55 (14%)
Frame = +1
Query: 137 GCCKPPTACNY--------NMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLE 183
GCC+PP+ C Y ++ ++ ++DC ++ N + CY+CDSCKAGV +
Sbjct: 208 GCCRPPSQCGYPAVNASYYDLTFHPVSPNNDCKRYKNSRAIKCYDCDSCKAGVAQ 372
>AW780642
Length = 142
Score = 48.5 bits (114), Expect = 3e-06
Identities = 22/47 (46%), Positives = 25/47 (52%), Gaps = 7/47 (14%)
Frame = +2
Query: 138 CCKPPTACNYNM-------EAVMMTQDSDCYKWSNEPTLLCYECDSC 177
CCKPPTAC Y V+ DS CY WSN+ LCY D+C
Sbjct: 2 CCKPPTACGYYYVNPILWTNPVIPMADSYCYLWSNDLNHLCYHSDAC 142
>TC222402
Length = 288
Score = 42.4 bits (98), Expect = 2e-04
Identities = 16/18 (88%), Positives = 17/18 (93%)
Frame = +1
Query: 224 YPYGENRMTKVRPRWDYH 241
+ YGENRMTKVRPRWDYH
Sbjct: 4 HEYGENRMTKVRPRWDYH 57
>CO979245
Length = 730
Score = 33.5 bits (75), Expect = 0.090
Identities = 16/57 (28%), Positives = 19/57 (33%)
Frame = -3
Query: 22 FYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRFDYIWFWCD**RWWCRSAW 78
F+C C WF WC CC + WFWC WW +W
Sbjct: 239 FWCCCHPWWWFCWC---CC--------------------PWCWFWCCFFYWWICRSW 138
>TC213566 weakly similar to UP|Q79FY9 (Q79FY9) PE-PGRS FAMILY PROTEIN,
partial (4%)
Length = 676
Score = 30.8 bits (68), Expect = 0.58
Identities = 19/63 (30%), Positives = 22/63 (34%), Gaps = 2/63 (3%)
Frame = +3
Query: 19 CNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRFDYIW--FWCD**RWWCRS 76
C F C C + WC F C C F + W FW +WWC S
Sbjct: 357 CFLFVCRC-----WCWCFFGWCWC-----------------FGWCWGYFWSR--QWWCIS 464
Query: 77 AW* 79
W*
Sbjct: 465 WW* 473
>BE210462 similar to GP|4102692|gb|A late-embryogenesis abundant protein
{Glycine max}, partial (70%)
Length = 310
Score = 30.8 bits (68), Expect = 0.58
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = -1
Query: 19 CNRFYCTCDIISWFHWCLFSCCMCTMVVL 47
C+RF+C C +I +H LF C C + +L
Sbjct: 85 CSRFFCFCVLIG-YHLLLFCSCCCFLCLL 2
>TC228918 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (4%)
Length = 1181
Score = 30.8 bits (68), Expect = 0.58
Identities = 18/47 (38%), Positives = 20/47 (42%)
Frame = -3
Query: 2 DGKEQYNLCKFSPNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLG 48
+GK Y C SP T C C C + F C SCC C LG
Sbjct: 786 EGKLTYFRCCISP-TLFCCPSICACCMFVVFG*CPISCCCCCEGKLG 649
>TC221636 similar to UP|Q79G09 (Q79G09) PE-PGRS FAMILY PROTEIN, partial (5%)
Length = 756
Score = 30.8 bits (68), Expect = 0.58
Identities = 17/60 (28%), Positives = 22/60 (36%), Gaps = 2/60 (3%)
Frame = +1
Query: 22 FYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRFDYIW--FWCD**RWWCRSAW* 79
F+C + + WC F C C F + W FW +WWC S W*
Sbjct: 571 FWCFLFVCRCWCWCFFGWCWC-----------------FGWCWGYFWSR--QWWCISWW* 693
>TC205771 similar to UP|Q9JM93 (Q9JM93) SRp25 nuclear protein
(ADP-ribosylation factor-like 6 interacting protein 4),
partial (6%)
Length = 490
Score = 30.4 bits (67), Expect = 0.76
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = +2
Query: 8 NLCKFSPNTTSCNRFYCTCDIISWFHWCLFS 38
N+ +F + S +C +SWFHWCL S
Sbjct: 134 NISQFLTWSCSTTTSSFSCCFLSWFHWCLAS 226
>TC224983 similar to UP|Q8LUS1 (Q8LUS1) NADH dehydrogenase 1 (Fragment),
partial (37%)
Length = 1412
Score = 29.6 bits (65), Expect = 1.3
Identities = 13/40 (32%), Positives = 20/40 (49%), Gaps = 2/40 (5%)
Frame = +3
Query: 30 SWF--HWCLFSCCMCTMVVLGDYVIANSSTIRFDYIWFWC 67
SWF +CLF + +VV G + A + ++ FWC
Sbjct: 1026 SWFANKFCLFKVMILALVVHGSHTCALPFILEVPFLCFWC 1145
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.343 0.149 0.575
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,138,792
Number of Sequences: 63676
Number of extensions: 300652
Number of successful extensions: 4568
Number of sequences better than 10.0: 70
Number of HSP's better than 10.0 without gapping: 4444
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4548
length of query: 241
length of database: 12,639,632
effective HSP length: 94
effective length of query: 147
effective length of database: 6,654,088
effective search space: 978150936
effective search space used: 978150936
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC126015.14


