

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.12 - phase: 0
(1620 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
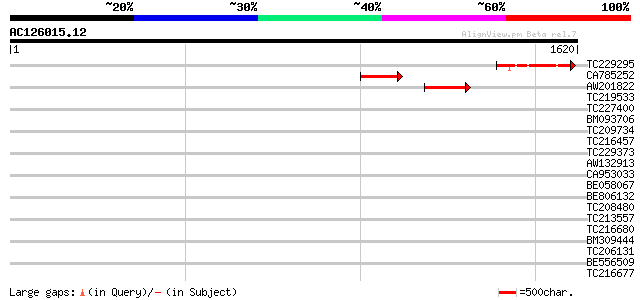
Score E
Sequences producing significant alignments: (bits) Value
TC229295 335 7e-92
CA785252 157 3e-38
AW201822 137 4e-32
TC219533 homologue to UP|Q8L5P7 (Q8L5P7) LHY protein, partial (10%) 41 0.004
TC227400 similar to UP|Q6R0G3 (Q6R0G3) MYB transcription factor,... 41 0.004
BM093706 39 0.014
TC209734 similar to GB|AAS58514.1|45357110|AY550303 MYB transcri... 39 0.024
TC216457 similar to GB|AAS58510.1|45357102|AY550299 MYB transcri... 38 0.041
TC229373 similar to UP|Q6QAC9 (Q6QAC9) MYB transcription factor,... 38 0.041
AW132913 38 0.041
CA953033 38 0.041
BE058067 similar to PIR|D86350|D863 F8K7.13 protein - Arabidopsi... 37 0.070
BE806132 similar to PIR|D86350|D863 F8K7.13 protein - Arabidopsi... 36 0.12
TC208480 36 0.16
TC213557 similar to UP|Q9LSZ3 (Q9LSZ3) Arabidopsis thaliana geno... 36 0.16
TC216680 homologue to GB|AAS58514.1|45357110|AY550303 MYB transc... 35 0.35
BM309444 35 0.35
TC206131 similar to UP|Q9LYF9 (Q9LYF9) Clathrin binding protein-... 34 0.46
BE556509 34 0.60
TC216677 similar to GB|AAS58514.1|45357110|AY550303 MYB transcri... 33 0.78
>TC229295
Length = 909
Score = 335 bits (860), Expect = 7e-92
Identities = 170/232 (73%), Positives = 195/232 (83%), Gaps = 6/232 (2%)
Frame = +1
Query: 1392 EENGSHHPKLNNKSSNLNFTGHQNSDENLNFLKF------GLENVPVMSYGYWEGNAIQS 1445
EENG+HHPKL++KSSN TGH ++D NL LKF GLENVP+ SYGYW+GN IQ+
Sbjct: 1 EENGTHHPKLSSKSSNPKITGHHSADGNLKILKFDHNDYVGLENVPMRSYGYWDGNRIQT 180
Query: 1446 RQSGLSSLPDSSFLLAKYPAAFSNYPTSSSNLEQQPPLQAFAKNSQRHLTGASTFTARDV 1505
GLS+LPDS+ LLAKYPAAFSNY TSS+ LEQ P LQ ++KN++R L GASTFT RD+
Sbjct: 181 ---GLSTLPDSAILLAKYPAAFSNYLTSSAKLEQ-PSLQTYSKNNERLLNGASTFTTRDI 348
Query: 1506 NGSNAMLDYQMFRGRDGPQVQPFMVDVQHRQDLFSEMQRRHSFEAISSLQQQGRGMMGMN 1565
NGSNA++DYQMFR RDGP+VQPFMVDV+H QD+FSEMQRR+ FEAISSLQQQ RGM N
Sbjct: 349 NGSNALIDYQMFR-RDGPKVQPFMVDVKHCQDVFSEMQRRNGFEAISSLQQQSRGM---N 516
Query: 1566 SVGRPGILVGGSCSGVSDPVAAIKMHYSNSEKYGGQNGSVVRDDESWGGKGD 1617
VGRPGILVGGSCSGVSDPVAAIKMHYSNS+KYGGQ GS+ R+DESWGGKGD
Sbjct: 517 GVGRPGILVGGSCSGVSDPVAAIKMHYSNSDKYGGQTGSIAREDESWGGKGD 672
>CA785252
Length = 434
Score = 157 bits (398), Expect = 3e-38
Identities = 80/119 (67%), Positives = 95/119 (79%)
Frame = -1
Query: 1003 PVPGINGSPLNDDANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDALNTFHDESNP 1062
P+P GSP+NDDANGGESDTDDACVVE GSVV DKSG KTDEDLP NT+HDES+P
Sbjct: 434 PIPENVGSPVNDDANGGESDTDDACVVETGSVVGTDKSGTKTDEDLPLYGTNTYHDESHP 255
Query: 1063 LEATSLSAKLNESREISGTEVCLENVDVASVACAINVESKLGSDVSGVGLCTTDKSGSV 1121
+EA +LSA+LNES+EI GTEV LE+ +V S A IN++S+LG D S V LC ++KSGSV
Sbjct: 254 VEARNLSAELNESKEIIGTEVDLEDANVTSGAYQINIDSELGCDGSEVFLCVSNKSGSV 78
>AW201822
Length = 404
Score = 137 bits (345), Expect = 4e-32
Identities = 75/136 (55%), Positives = 96/136 (70%), Gaps = 3/136 (2%)
Frame = +1
Query: 1185 EHVADAGVVVELKNCVLESSTAANVSFSPVVNSCSGLSFGSENKHVSFGKPHTSALSMSM 1244
+H AD+GV+V++K+ V + ST N S S + NSCSGLSF SENKHV G P SAL SM
Sbjct: 10 KHEADSGVIVDMKSSVHDLSTMINSSISSLGNSCSGLSFSSENKHVPLGNPRVSAL--SM 183
Query: 1245 SDLQATANSLLLKAAAAQCEKTVSQDRLSSTCDIQGGRDMRCHSSGSNGDHQLPLSGS-- 1302
+L A + + A QCEKT SQD++SSTCDI+GGRDM C +S SNGDHQ ++G+
Sbjct: 184 DNLHALLQNTV--AVDVQCEKTASQDQMSSTCDIRGGRDMHCQNSISNGDHQ-HITGNLS 354
Query: 1303 -HVETVSVLQGYSMQV 1317
HV+ VS+LQGY +QV
Sbjct: 355 DHVDAVSILQGYPLQV 402
>TC219533 homologue to UP|Q8L5P7 (Q8L5P7) LHY protein, partial (10%)
Length = 641
Score = 41.2 bits (95), Expect = 0.004
Identities = 19/56 (33%), Positives = 35/56 (61%), Gaps = 6/56 (10%)
Frame = +3
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKT--RKCLGLNL 1000
WT++E FL+A+ +G+ +++I +GTK + H ++FF+K R+C G N+
Sbjct: 312 WTEEEHNRFLEALKLYGRAWQRIEEHIGTKTAVQIRSHAQKFFTKLR*RECEGYNI 479
Score = 32.7 bits (73), Expect = 1.3
Identities = 19/65 (29%), Positives = 30/65 (45%)
Frame = +3
Query: 725 PLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSEC 784
P I K+R WT EE FLE +G+ +++I + KT ++H +
Sbjct: 282 PYTITKQRER---WTEEEHNRFLEALKLYGRAWQRIEEHIGTKTAVQ-----IRSHAQKF 437
Query: 785 FEKLK 789
F KL+
Sbjct: 438 FTKLR 452
>TC227400 similar to UP|Q6R0G3 (Q6R0G3) MYB transcription factor, partial (30%)
Length = 1488
Score = 41.2 bits (95), Expect = 0.004
Identities = 23/65 (35%), Positives = 34/65 (51%), Gaps = 4/65 (6%)
Frame = +1
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRKCLGLNLANPVPG 1006
WTD+E FL+A+ +G+ + +I VGTK + H ++FFSK + N N V
Sbjct: 232 WTDEEHKKFLEALKLYGRAWRRIEEHVGTKTAVQIRSHAQKFFSKLLRDPTGNNTNTVES 411
Query: 1007 INGSP 1011
I P
Sbjct: 412 IEIPP 426
Score = 37.4 bits (85), Expect = 0.054
Identities = 23/73 (31%), Positives = 33/73 (44%)
Frame = +1
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P I K+R WT EE + FLE +G+ +R+I + KT ++H
Sbjct: 193 VRKPYTITKQRER---WTDEEHKKFLEALKLYGRAWRRIEEHVGTKTAVQ-----IRSHA 348
Query: 782 SECFEKLKRKDIG 794
+ F KL R G
Sbjct: 349 QKFFSKLLRDPTG 387
>BM093706
Length = 421
Score = 39.3 bits (90), Expect = 0.014
Identities = 26/101 (25%), Positives = 47/101 (45%), Gaps = 4/101 (3%)
Frame = +3
Query: 895 EAMSSCITSSIDPVDGNKETKFLKANPLFKQPLTPDISQNADDETCSDESCGEATEWTDD 954
E+ S I S + N E + P ++P T I++ + +WT++
Sbjct: 78 ESTRSTIFGSASNIHSNAEKQAENVAPKVRKPYT--ITKQRE-------------KWTEE 212
Query: 955 ETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSK 991
E FL+A+ +G+ + +I +GTK + H ++FFSK
Sbjct: 213 EHQKFLEALKLYGRGWRQIEEHIGTKTAVQIRSHAQKFFSK 335
Score = 36.2 bits (82), Expect = 0.12
Identities = 21/70 (30%), Positives = 33/70 (47%)
Frame = +3
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P I K+R WT EE + FLE +G+ +R+I + KT ++H
Sbjct: 162 VRKPYTITKQREK---WTEEEHQKFLEALKLYGRGWRQIEEHIGTKTAVQ-----IRSHA 317
Query: 782 SECFEKLKRK 791
+ F K+ R+
Sbjct: 318 QKFFSKVVRE 347
>TC209734 similar to GB|AAS58514.1|45357110|AY550303 MYB transcription factor
{Arabidopsis thaliana;} , partial (47%)
Length = 1324
Score = 38.5 bits (88), Expect = 0.024
Identities = 21/70 (30%), Positives = 35/70 (50%)
Frame = +2
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
+ P I K R WT +E + FLE F +D++KI +F+ KT ++H
Sbjct: 299 IRKPYTITKSRES---WTEQEHDKFLEALQLFDRDWKKIEAFVGSKTV-----IQIRSHA 454
Query: 782 SECFEKLKRK 791
+ F K+++K
Sbjct: 455 QKYFLKVQKK 484
>TC216457 similar to GB|AAS58510.1|45357102|AY550299 MYB transcription factor
{Arabidopsis thaliana;} , partial (27%)
Length = 2351
Score = 37.7 bits (86), Expect = 0.041
Identities = 16/46 (34%), Positives = 29/46 (62%), Gaps = 4/46 (8%)
Frame = +1
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCVGTK----AQEHCKRFFSK 991
+WT++E FL+A+ +G+ + +I +GTK + H ++FFSK
Sbjct: 472 KWTEEEHQKFLEALKLYGRGWRQIEEHIGTKNAVQIRSHAQKFFSK 609
Score = 36.6 bits (83), Expect = 0.092
Identities = 23/78 (29%), Positives = 38/78 (48%)
Frame = +1
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P I K+R WT EE + FLE +G+ +R+I +H T + ++ ++H
Sbjct: 436 VRKPYTITKQREK---WTEEEHQKFLEALKLYGRGWRQIE---EHIGTKNAVQI--RSHA 591
Query: 782 SECFEKLKRKDIGKLGKS 799
+ F K+ R+ G S
Sbjct: 592 QKFFSKVVRESEGSAESS 645
>TC229373 similar to UP|Q6QAC9 (Q6QAC9) MYB transcription factor, partial
(70%)
Length = 1250
Score = 37.7 bits (86), Expect = 0.041
Identities = 22/69 (31%), Positives = 33/69 (46%)
Frame = +2
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P I K R WT EE + FLE F +D++KI F+ KT ++H
Sbjct: 257 VRKPYTITKSRES---WTEEEHDKFLEALQLFDRDWKKIEDFVGSKTV-----IQIRSHA 412
Query: 782 SECFEKLKR 790
+ F K+++
Sbjct: 413 QKYFLKVQK 439
>AW132913
Length = 400
Score = 37.7 bits (86), Expect = 0.041
Identities = 16/46 (34%), Positives = 29/46 (62%), Gaps = 4/46 (8%)
Frame = +3
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCVGTK----AQEHCKRFFSK 991
+WT++E FL+A+ +G+ + +I +GTK + H ++FFSK
Sbjct: 231 KWTEEEHQKFLEALKLYGRGWRQIEEHIGTKNAVQIRSHAQKFFSK 368
Score = 36.2 bits (82), Expect = 0.12
Identities = 22/73 (30%), Positives = 37/73 (50%)
Frame = +3
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P I K+R WT EE + FLE +G+ +R+I +H T + ++ ++H
Sbjct: 195 VRKPYTITKQREK---WTEEEHQKFLEALKLYGRGWRQIE---EHIGTKNAVQI--RSHA 350
Query: 782 SECFEKLKRKDIG 794
+ F K+ R+ G
Sbjct: 351 QKFFSKVVRESEG 389
>CA953033
Length = 421
Score = 37.7 bits (86), Expect = 0.041
Identities = 16/46 (34%), Positives = 29/46 (62%), Gaps = 4/46 (8%)
Frame = +3
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCVGTK----AQEHCKRFFSK 991
+WT++E FL+A+ +G+ + +I +GTK + H ++FFSK
Sbjct: 210 KWTEEEHQKFLEALKLYGRGWRQIEEHIGTKNAVQIRSHAQKFFSK 347
Score = 36.6 bits (83), Expect = 0.092
Identities = 23/78 (29%), Positives = 38/78 (48%)
Frame = +3
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P I K+R WT EE + FLE +G+ +R+I +H T + ++ ++H
Sbjct: 174 VRKPYTITKQREK---WTEEEHQKFLEALKLYGRGWRQIE---EHIGTKNAVQI--RSHA 329
Query: 782 SECFEKLKRKDIGKLGKS 799
+ F K+ R+ G S
Sbjct: 330 QKFFSKVVRESEGSAESS 383
>BE058067 similar to PIR|D86350|D863 F8K7.13 protein - Arabidopsis thaliana,
partial (11%)
Length = 423
Score = 37.0 bits (84), Expect = 0.070
Identities = 20/74 (27%), Positives = 38/74 (51%), Gaps = 2/74 (2%)
Frame = +3
Query: 947 EATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTRKCLGL--NLANPV 1004
+ WTD ET L+A+ + +++ +I+ VGTK++ C F + G N+ P
Sbjct: 15 DGDSWTDQETLLLLEAMEIYNENWNEIAEHVGTKSKAQCILHFLRLPMEDGKLENINVPS 194
Query: 1005 PGINGSPLNDDANG 1018
++ + +N D +G
Sbjct: 195 MSLSSNAINRDHSG 236
>BE806132 similar to PIR|D86350|D863 F8K7.13 protein - Arabidopsis thaliana,
partial (11%)
Length = 406
Score = 36.2 bits (82), Expect = 0.12
Identities = 14/39 (35%), Positives = 24/39 (60%)
Frame = +3
Query: 947 EATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHC 985
+ WTD ET L+AV + +++ +I+ VGTK++ C
Sbjct: 9 DGDSWTDQETLLLLEAVEVYNENWNEIAEHVGTKSKAQC 125
>TC208480
Length = 1576
Score = 35.8 bits (81), Expect = 0.16
Identities = 63/307 (20%), Positives = 125/307 (40%), Gaps = 3/307 (0%)
Frame = +3
Query: 282 DDKLLGKVGNADNDVSNLTDSPAPGFQNHLQKFYLNLDKLDVDSLNSLGSSIVELVQSDD 341
+DKLL KV + + ++NL Q H ++ + L S +S+++
Sbjct: 249 NDKLLQKVVSLEEVINNLQTDNELQTQKHTSL------EMRIAQLQSENNSLLQ------ 392
Query: 342 PSSDDSGLVRSNAINKLLIWKADISKVLEMTESEIDLLENELKSLKSESVDRSECPVASG 401
++GLV N+LL K +S E E +I+LLENEL S + V E +A
Sbjct: 393 ---KEAGLVEKT--NQLLNEKVVLSLKAESLERKINLLENELSSFSEKEVG-LETRIAQL 554
Query: 402 SQQADSSSKFYEERVEVSQKVIRPVPLKIISSD--EPNTVKMPQSTNLCSIHENDKEEDI 459
+ +S + VE + +++ + + + E + N EN +E I
Sbjct: 555 QSENNSLLQKEATLVERTNQLLNEKAVLSLKGESLEQKIYLLESDLNSLVKKENSTKETI 734
Query: 460 DS-PGSATSKFVEPLPVNAVSSSYTRGYDNLSRDMNAVQSTMMKCFVRCNRKNTSVSACN 518
+ G+ + + ++ L ++++QST+ N +N++ S+C
Sbjct: 735 SNLNGNIAVLQAQVEELEESRNNLFLENQQLREKVSSLQSTVQ------NHENSNTSSC- 893
Query: 519 NVNTPTEVKDSLGDVTFGANLCSSYGDTYKSIIASNKESANRAHKLFTKLVPKECKKHGN 578
+ VKD + + + + ++A N E + +L +L + + G+
Sbjct: 894 --SWDASVKDLASENEDLKSEIEAAFTLVEKLMAENAELVEKVTELCVELDHRSAEV-GH 1064
Query: 579 MGVSNDS 585
GV+ +
Sbjct: 1065SGVTESN 1085
>TC213557 similar to UP|Q9LSZ3 (Q9LSZ3) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MSD21, partial (24%)
Length = 746
Score = 35.8 bits (81), Expect = 0.16
Identities = 24/77 (31%), Positives = 39/77 (50%)
Frame = +2
Query: 1356 SSNSEKTSRNGDVKLFGKILTNPSSTQNPNLTAKRSEENGSHHPKLNNKSSNLNFTGHQN 1415
+S+ + + GD+K+ GK + Q T++ N H+PKL ++NF G+ N
Sbjct: 407 ASSPVNSPKVGDLKVSGK-----NDDQLNGFTSRGKGFNSQHYPKLQADECDINFNGN-N 568
Query: 1416 SDENLNFLKFGLENVPV 1432
+ N N K GL VP+
Sbjct: 569 VECNTNLKKKGLCLVPL 619
>TC216680 homologue to GB|AAS58514.1|45357110|AY550303 MYB transcription
factor {Arabidopsis thaliana;} , partial (29%)
Length = 726
Score = 34.7 bits (78), Expect = 0.35
Identities = 20/66 (30%), Positives = 32/66 (48%)
Frame = +2
Query: 725 PLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSEC 784
P I K R WT E + FLE F +D++KI +F+ KT ++H +
Sbjct: 188 PYTITKSRES---WTEPEHDKFLEALQLFDRDWKKIEAFVGSKTV-----IQIRSHAQKY 343
Query: 785 FEKLKR 790
F K+++
Sbjct: 344 FLKVQK 361
>BM309444
Length = 433
Score = 34.7 bits (78), Expect = 0.35
Identities = 14/50 (28%), Positives = 28/50 (56%)
Frame = +3
Query: 949 TEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTRKCLGL 998
++W+ +E F +A +GKD++K++ V ++ E + +S R L L
Sbjct: 234 SQWSKEELERFYEAYRKYGKDWKKVAAVVRNRSTEMVEALYSMNRAYLSL 383
>TC206131 similar to UP|Q9LYF9 (Q9LYF9) Clathrin binding protein-like,
partial (52%)
Length = 2220
Score = 34.3 bits (77), Expect = 0.46
Identities = 35/128 (27%), Positives = 52/128 (40%), Gaps = 8/128 (6%)
Frame = +3
Query: 140 RSGSLSSRGSGFSHSSSSR-----SMAG---TDSYEGKPNLKHKNVTAVESNSGEATACV 191
+SGS SS GSG S SS + S G DSY K + + + +S+ A
Sbjct: 747 KSGSASSYGSGSSFQSSGKYGGFGSRDGDRFNDSYRDKGSYEEEKDYQGKSHHATAGDNQ 926
Query: 192 TSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSAGNMEPCSSISPNL 251
+S A S K + N + +V +N GSV S + P +S ++
Sbjct: 927 ENSFKKGSARSASKSQEN--------KSSRVSKSSTNANNYGSVPSQSSSVPANSTEDDM 1082
Query: 252 VDKSPKVT 259
D P+ T
Sbjct: 1083DDFDPRGT 1106
>BE556509
Length = 367
Score = 33.9 bits (76), Expect = 0.60
Identities = 13/38 (34%), Positives = 23/38 (60%)
Frame = +3
Query: 37 RSSRDNRGGPFGQRDWRGHSWEASNGSPNLSRRPQDMN 74
+S RD R + +W G +E+ +G+ N +R P+D+N
Sbjct: 162 QSQRDKRDRKTSENNWLGMEYESCHGTLNTNRSPEDLN 275
>TC216677 similar to GB|AAS58514.1|45357110|AY550303 MYB transcription factor
{Arabidopsis thaliana;} , partial (51%)
Length = 1373
Score = 33.5 bits (75), Expect = 0.78
Identities = 19/69 (27%), Positives = 33/69 (47%)
Frame = +3
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
+ P I K R WT E + FLE F +D++KI +F+ K+ ++H
Sbjct: 201 IRKPYTITKSREN---WTEPEHDKFLEAIQLFDRDWKKIEAFVGSKSV-----IQIRSHA 356
Query: 782 SECFEKLKR 790
+ F K+++
Sbjct: 357 QKYFLKVQK 383
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.128 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 69,211,373
Number of Sequences: 63676
Number of extensions: 985998
Number of successful extensions: 4421
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 4300
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4408
length of query: 1620
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1510
effective length of database: 5,635,272
effective search space: 8509260720
effective search space used: 8509260720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC126015.12


