

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.11 + phase: 0
(371 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
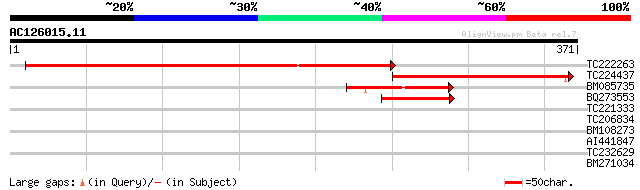
Score E
Sequences producing significant alignments: (bits) Value
TC222263 similar to UP|Q8VYW0 (Q8VYW0) AT3g52260/T25B15_30, part... 415 e-116
TC224437 weakly similar to UP|Q97H99 (Q97H99) Predicted pseudour... 211 4e-55
BM085735 58 6e-09
BQ273553 similar to GP|6721111|gb|A T4O12.25 {Arabidopsis thalia... 45 7e-05
TC221333 similar to UP|Q8W4Q8 (Q8W4Q8) AT5g51140/MWD22_8, partia... 30 1.3
TC206834 similar to UP|Q944I3 (Q944I3) AT3g01400/T13O15_4, parti... 30 1.3
BM108273 similar to GP|18376331|emb related to hydroxyproline-ri... 29 3.0
AI441847 homologue to PIR|T11622|T11 extensin class 1 precursor ... 28 6.6
TC232629 28 6.6
BM271034 similar to GP|10177899|dbj contains similarity to integ... 28 8.7
>TC222263 similar to UP|Q8VYW0 (Q8VYW0) AT3g52260/T25B15_30, partial (65%)
Length = 729
Score = 415 bits (1067), Expect = e-116
Identities = 203/242 (83%), Positives = 222/242 (90%)
Frame = +3
Query: 11 WPEFNDGLSYDDVVRPSDAGLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPN 70
WPE NDGLSYDDVVR SDAG TLIEFYSTKYKSSAPLQGWLQRIKS QIT+D VVTD N
Sbjct: 3 WPECNDGLSYDDVVRASDAGATLIEFYSTKYKSSAPLQGWLQRIKSGQITVDGGVVTDSN 182
Query: 71 TILRVGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQL 130
T+LRVGSKL+YHR+PWKEPDAPH I+VLYEDDD+IALNKPSGLQVLPGGL+QQRT+LTQL
Sbjct: 183 TVLRVGSKLIYHRLPWKEPDAPHMIDVLYEDDDMIALNKPSGLQVLPGGLYQQRTILTQL 362
Query: 131 QWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGR 190
QWE NN+ T E H++ HPVPVHRLGRGTSGILLCAKTKLARARLAS+FADGTS + GK R
Sbjct: 363 QWEANNQGTCEMHKRLHPVPVHRLGRGTSGILLCAKTKLARARLASHFADGTSHVGGK-R 539
Query: 191 HTNQELGKIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVASQSGKPALSVVHV 250
T QELGKIAK+YRALVSGI+ ND+VTINQPIG+VKYPGVAKGLYVAS+SGKPALSVV +
Sbjct: 540 DTKQELGKIAKMYRALVSGIVENDKVTINQPIGIVKYPGVAKGLYVASESGKPALSVVDI 719
Query: 251 LE 252
LE
Sbjct: 720 LE 725
>TC224437 weakly similar to UP|Q97H99 (Q97H99) Predicted pseudouridylate
synthase, YLYB B.subtilis ortholog, partial (11%)
Length = 543
Score = 211 bits (537), Expect = 4e-55
Identities = 101/121 (83%), Positives = 109/121 (89%), Gaps = 2/121 (1%)
Frame = +2
Query: 251 LERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFA 310
LE N++ENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLY VGGQPKCLDCDF+DESFA
Sbjct: 26 LETNIQENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYGVGGQPKCLDCDFVDESFA 205
Query: 311 EDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLPSILQTHEEA--IAQAQQ 368
EDGGY RP+KPVPGDCGY+LHAH+L LSHP TNE+IE+IAPLPS L T EEA IA QQ
Sbjct: 206 EDGGYQRPTKPVPGDCGYYLHAHKLVLSHPTTNELIEIIAPLPSALHTAEEAKEIAIMQQ 385
Query: 369 T 369
T
Sbjct: 386 T 388
>BM085735
Length = 422
Score = 58.2 bits (139), Expect = 6e-09
Identities = 35/73 (47%), Positives = 46/73 (62%), Gaps = 3/73 (4%)
Frame = -1
Query: 221 PIGMVKYPGVA---KGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIH 277
PIG K+ A + L +++ K A + + N+ + + QVKIQSGRPHQIRIH
Sbjct: 266 PIGYPKFCLCASCYESLLRSNKRKKKATIKLEIQNLNIPD-CLVCQVKIQSGRPHQIRIH 90
Query: 278 LSFIGHPLLGDPL 290
LSFIGHPLLG+ L
Sbjct: 89 LSFIGHPLLGEYL 51
>BQ273553 similar to GP|6721111|gb|A T4O12.25 {Arabidopsis thaliana}, partial
(6%)
Length = 420
Score = 44.7 bits (104), Expect = 7e-05
Identities = 22/48 (45%), Positives = 31/48 (63%)
Frame = +2
Query: 244 ALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLY 291
A S V+E + +LV+ K+++GR HQIR H ++G PLLGD LY
Sbjct: 14 AASRYKVIEILAGGSCSLVEWKLETGRTHQIRAHAKYLGVPLLGDELY 157
>TC221333 similar to UP|Q8W4Q8 (Q8W4Q8) AT5g51140/MWD22_8, partial (19%)
Length = 629
Score = 30.4 bits (67), Expect = 1.3
Identities = 11/19 (57%), Positives = 14/19 (72%)
Frame = +2
Query: 273 QIRIHLSFIGHPLLGDPLY 291
QIR+HL + GHP+ D LY
Sbjct: 11 QIRVHLQYSGHPIANDMLY 67
>TC206834 similar to UP|Q944I3 (Q944I3) AT3g01400/T13O15_4, partial (89%)
Length = 1740
Score = 30.4 bits (67), Expect = 1.3
Identities = 17/41 (41%), Positives = 22/41 (53%)
Frame = -3
Query: 32 TLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTI 72
T I Y YK+S+PL+ +LQ S + DAR V TI
Sbjct: 1711 TRIILYHKLYKASSPLRRYLQSFNSPSLLEDARCVLTKFTI 1589
>BM108273 similar to GP|18376331|emb related to hydroxyproline-rich
glycoprotein {Neurospora crassa}, partial (2%)
Length = 414
Score = 29.3 bits (64), Expect = 3.0
Identities = 25/76 (32%), Positives = 29/76 (37%), Gaps = 7/76 (9%)
Frame = -3
Query: 272 HQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAEDGGYTRPSKPVPGDCGYHLH 331
H HL F+GHPLL P +G F G Y S P P YH H
Sbjct: 253 HHHHQHLXFLGHPLL--PPLGIG---------HYSYHFQL*GTYMXESHPPP----YHSH 119
Query: 332 A-------HRLFLSHP 340
A H +F S+P
Sbjct: 118 AFFPSLTPHLVFFSYP 71
>AI441847 homologue to PIR|T11622|T11 extensin class 1 precursor - cowpea,
partial (29%)
Length = 429
Score = 28.1 bits (61), Expect = 6.6
Identities = 13/31 (41%), Positives = 17/31 (53%)
Frame = +3
Query: 328 YHLHAHRLFLSHPITNEIIEMIAPLPSILQT 358
+HL L L HP II ++ PLP +L T
Sbjct: 294 HHLTITNLLLLHPQLPTIINLLLPLPPLLTT 386
>TC232629
Length = 514
Score = 28.1 bits (61), Expect = 6.6
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Frame = +1
Query: 93 HTIEVLYEDDDIIALNKPSGLQVLPGGL-FQQRTVLTQL 130
HT++VLY+D ++ L P +PG L F++ VL QL
Sbjct: 325 HTLQVLYQDFSLVTLQVP----YVPGFLAFREAPVLLQL 429
>BM271034 similar to GP|10177899|dbj contains similarity to integral membrane
protein~gene_id:K16H17.9 {Arabidopsis thaliana}, partial
(12%)
Length = 425
Score = 27.7 bits (60), Expect = 8.7
Identities = 12/26 (46%), Positives = 14/26 (53%)
Frame = -2
Query: 134 TNNKSTIEAHQKPHPVPVHRLGRGTS 159
TN K H + HP P+ R GRG S
Sbjct: 91 TNQKGNRHHHGQRHPFPILRWGRGLS 14
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.137 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,943,230
Number of Sequences: 63676
Number of extensions: 223177
Number of successful extensions: 1070
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1063
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1069
length of query: 371
length of database: 12,639,632
effective HSP length: 99
effective length of query: 272
effective length of database: 6,335,708
effective search space: 1723312576
effective search space used: 1723312576
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126015.11


