

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126010.6 + phase: 0
(518 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
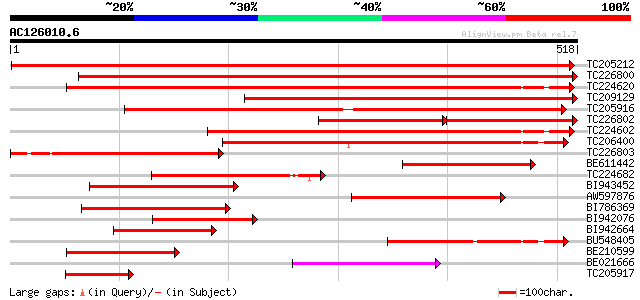
Score E
Sequences producing significant alignments: (bits) Value
TC205212 homologue to UP|GLYM_SOLTU (P50433) Serine hydroxymethy... 937 0.0
TC226800 similar to UP|GLYM_PEA (P34899) Serine hydroxymethyltra... 824 0.0
TC224620 similar to UP|O23254 (O23254) Serine hydroxymethyltrans... 536 e-152
TC209129 similar to UP|GLYM_PEA (P34899) Serine hydroxymethyltra... 535 e-152
TC205916 similar to UP|Q9SUU0 (Q9SUU0) Glycine hydroxymethyltran... 503 e-143
TC226802 similar to UP|Q940M9 (Q940M9) AT4g37930/F20D10_50, part... 213 e-103
TC224602 similar to UP|O23254 (O23254) Serine hydroxymethyltrans... 353 8e-98
TC206400 similar to UP|Q9LM59 (Q9LM59) F2E2.7 (At1g22020/F2E2_3)... 302 2e-82
TC226803 similar to UP|GLYM_PEA (P34899) Serine hydroxymethyltra... 278 5e-75
BE611442 similar to SP|P50433|GLYM Serine hydroxymethyltransfera... 188 4e-48
TC224682 similar to UP|O23254 (O23254) Serine hydroxymethyltrans... 182 4e-46
BI943452 176 2e-44
AW597876 similar to SP|P50433|GLYM Serine hydroxymethyltransfera... 171 6e-43
BI786369 similar to GP|9280677|gb| F2E2.7 {Arabidopsis thaliana}... 171 6e-43
BI942076 similar to SP|P34899|GLYM_ Serine hydroxymethyltransfer... 171 9e-43
BI942664 129 4e-30
BU548405 125 5e-29
BE210599 similar to GP|21592544|gb hydroxymethyltransferase {Ara... 121 9e-28
BE021666 similar to GP|11762130|gb AT4g13930 {Arabidopsis thalia... 115 4e-26
TC205917 similar to UP|Q94JQ3 (Q94JQ3) AT4g32520/F8B4_220, parti... 82 6e-16
>TC205212 homologue to UP|GLYM_SOLTU (P50433) Serine
hydroxymethyltransferase, mitochondrial precursor
(Serine methylase) (Glycine hydroxymethyltransferase)
(SHMT) , complete
Length = 1918
Score = 937 bits (2423), Expect = 0.0
Identities = 463/515 (89%), Positives = 496/515 (95%)
Frame = +1
Query: 2 AMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEIDP 61
AMAMALRRLSSSI+K RPLF+A S+YYKSSLPDEAVYDKE V+WPKQLN+ LE +DP
Sbjct: 1 AMAMALRRLSSSIDKPLRPLFNAGSLYYKSSLPDEAVYDKERPGVTWPKQLNAPLEVVDP 180
Query: 62 EIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDM 121
EIADIIELEKARQWKGLELIPSENFTS+SVMQAVGS+MTNKYSEGYPGARYYGGNEYIDM
Sbjct: 181 EIADIIELEKARQWKGLELIPSENFTSVSVMQAVGSVMTNKYSEGYPGARYYGGNEYIDM 360
Query: 122 AETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSH 181
AETLCQKRALEAFRLDPAKWGVNVQPLSGSP+NFHVYTALLKPH+RIMALDLPHGGHLSH
Sbjct: 361 AETLCQKRALEAFRLDPAKWGVNVQPLSGSPANFHVYTALLKPHERIMALDLPHGGHLSH 540
Query: 182 GYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDY 241
GYQTDTKKISAVSIFFETMPYRL+ESTGYIDYDQ+EKSATLFRPKLIVAGASAYARLYDY
Sbjct: 541 GYQTDTKKISAVSIFFETMPYRLNESTGYIDYDQMEKSATLFRPKLIVAGASAYARLYDY 720
Query: 242 ARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRK 301
R+RKVCDKQKA++LADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIF+RK
Sbjct: 721 ERVRKVCDKQKAILLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFYRK 900
Query: 302 GLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVL 361
G+KE+NKQGKEV YDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEY+AYQEQVL
Sbjct: 901 GVKEINKQGKEVLYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYRAYQEQVL 1080
Query: 362 SNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGD 421
SN KFAQALSE+ YELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGD
Sbjct: 1081SNSFKFAQALSERSYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGD 1260
Query: 422 VSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQ 481
VSAMVPGGIRMGTPALTSRGFVEEDFVKVAE+FDA+V +A+KIK ESKGTKLKDF+ T++
Sbjct: 1261VSAMVPGGIRMGTPALTSRGFVEEDFVKVAEFFDAAVKIAVKIKGESKGTKLKDFLATIE 1440
Query: 482 SSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
SSS QSEI+KLR DVEE+AKQFPTIGF+K++MK+
Sbjct: 1441SSSTFQSEIAKLRLDVEEYAKQFPTIGFDKATMKH 1545
>TC226800 similar to UP|GLYM_PEA (P34899) Serine hydroxymethyltransferase,
mitochondrial precursor (Serine methylase) (Glycine
hydroxymethyltransferase) (SHMT) , partial (88%)
Length = 1673
Score = 824 bits (2128), Expect = 0.0
Identities = 406/455 (89%), Positives = 432/455 (94%)
Frame = +1
Query: 64 ADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAE 123
ADIIELEKARQWKG ELIPSENFTSLSVMQAVGS+MTNKYSEGYPGARYYGGNEYIDMAE
Sbjct: 1 ADIIELEKARQWKGFELIPSENFTSLSVMQAVGSVMTNKYSEGYPGARYYGGNEYIDMAE 180
Query: 124 TLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGY 183
TLCQKRALEAF+LDPAKWGVNVQ LSGSPSNF VYTALLKPH+RIMALDLPHGGHLSHGY
Sbjct: 181 TLCQKRALEAFQLDPAKWGVNVQSLSGSPSNFQVYTALLKPHERIMALDLPHGGHLSHGY 360
Query: 184 QTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDYAR 243
QTDTKKISAVSIFFETMPYRL+ESTGYIDYDQLEKSA LFRPKLIVAGASAYARLYDYAR
Sbjct: 361 QTDTKKISAVSIFFETMPYRLNESTGYIDYDQLEKSAALFRPKLIVAGASAYARLYDYAR 540
Query: 244 IRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGL 303
+RKVCDKQKAV+LADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKG+
Sbjct: 541 VRKVCDKQKAVLLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGV 720
Query: 304 KEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSN 363
KE+NKQGKEV YDYEDKINQAVFPGLQGGPHNHTI+GLAVALKQA TPE+K YQ+QVLSN
Sbjct: 721 KEINKQGKEVLYDYEDKINQAVFPGLQGGPHNHTISGLAVALKQAMTPEFKNYQKQVLSN 900
Query: 364 CAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVS 423
C+ FAQ+L EKGYELVSGGT+NHLVLVNL+NKGIDGSRVEKVLEAVHIAANKNTVPGDVS
Sbjct: 901 CSAFAQSLLEKGYELVSGGTDNHLVLVNLRNKGIDGSRVEKVLEAVHIAANKNTVPGDVS 1080
Query: 424 AMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQSS 483
AMVPGGIRMGTPALTSRGFVEEDF KVAEYFDA+V LAL+IK + GTKLKDFV +QS
Sbjct: 1081AMVPGGIRMGTPALTSRGFVEEDFEKVAEYFDAAVKLALQIKENTNGTKLKDFVAAMQSD 1260
Query: 484 SYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK 518
VQS+I+ LRH+VE++AKQFPTIGF+ +MKY K
Sbjct: 1261EQVQSKIANLRHEVEDYAKQFPTIGFDIETMKYGK 1365
>TC224620 similar to UP|O23254 (O23254) Serine hydroxymethyltransferase
(Serine methylase) (Glycine hydroxymethyltransferase)
(SHMT) , complete
Length = 1887
Score = 536 bits (1380), Expect = e-152
Identities = 273/466 (58%), Positives = 341/466 (72%), Gaps = 2/466 (0%)
Frame = +3
Query: 53 NSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARY 112
N+ L +DPEI D+IE EK RQ +G+ELI SENFTS +V++A+GS +TNKYSEG PG RY
Sbjct: 108 NTPLATVDPEIHDLIEKEKRRQCRGIELIASENFTSFAVIEALGSALTNKYSEGMPGNRY 287
Query: 113 YGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALD 172
YGGNEYID E LC+ RAL+AF LD WGVNVQP SGSP+NF YTA+L PHDRIM LD
Sbjct: 288 YGGNEYIDQIENLCRSRALQAFHLDAQSWGVNVQPYSGSPANFAAYTAVLNPHDRIMGLD 467
Query: 173 LPHGGHLSHGYQTD-TKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAG 231
LP GGHL+HGY T KKISA SI+FE++PY+++ +TGYIDYD+LE+ A FRPKLI+ G
Sbjct: 468 LPSGGHLTHGYYTSGGKKISATSIYFESLPYKVNSTTGYIDYDRLEEKALDFRPKLIICG 647
Query: 232 ASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRG 291
SAY R +DY R R+V DK A++L DMAH SGLVAA + SPF+Y D+VTTTTHKSLRG
Sbjct: 648 GSAYPRDWDYKRFREVADKCGALLLCDMAHTSGLVAAQEVNSPFEYCDIVTTTTHKSLRG 827
Query: 292 PRGAMIFFRKGLKEVNK-QGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATT 350
PR MIF+RKG K K Q + YD+EDKIN AVFP LQGGPHNH I LAVALKQA +
Sbjct: 828 PRAGMIFYRKGPKPPKKGQPENAVYDFEDKINFAVFPSLQGGPHNHQIGALAVALKQAAS 1007
Query: 351 PEYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVH 410
P +KAY +QV +N + L KGY LV+GGTENHLVL +L+ G+ G++VEK+ + +
Sbjct: 1008PGFKAYAKQVKANAVALGKYLMGKGYSLVTGGTENHLVLWDLRPLGLTGNKVEKLCDLCN 1187
Query: 411 IAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKG 470
I NKN V GD SA+ PGG+R+G PA+TSRG VE+DF ++ E+ +V L L+I+ E G
Sbjct: 1188ITVNKNAVFGDSSALAPGGVRIGAPAMTSRGLVEKDFEQIGEFLHRAVTLTLEIQKE-HG 1364
Query: 471 TKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
LKDF + L ++ ++ L+ DVE+F+ F GF S MKY
Sbjct: 1365KLLKDFNKGLVNNKAIED----LKADVEKFSALFDMPGFLVSEMKY 1490
>TC209129 similar to UP|GLYM_PEA (P34899) Serine hydroxymethyltransferase,
mitochondrial precursor (Serine methylase) (Glycine
hydroxymethyltransferase) (SHMT) , partial (58%)
Length = 1054
Score = 535 bits (1379), Expect = e-152
Identities = 263/304 (86%), Positives = 292/304 (95%)
Frame = +2
Query: 215 QLEKSATLFRPKLIVAGASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSP 274
+LE +A LFRPKLIVAGA+AYARLYDYARIRKVCDKQKAV+LADMAHISGLVAAGVIPSP
Sbjct: 68 KLESTAKLFRPKLIVAGATAYARLYDYARIRKVCDKQKAVLLADMAHISGLVAAGVIPSP 247
Query: 275 FDYADVVTTTTHKSLRGPRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPH 334
FDYADVVTTTTHKSLRGPRGAMIFFRKG+KE+N++G+EV YDYEDKIN+AVFPGLQ GPH
Sbjct: 248 FDYADVVTTTTHKSLRGPRGAMIFFRKGVKEINEKGEEVMYDYEDKINRAVFPGLQSGPH 427
Query: 335 NHTITGLAVALKQATTPEYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKN 394
H+ITGLAVALKQATTP Y+AYQEQVL NC+KFAQALSEKGYELVSGGTENHL+LVNLK+
Sbjct: 428 FHSITGLAVALKQATTPNYRAYQEQVLRNCSKFAQALSEKGYELVSGGTENHLLLVNLKS 607
Query: 395 KGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYF 454
KGIDGSRV+KVLE+VHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGF EEDFV VAE+F
Sbjct: 608 KGIDGSRVQKVLESVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFAEEDFVMVAEFF 787
Query: 455 DASVNLALKIKAESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSM 514
DA+VNLA+KIK+E+KG+KLKDF+ T+QSSSY QSEI+KLRHDVEE+AKQFPTIGF+K +M
Sbjct: 788 DAAVNLAVKIKSETKGSKLKDFLATIQSSSYFQSEIAKLRHDVEEYAKQFPTIGFDKETM 967
Query: 515 KYNK 518
KYNK
Sbjct: 968 KYNK 979
>TC205916 similar to UP|Q9SUU0 (Q9SUU0) Glycine hydroxymethyltransferase
-like protein , partial (86%)
Length = 1347
Score = 503 bits (1295), Expect = e-143
Identities = 250/404 (61%), Positives = 306/404 (74%), Gaps = 1/404 (0%)
Frame = +3
Query: 106 GYPGARYYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPH 165
G PG RYYGGNEYID E LCQ+RAL AF +D KWGVNVQ LSGSP+NF VYTA+LKPH
Sbjct: 3 GLPGKRYYGGNEYIDELEILCQQRALAAFHVDENKWGVNVQTLSGSPANFAVYTAVLKPH 182
Query: 166 DRIMALDLPHGGHLSHGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRP 225
DRIM LDLPHGGHLSHG+ T K++SA SI+FE+MPYRLDESTG IDYD LEK+ATLFRP
Sbjct: 183 DRIMGLDLPHGGHLSHGFMTPKKRVSATSIYFESMPYRLDESTGLIDYDMLEKTATLFRP 362
Query: 226 KLIVAGASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTT 285
KLIVAGASAY R DY R+RK+ D+ A ++ DMAHISGLVAA V+ +PF+Y D+VTTTT
Sbjct: 363 KLIVAGASAYPRDIDYPRMRKIADEVGAFLMMDMAHISGLVAASVLSNPFEYCDIVTTTT 542
Query: 286 HKSLRGPRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVAL 345
HKSLRGPRG MIFF+K D E IN AVFPGLQGGPHNHTI GLAV L
Sbjct: 543 HKSLRGPRGGMIFFKKDTVH--------GVDLEPAINNAVFPGLQGGPHNHTIGGLAVCL 698
Query: 346 KQATTPEYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKV 405
K A +PE+K YQ QV++NC A+ L E GY+LVSGG++NHLVLV+L+ G+DG+RVEK+
Sbjct: 699 KYAQSPEFKNYQNQVVANCRALAKRLIEHGYKLVSGGSDNHLVLVDLRPSGLDGARVEKI 878
Query: 406 LEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIK 465
L+ I NKN+VPGD SA+VPGGIR+G PA+T+RG E++F +A++ V ++L+ K
Sbjct: 879 LDLASITLNKNSVPGDKSALVPGGIRIGAPAMTTRGLGEKEFSLIADFIHEGVQISLEAK 1058
Query: 466 AESKGTKLKDFVETLQSSSYVQSE-ISKLRHDVEEFAKQFPTIG 508
+ GTKL+DF++ + SS + E +S+LR VE Q+P G
Sbjct: 1059SLVSGTKLQDFLKFVTSSEFPLGEKVSELRRKVEALTTQYPIPG 1190
>TC226802 similar to UP|Q940M9 (Q940M9) AT4g37930/F20D10_50, partial (44%)
Length = 1042
Score = 213 bits (542), Expect(2) = e-103
Identities = 101/118 (85%), Positives = 112/118 (94%)
Frame = +3
Query: 283 TTTHKSLRGPRGAMIFFRKGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLA 342
TTTHKSLRGPRGAMIFFRKG+KE+NKQGKEV YDYEDKINQAVFPGLQGGPHNHTI+GLA
Sbjct: 3 TTTHKSLRGPRGAMIFFRKGVKEINKQGKEVLYDYEDKINQAVFPGLQGGPHNHTISGLA 182
Query: 343 VALKQATTPEYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGS 400
VALKQA TPE+K YQ+QVLSNC+ FAQ+L EKGYELVSGGT+NHLVLVNL+NKG +G+
Sbjct: 183 VALKQAMTPEFKNYQKQVLSNCSAFAQSLLEKGYELVSGGTDNHLVLVNLRNKG*EGT 356
Score = 182 bits (461), Expect(2) = e-103
Identities = 90/119 (75%), Positives = 103/119 (85%)
Frame = +1
Query: 400 SRVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVN 459
+RVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDF KVAEYFDA+V
Sbjct: 337 TRVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFEKVAEYFDAAVK 516
Query: 460 LALKIKAESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK 518
LAL+IK + GTKLKDFV +QS +QS+I+ L H+VE++AK+FPTIGF +MKY K
Sbjct: 517 LALQIKENTNGTKLKDFVAAMQSDEQIQSKIANLCHEVEDYAKKFPTIGFNIETMKYGK 693
>TC224602 similar to UP|O23254 (O23254) Serine hydroxymethyltransferase
(Serine methylase) (Glycine hydroxymethyltransferase)
(SHMT) , partial (71%)
Length = 1221
Score = 353 bits (907), Expect = 8e-98
Identities = 185/338 (54%), Positives = 238/338 (69%), Gaps = 2/338 (0%)
Frame = +3
Query: 181 HGYQTDT-KKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLY 239
HGY T KKISA SI+FE++PY+++ +TGYIDYD+LE+ A FRPKLI+ G SAY R +
Sbjct: 3 HGYYTSGGKKISATSIYFESLPYKVNSTTGYIDYDRLEEKALDFRPKLIICGGSAYPRDW 182
Query: 240 DYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFF 299
DY R R++ DK A++L DMAH SGLVAA + SPF+Y D+VTTTTHKSLRGPR MIF+
Sbjct: 183 DYKRFREIADKCGALLLCDMAHTSGLVAAQEVNSPFEYCDIVTTTTHKSLRGPRAGMIFY 362
Query: 300 RKGLKEVNK-QGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQE 358
RKG K K Q + YD+EDKIN AVFP LQGGPHNH I LAVALKQA +P +KAY +
Sbjct: 363 RKGPKPPKKGQPENAVYDFEDKINFAVFPSLQGGPHNHQIGALAVALKQAASPGFKAYAK 542
Query: 359 QVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTV 418
QV +N L KGY LV+GGTENHLVL +L+ G+ G++VEK+ + +I NKN V
Sbjct: 543 QVKANAVALGNYLMGKGYSLVTGGTENHLVLWDLRPLGLTGNKVEKLCDLCNITVNKNAV 722
Query: 419 PGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVE 478
GD SA+ PGG+R+G PA+TSRG VE+DF ++ E+ +V L L+I+ E G LKDF +
Sbjct: 723 FGDSSALAPGGVRIGAPAMTSRGLVEKDFEQIGEFLHRAVTLTLEIQKE-HGKLLKDFNK 899
Query: 479 TLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
L ++ ++ L+ DVE+F+ F GF S MKY
Sbjct: 900 GLVNNKAIED----LKADVEKFSATFDMPGFLVSEMKY 1001
>TC206400 similar to UP|Q9LM59 (Q9LM59) F2E2.7 (At1g22020/F2E2_3), partial
(49%)
Length = 1572
Score = 302 bits (774), Expect = 2e-82
Identities = 160/322 (49%), Positives = 216/322 (66%), Gaps = 6/322 (1%)
Frame = +3
Query: 195 IFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDYARIRKVCDKQKAV 254
IFFE++PY+++ TGYIDYD+LE+ A FRPK+++ G S+Y R +DYAR R + DK AV
Sbjct: 6 IFFESLPYKVNPQTGYIDYDKLEERALDFRPKILICGGSSYPREWDYARFRHIADKCGAV 185
Query: 255 MLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKEVNK-----Q 309
+L DMA ISG++AA +PFDY D+VT+TTHKSLRGPRG +IF+RKG K N+ Q
Sbjct: 186 LLCDMAQISGIIAAKECVNPFDYCDIVTSTTHKSLRGPRGGIIFYRKGTKPRNRGILLSQ 365
Query: 310 GKEVF-YDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSNCAKFA 368
G E YD+E+KIN AVFP +QGGPHN+ I LA+ALKQ TPEYKAY +QV N A
Sbjct: 366 GHESDQYDFEEKINFAVFPSMQGGPHNNHIAALAIALKQVATPEYKAYMQQVKKNAQALA 545
Query: 369 QALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPG 428
AL + LV+GGT+NHL+L +L+ G+ G EKV E HI NK + GD ++PG
Sbjct: 546 CALLRRKCRLVTGGTDNHLILWDLRPLGLTGKFYEKVCETCHITLNKIAIFGDNGTIIPG 725
Query: 429 GIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQSSSYVQS 488
G+R+GTPA+TSRG +E F +AE+ + +A ++ E G K ++ L+S+
Sbjct: 726 GVRVGTPAMTSRGCLEAHFETMAEFLIRAAQIASILQRE-HGKLQKTTLKGLESN----R 890
Query: 489 EISKLRHDVEEFAKQFPTIGFE 510
++ +LR VE FA QF GF+
Sbjct: 891 DVVELRARVEAFATQFAMPGFD 956
>TC226803 similar to UP|GLYM_PEA (P34899) Serine hydroxymethyltransferase,
mitochondrial precursor (Serine methylase) (Glycine
hydroxymethyltransferase) (SHMT) , partial (26%)
Length = 756
Score = 278 bits (710), Expect = 5e-75
Identities = 142/195 (72%), Positives = 160/195 (81%)
Frame = +1
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
MAMA+ LRRLSS++NK PL SA+S+Y++ S A +K+ SR W KQLN LE ID
Sbjct: 184 MAMALPLRRLSSTLNK---PLASATSIYHRMSSSLSA-QEKDKSRADWIKQLNDPLETID 351
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEIADIIELEKARQWKG ELIPSENFTSLSVMQAVGS+MTNKYSEGYPGARYYGGNEYID
Sbjct: 352 PEIADIIELEKARQWKGFELIPSENFTSLSVMQAVGSVMTNKYSEGYPGARYYGGNEYID 531
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAETLCQK ALEAFRLDPAKWGVNVQ LSGS S F VYTAL PH+RIMAL LP GGH+
Sbjct: 532 MAETLCQKPALEAFRLDPAKWGVNVQSLSGSRSIFQVYTALCVPHERIMALGLPPGGHVV 711
Query: 181 HGYQTDTKKISAVSI 195
HG++T+ ++ + I
Sbjct: 712 HGFRTNPRRYQLIDI 756
>BE611442 similar to SP|P50433|GLYM Serine hydroxymethyltransferase
mitochondrial precursor (EC 2.1.2.1) (Serine methylase),
partial (23%)
Length = 364
Score = 188 bits (478), Expect = 4e-48
Identities = 92/121 (76%), Positives = 110/121 (90%)
Frame = +2
Query: 360 VLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVP 419
VL NC+KFAQALSEKGY+LVSGGT+NHL+LVNLK+KGIDGSRV+KVLE+VHIAANKNTV
Sbjct: 2 VLRNCSKFAQALSEKGYDLVSGGTDNHLLLVNLKSKGIDGSRVQKVLESVHIAANKNTVH 181
Query: 420 GDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVET 479
GDVS MVPGGIRMGTPALTS GF EEDF VA+ FDA+VNLA+ I++E++G++L DF+ T
Sbjct: 182 GDVSTMVPGGIRMGTPALTSTGFAEEDFEVVAKMFDAAVNLAVNIRSETRGSELSDFLAT 361
Query: 480 L 480
+
Sbjct: 362 V 364
>TC224682 similar to UP|O23254 (O23254) Serine hydroxymethyltransferase
(Serine methylase) (Glycine hydroxymethyltransferase)
(SHMT) , partial (27%)
Length = 689
Score = 182 bits (461), Expect = 4e-46
Identities = 94/164 (57%), Positives = 117/164 (71%), Gaps = 5/164 (3%)
Frame = +1
Query: 130 ALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQTD-TK 188
AL+AF LD WGVNVQP SGSP+NF YTA+L PHDRIM LDLP GGHL+HGY T K
Sbjct: 1 ALQAFHLDAQSWGVNVQPYSGSPANFAAYTAVLNPHDRIMGLDLPSGGHLTHGYYTSGGK 180
Query: 189 KISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDYARIRKVC 248
KISA SI+FE++PY+++ +TGYIDYD+LE+ A FRPKLI+ G SAY R +DY R R++
Sbjct: 181 KISATSIYFESLPYKVNSTTGYIDYDRLEEKALDFRPKLIICGGSAYPRDWDYKRFREIA 360
Query: 249 DKQKAVMLADMAHISGLVAAGVIP----SPFDYADVVTTTTHKS 288
DK A++L + H G P +PF Y +VTTTTHK+
Sbjct: 361 DKCGALLL--LRH--GAHXRPCWPPKN*TPFXYCXIVTTTTHKT 480
>BI943452
Length = 420
Score = 176 bits (447), Expect = 2e-44
Identities = 87/137 (63%), Positives = 106/137 (76%), Gaps = 1/137 (0%)
Frame = +1
Query: 74 QWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETLCQKRALEA 133
Q++G+ELI SENF +VM+A+GS +TNKYSEG PGARYYGGN+YID ETLC +RAL A
Sbjct: 10 QFRGIELIASENFVCRAVMEALGSHLTNKYSEGMPGARYYGGNQYIDEIETLCCERALNA 189
Query: 134 FRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQT-DTKKISA 192
F LDP WGVNVQP S + +NF VYT LL P DRIM LD P GG+ SHGY T + KK+S
Sbjct: 190 FGLDPKCWGVNVQPYSCTSANFAVYTXLLLPGDRIMGLDTPSGGNTSHGYYTPNGKKVSG 369
Query: 193 VSIFFETMPYRLDESTG 209
SIFFE++ Y+++ STG
Sbjct: 370 ASIFFESLAYKVNPSTG 420
>AW597876 similar to SP|P50433|GLYM Serine hydroxymethyltransferase
mitochondrial precursor (EC 2.1.2.1) (Serine methylase),
partial (24%)
Length = 427
Score = 171 bits (434), Expect = 6e-43
Identities = 94/142 (66%), Positives = 108/142 (75%), Gaps = 1/142 (0%)
Frame = +2
Query: 313 VFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSNCAKFAQALS 372
V YDYEDKIN+AVFPGLQ GPH H+ITGLAVAL QATTP Y AYQE VL + + +
Sbjct: 2 VMYDYEDKINRAVFPGLQSGPHFHSITGLAVALMQATTPNYGAYQEHVLPLLLQISTST* 181
Query: 373 -EKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPGGIR 431
KG+ LVSGGTEN+L+LVNLK+KGIDGSRV VLE+VHIAA NTVPGDV M P GI
Sbjct: 182 *SKGFHLVSGGTENNLLLVNLKSKGIDGSRVL*VLESVHIAAYTNTVPGDVCDMSPVGII 361
Query: 432 MGTPALTSRGFVEEDFVKVAEY 453
+GT ALT G ++D VKVA +
Sbjct: 362 IGTTALTY*GSADDD*VKVAVF 427
>BI786369 similar to GP|9280677|gb| F2E2.7 {Arabidopsis thaliana}, partial
(22%)
Length = 422
Score = 171 bits (434), Expect = 6e-43
Identities = 87/137 (63%), Positives = 103/137 (74%), Gaps = 1/137 (0%)
Frame = +1
Query: 66 IIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETL 125
I+ EK RQ+KG+ELI SENF +VM+A+GS ++NKYSEG PGA+YY GN+YID E L
Sbjct: 10 IMGKEKQRQFKGIELIASENFVCRAVMEALGSHLSNKYSEGMPGAKYYTGNQYIDEIEFL 189
Query: 126 CQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQT 185
C +RAL AF L P WGVNVQP S + +NF VYT +L P DRIM LD P GGHLSHGY T
Sbjct: 190 CCQRALLAFDLHPNNWGVNVQPYSCTSANFAVYTGILHPGDRIMGLDSPSGGHLSHGYYT 369
Query: 186 -DTKKISAVSIFFETMP 201
KK+SA SIFFET+P
Sbjct: 370 LGGKKVSAASIFFETLP 420
>BI942076 similar to SP|P34899|GLYM_ Serine hydroxymethyltransferase
mitochondrial precursor (EC 2.1.2.1) (Serine methylase),
partial (17%)
Length = 327
Score = 171 bits (432), Expect = 9e-43
Identities = 79/96 (82%), Positives = 87/96 (90%)
Frame = +2
Query: 131 LEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQTDTKKI 190
L +F DPAKWGVNVQPLSGS +NF VYTALLKPHDRIM LDLPHGGHLSHGYQTDT K+
Sbjct: 38 LISFHSDPAKWGVNVQPLSGSSANFQVYTALLKPHDRIMGLDLPHGGHLSHGYQTDTNKV 217
Query: 191 SAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPK 226
SAVS+FFETMPYRL+E+TG+IDYDQLE +A LFRPK
Sbjct: 218 SAVSLFFETMPYRLNENTGHIDYDQLESTAKLFRPK 325
>BI942664
Length = 284
Score = 129 bits (323), Expect = 4e-30
Identities = 62/94 (65%), Positives = 70/94 (73%)
Frame = +2
Query: 96 GSIMTNKYSEGYPGARYYGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNF 155
GS +TNKYSEG PG+RYYGGN+YID ETLC +RAL AF LDP WGVNVQP S + +NF
Sbjct: 2 GSHLTNKYSEGMPGSRYYGGNQYIDEIETLCCERALNAFGLDPKCWGVNVQPYSCTCANF 181
Query: 156 HVYTALLKPHDRIMALDLPHGGHLSHGYQTDTKK 189
VYT LL P DRIM LD P GG+ +HGY T K
Sbjct: 182 SVYTGLLLPGDRIMGLDTPSGGNTNHGYYTPNGK 283
>BU548405
Length = 688
Score = 125 bits (314), Expect = 5e-29
Identities = 70/165 (42%), Positives = 100/165 (60%)
Frame = -1
Query: 346 KQATTPEYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKV 405
KQ TPEYKAY +QV N A AL + + LV+ GT+NHL+L +L G+ EKV
Sbjct: 688 KQVATPEYKAYMQQVKRNAQALASALLRRNFRLVTDGTDNHLLLWDLTALGLIDRNYEKV 509
Query: 406 LEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIK 465
EA I NK + G +S PGG+R+GTPA+TSRG +EEDF +A++ + + ++
Sbjct: 508 CEACRITLNKCAIYGSIS---PGGVRIGTPAMTSRGCLEEDFETIADFLRRAAQITSIVQ 338
Query: 466 AESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFE 510
E G KDF++ LQ++ +IS+LR+ VE F+ QF GF+
Sbjct: 337 RE-HGKSCKDFLKGLQNN----KDISELRNRVETFSSQFAMPGFD 218
>BE210599 similar to GP|21592544|gb hydroxymethyltransferase {Arabidopsis
thaliana}, partial (22%)
Length = 331
Score = 121 bits (303), Expect = 9e-28
Identities = 59/103 (57%), Positives = 75/103 (72%)
Frame = +2
Query: 53 NSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARY 112
N+ L +DP+I D+I+ EK RQ +G+ LI ENFT LSV++A+GS +TNKYSEG PG RY
Sbjct: 23 NTPLATVDPDIHDLIDKEKHRQCRGI*LIAFENFTYLSVIEALGSALTNKYSEGMPGNRY 202
Query: 113 YGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNF 155
Y G+E+ID E LC+ RAL+AF LD WGV+VQP SP F
Sbjct: 203 YCGHEFIDQIENLCRSRALQAFHLDAQSWGVDVQPYYCSPVYF 331
>BE021666 similar to GP|11762130|gb AT4g13930 {Arabidopsis thaliana}, partial
(25%)
Length = 439
Score = 115 bits (289), Expect = 4e-26
Identities = 65/136 (47%), Positives = 81/136 (58%), Gaps = 1/136 (0%)
Frame = +2
Query: 259 MAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKEVNK-QGKEVFYDY 317
M+H GL AA + PF + DVVTTTTH LR PR MIF+RKGLK K Q + YD
Sbjct: 17 MSHSIGLEAALEVNIPFQFCDVVTTTTHMILRCPRAVMIFYRKGLKPTKKGQPENAVYDI 196
Query: 318 EDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSNCAKFAQALSEKGYE 377
ED IN AVF LQGGP NH I L+V L QA P + + ++ + KG +
Sbjct: 197 EDMINFAVFLSLQGGPDNHHIGALSVVLNQAVLPVLRPTRSMLMRTPLRLENT*WGKGTD 376
Query: 378 LVSGGTENHLVLVNLK 393
LV+GGT+NHL L +L+
Sbjct: 377 LVTGGTDNHLCL*DLR 424
>TC205917 similar to UP|Q94JQ3 (Q94JQ3) AT4g32520/F8B4_220, partial (12%)
Length = 671
Score = 82.0 bits (201), Expect = 6e-16
Identities = 40/62 (64%), Positives = 48/62 (76%)
Frame = +3
Query: 52 LNSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGAR 111
L+ L E DP++ II+ EK RQ+K LELI SENFTS +VM+AVGS +TNKYSEG PG R
Sbjct: 363 LDYGLSEADPDVRAIIDKEKDRQFKSLELIASENFTSRAVMEAVGSCLTNKYSEGLPGKR 542
Query: 112 YY 113
YY
Sbjct: 543 YY 548
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.132 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,635,071
Number of Sequences: 63676
Number of extensions: 225001
Number of successful extensions: 1006
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 987
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 990
length of query: 518
length of database: 12,639,632
effective HSP length: 102
effective length of query: 416
effective length of database: 6,144,680
effective search space: 2556186880
effective search space used: 2556186880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126010.6


