

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126008.6 + phase: 0
(530 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
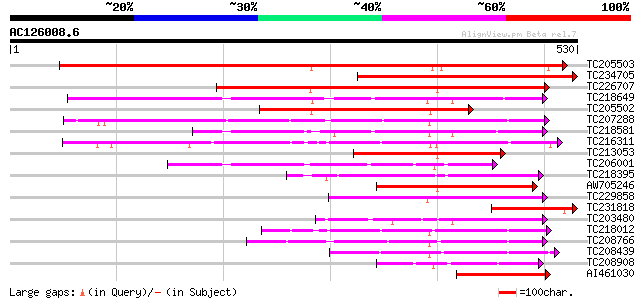
Score E
Sequences producing significant alignments: (bits) Value
TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete 455 e-128
TC234705 weakly similar to GB|CAD70577.1|32698459|MMU549317 solu... 341 5e-94
TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter... 290 8e-79
TC218649 similar to UP|Q39416 (Q39416) Integral membrane protein... 233 1e-61
TC205502 homologue to UP|Q7XA50 (Q7XA50) Sorbitol-like transport... 182 2e-46
TC207288 UP|Q7XA51 (Q7XA51) Monosaccharide transporter, complete 174 9e-44
TC218581 similar to GB|AAM70554.1|21700861|AY124845 At1g75220/F2... 167 8e-42
TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete 144 1e-34
TC213053 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter... 143 2e-34
TC206001 similar to UP|Q8VZI5 (Q8VZI5) AT5g18840/F17K4_90, parti... 134 1e-31
TC218395 similar to UP|Q9FIF2 (Q9FIF2) Sugar transporter-like pr... 128 6e-30
AW705246 127 1e-29
TC229858 weakly similar to UP|Q8VHD6 (Q8VHD6) Glucose transporte... 118 7e-27
TC231818 weakly similar to UP|Q94CI6 (Q94CI6) Sugar-porter famil... 112 4e-25
TC203480 similar to UP|Q8LPQ8 (Q8LPQ8) AT4g35300/F23E12_140, par... 110 1e-24
TC218012 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporte... 110 1e-24
TC208766 similar to UP|STA_RICCO (Q10710) Sugar carrier protein ... 110 2e-24
TC208439 weakly similar to UP|O04078 (O04078) Monosaccharid tran... 108 8e-24
TC208908 weakly similar to UP|Q8LBI9 (Q8LBI9) Sugar transporter-... 104 9e-23
AI461030 101 7e-22
>TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete
Length = 1928
Score = 455 bits (1171), Expect = e-128
Identities = 238/493 (48%), Positives = 333/493 (67%), Gaps = 18/493 (3%)
Frame = +1
Query: 47 KRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSL 106
K+ KY ACA+ AS+ ++LLGYD+GVMSGA I+IK DLK+++ Q+E L+GI+++ SL
Sbjct: 232 KKRKRNKYAFACAMLASMTSILLGYDIGVMSGAAIYIKRDLKVSDEQIEILLGIINLYSL 411
Query: 107 LGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMIS 166
+GS GRTSD IGR++T+ L +F +G M P Y LM GR +AGIGIG+ +MI+
Sbjct: 412 IGSCLAGRTSDWIGRRYTIGLGGAIFLVGSTLMGFYPHYSFLMCGRFVAGIGIGYALMIA 591
Query: 167 PIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVF 226
P+Y AE+SP +RG LT+FPE+FIN GI+LGY+SNY FS L++ + WR+ML VG +PSV
Sbjct: 592 PVYTAEVSPASSRGFLTSFPEVFINGGILLGYISNYGFSKLTLKVGWRMMLGVGAIPSVV 771
Query: 227 IGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANS------ 280
+ + +PESPRWLVM+ R+ EAR VL KT++ ++E + RLAEI+QAAG S
Sbjct: 772 LTEGVLAMPESPRWLVMRGRLGEARKVLNKTSDSKEEAQLRLAEIKQAAGIPESCNDDVV 951
Query: 281 ---GKYEDKPVWREL-LSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIED 336
+ + VW+EL L P PA+R ++I LGI FQQ SG+DA V YSP I AGI +
Sbjct: 952 QVNKQSNGEGVWKELFLYPTPAIRHIVIAALGIHFFQQASGVDAVVLYSPRIFEKAGITN 1131
Query: 337 KSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE--K 394
+ L ATVAVG KTVFIL A +D+VGR+PLL++S GM L + ++L++ + +
Sbjct: 1132DTHKLLATVAVGFVKTVFILAATFTLDRVGRRPLLLSSVGGMVLSLLTLAISLTVIDHSE 1311
Query: 395 GPLVIALG--ILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLV 452
L+ A+G I V VA FS+G GP+ WV +SEIFPLR+RAQ +A G NR S +V
Sbjct: 1312RKLMWAVGSSIAMVLAYVATFSIGAGPITWVYSSEIFPLRLRAQGAAAGVAVNRTTSAVV 1491
Query: 453 AMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF----ENEHGS 508
+M+FLS++ AI+ GG FFL+ I+ + +F +T++PET+GK+LE +E F + S
Sbjct: 1492SMTFLSLTRAITIGGAFFLYCGIATVGWIFFYTVLPETRGKTLEDMEGSFGTFRSKSNAS 1671
Query: 509 QGKEMELGDVEQL 521
+ E E G V Q+
Sbjct: 1672KAVENENGQVAQV 1710
>TC234705 weakly similar to GB|CAD70577.1|32698459|MMU549317 solute carrier
family 2 (facilitated glucose transporter), member 12
{Mus musculus;} , partial (5%)
Length = 992
Score = 341 bits (874), Expect = 5e-94
Identities = 175/205 (85%), Positives = 186/205 (90%)
Frame = +1
Query: 326 PEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCM 385
PEI AAGIED SKLLAATVAVG+ KT+FILVAI+LIDK+GRKPLL+ STIGMT CLFCM
Sbjct: 1 PEIFQAAGIEDNSKLLAATVAVGVAKTIFILVAIILIDKLGRKPLLMISTIGMTVCLFCM 180
Query: 386 GVTLSLFEKGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVAN 445
G TL+L KG IAL ILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVAN
Sbjct: 181 GATLALLGKGSFAIALAILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVAN 360
Query: 446 RVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENE 505
RVCSGLVAMSFLSVS+AIS GTFF+F+AISALAI FV TLVPETKGKSLEQIEMMF+NE
Sbjct: 361 RVCSGLVAMSFLSVSEAISVAGTFFVFAAISALAIAFVVTLVPETKGKSLEQIEMMFQNE 540
Query: 506 HGSQGKEMELGDVEQLVQNKTGLTN 530
+ QGKEMELGDVEQLVQNKTGLTN
Sbjct: 541 YEIQGKEMELGDVEQLVQNKTGLTN 615
>TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 2338
Score = 290 bits (743), Expect = 8e-79
Identities = 155/325 (47%), Positives = 211/325 (64%), Gaps = 14/325 (4%)
Frame = +3
Query: 194 IMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSV 253
I++GY+SNY FS L++ + WR+ML VG +PS+ IG A+ +PESPRWLV + R+ EA+ V
Sbjct: 3 ILIGYISNYGFSKLALRLGWRLMLGVGAIPSILIGVAVLAMPESPRWLVAKGRLGEAKRV 182
Query: 254 LLKTNEDEKEVEERLAEIQQAAGFAN---------SGKYEDKPVWREL-LSPPPALRRML 303
L K +E E+E RLA+I+ AG S + VWREL L P PA+R +
Sbjct: 183 LYKISESEEEARLRLADIKDTAGIPQDCDDDVVLVSKQTHGHGVWRELFLHPTPAVRHIF 362
Query: 304 ITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLID 363
I LGI F Q +GIDA V YSP I AGI+ + L ATVAVG KTV ILVA +D
Sbjct: 363 IASLGIHFFAQATGIDAVVLYSPRIFEKAGIKSDNYRLLATVAVGFVKTVSILVATFFLD 542
Query: 364 KVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLV----IALGILFVCGNVAFFSVGLGP 419
+ GR+ LL+ S G+ L +G++L++ + + L I V VA FS+G GP
Sbjct: 543 RAGRRVLLLCSVSGLILSLLTLGLSLTVVDHSQTTLNWAVGLSIAAVLSYVATFSIGSGP 722
Query: 420 VCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALA 479
+ WV +SEIFPLR+RAQ A+GA NRV SG++AM+FLS+ AI+ GG FFLF+ ++A+A
Sbjct: 723 ITWVYSSEIFPLRLRAQGVAIGAAVNRVTSGVIAMTFLSLQKAITIGGAFFLFAGVAAVA 902
Query: 480 IVFVFTLVPETKGKSLEQIEMMFEN 504
+F +TL+PET+GK+LE+IE F N
Sbjct: 903 WIFHYTLLPETRGKTLEEIEKSFGN 977
>TC218649 similar to UP|Q39416 (Q39416) Integral membrane protein, partial
(93%)
Length = 1695
Score = 233 bits (595), Expect = 1e-61
Identities = 147/458 (32%), Positives = 240/458 (52%), Gaps = 10/458 (2%)
Frame = +2
Query: 55 VIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSLGGGR 114
V AC + +L + G+ G S I DL ++ + + ++ +++G++ G+
Sbjct: 173 VFACVLIVALGPIQFGFTAGYTSPTQSAIINDLGLSVSEFSLFGSLSNVGAMVGAIASGQ 352
Query: 115 TSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEIS 174
++ IGRK ++ +A++ +G + ++ A L +GRLL G G+G P+YIAEIS
Sbjct: 353 IAEYIGRKGSLMIASIPNIIGWLAISFAKDSSFLYMGRLLEGFGVGIISYTVPVYIAEIS 532
Query: 175 PNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFII 234
P RG L + ++ + +GIML Y+ L + + WR++ +GILP + ALF I
Sbjct: 533 PPNLRGGLVSVNQLSVTIGIMLAYL-------LGIFVEWRILAIIGILPCTILIPALFFI 691
Query: 235 PESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSG---KYEDKPVWRE 291
PESPRWL EE + L + ++ + EI++A N+ ++ D R
Sbjct: 692 PESPRWLAKMGMTEEFETSLQVLRGFDTDISVEVNEIKRAVASTNTRITVRFADLKQRRY 871
Query: 292 LLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITK 351
L L+ G+G+ QQ+SGI+ ++YS I AGI AAT VG +
Sbjct: 872 WLP--------LMIGIGLLILQQLSGINGVLFYSSTIFRNAGISSSD---AATFGVGAVQ 1018
Query: 352 TVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTL----SLFEKGPLVIALGILFVC 407
+ + + L DK GR+ LLI S GM+ L + +T S+ E L L L +
Sbjct: 1019VLATSLTLWLADKSGRRLLLIVSATGMSFSLLVVAITFYIKASISETSSLYGILSTLSLV 1198
Query: 408 GNVAF---FSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAIS 464
G VA FS+G+G + W++ SEI P+ ++ A ++ +AN + S LV ++ + D S
Sbjct: 1199GVVAMVIAFSLGMGAMPWIIMSEILPINIKGLAGSVATLANWLFSWLVTLTANMLLD-WS 1375
Query: 465 FGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
GGTF +++ + AL +VFV VPETKGK++E+I+ F
Sbjct: 1376SGGTFTIYAVVCALTVVFVTIWVPETKGKTIEEIQWSF 1489
>TC205502 homologue to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(42%)
Length = 659
Score = 182 bits (463), Expect = 2e-46
Identities = 103/214 (48%), Positives = 135/214 (62%), Gaps = 14/214 (6%)
Frame = +1
Query: 234 IPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANS---------GKYE 284
+PESPRWLVM+ R+ EAR VL KT++ +E + RLAEI+QAAG S +
Sbjct: 16 MPESPRWLVMRGRLGEARKVLNKTSDSREEAQLRLAEIKQAAGIPESCNDDVVQVTKRST 195
Query: 285 DKPVWREL-LSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAA 343
+ VW+EL L P P +R ++I LGI FQQ SG+DA V YSP I AGI+D + L A
Sbjct: 196 GEGVWKELFLYPTPPIRHIVIAALGIHFFQQASGVDAVVLYSPRIFEKAGIKDDTHKLLA 375
Query: 344 TVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLF----EKGPLVI 399
TVAVG KTVFIL A +D+VGR+PLL++S GM L + ++L++ K +
Sbjct: 376 TVAVGFVKTVFILAATFTLDRVGRRPLLLSSVGGMVLSLLTLAISLTIIGHSERKLMWAV 555
Query: 400 ALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRV 433
AL I V VA FS+G GP+ WV +SEIFPLR+
Sbjct: 556 ALSIAMVLAYVATFSIGAGPITWVYSSEIFPLRL 657
>TC207288 UP|Q7XA51 (Q7XA51) Monosaccharide transporter, complete
Length = 1813
Score = 174 bits (441), Expect = 9e-44
Identities = 131/479 (27%), Positives = 229/479 (47%), Gaps = 25/479 (5%)
Frame = +1
Query: 51 TRKYVIACAIFASLNNVLLGYDVGVMSGAVI---FIKEDL-----------------KIT 90
T + C + A L L GYD+GV G F+KE K
Sbjct: 94 TAYFAFTCVVGA-LGGSLFGYDLGVSGGVPSMDDFLKEFFPKVYRRKQMHLHETDYCKYD 270
Query: 91 EVQVEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMI 150
+ + L +L+ + + GRK ++ + A+ F G I A + +L+I
Sbjct: 271 DQVLTLFTSSLYFSALVMTFFASFLTRKKGRKASIIVGALSFLAGAILNAAAKNIAMLII 450
Query: 151 GRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVH 210
GR+L G GIGFG P+Y++E++P RG++ + GI++ + NY F+
Sbjct: 451 GRVLLGGGIGFGNQAVPLYLSEMAPAKNRGAVNQLFQFTTCAGILIANLVNY-FTEKIHP 627
Query: 211 ISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAE 270
WR+ L + LP+ + E+P LV Q R+++A+ VL + E VE +
Sbjct: 628 YGWRISLGLAGLPAFAMLVGGICCAETPNSLVEQGRLDKAKQVLQRIRGTE-NVEAEFED 804
Query: 271 IQQAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILM 330
+++A+ A + K + + + P +++I LGI FQQ++G ++ ++Y+P I
Sbjct: 805 LKEASEEAQAVKSPFRTLLKRKYRP-----QLIIGALGIPAFQQLTGNNSILFYAPVIFQ 969
Query: 331 AAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLS 390
+ G + L ++ + G V ++++ L+DK GR+ + + M C+ G L+
Sbjct: 970 SLGFGANASLFSSFITNG-ALLVATVISMFLVDKYGRRKFFLEAGFEMICCMIITGAVLA 1146
Query: 391 L-----FEKGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVAN 445
+ E G V A ++ + V + GP+ W++ SE+FPL +R+ A ++ N
Sbjct: 1147VNFGHGKEIGKGVSAFLVVVIFLFVLAYGRSWGPLGWLVPSELFPLEIRSSAQSIVVCVN 1326
Query: 446 RVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFEN 504
+ + LVA FL + F G F LF+++ FVF L+PETK +E+I ++FEN
Sbjct: 1327MIFTALVAQLFLMSLCHLKF-GIFLLFASLIIFMSFFVFFLLPETKKVPIEEIYLLFEN 1500
>TC218581 similar to GB|AAM70554.1|21700861|AY124845 At1g75220/F22H5_6
{Arabidopsis thaliana;} , partial (66%)
Length = 1340
Score = 167 bits (424), Expect = 8e-42
Identities = 112/342 (32%), Positives = 182/342 (52%), Gaps = 11/342 (3%)
Frame = +2
Query: 172 EISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFAL 231
EI+PN G L + ++ I +GIML Y+ L + ++WR++ +GILP + L
Sbjct: 5 EIAPNHLIGGLGSVNQLSITIGIMLAYL-------LGLFVNWRILAILGILPCTVLIPGL 163
Query: 232 FIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWRE 291
F IPESPRWL I+E + L + ++ + EI+++ A++GK R
Sbjct: 164 FFIPESPRWLAKMGMIDEFETSLQVLRGFDTDISVEVHEIKRSV--ASTGK-------RA 316
Query: 292 LLSPPPALRRM----LITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAV 347
+ R+ L+ G+G+ QQ+SGI+ ++YS I AGI AATV +
Sbjct: 317 AIRFADLKRKRYWFPLMVGIGLLVLQQLSGINGILFYSTTIFANAGISSSE---AATVGL 487
Query: 348 GITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSL----FEKGPLVIALGI 403
G + + ++ L+DK GR+ LLI S+ MT L + + L E L LGI
Sbjct: 488 GAVQVIATGISTWLVDKSGRRLLLIISSSVMTVSLLIVSIAFYLEGVVSEDSHLFSILGI 667
Query: 404 LFVCGNVAF---FSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVS 460
+ + G VA FS+GLGP+ W++ SEI P+ ++ A ++ + N + S + M+ ++
Sbjct: 668 VSIVGLVAMVIGFSLGLGPIPWLIMSEILPVNIKGLAGSIATMGNWLISWGITMT-ANLL 844
Query: 461 DAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
S GGTF +++ ++A I F+ VPETKG++LE+I+ F
Sbjct: 845 LNWSSGGTFTIYTVVAAFTIAFIAMWVPETKGRTLEEIQFSF 970
>TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete
Length = 1925
Score = 144 bits (362), Expect = 1e-34
Identities = 130/509 (25%), Positives = 233/509 (45%), Gaps = 42/509 (8%)
Frame = +1
Query: 50 STRKYVIACAIFASLNNVLLGYDVGVMSGAV------------IFIKEDLKITEVQ---- 93
S +V I A++ ++ GYD+G+ G +F K++ T Q
Sbjct: 148 SLTPFVTVTCIVAAMGGLIFGYDIGISGGVTSMDPFLLKFFPSVFRKKNSDKTVNQYCQY 327
Query: 94 ----VEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLM 149
+ L + +LL SL + GRK +M ++F +G + A +L+
Sbjct: 328 DSQTLTMFTSSLYLAALLSSLVASTVTRRFGRKLSMLFGGLLFLVGALINGFAQHVWMLI 507
Query: 150 IGRLLAGIGIGFGVMIS---PIYIAEISPNLTRGSLTTFPEIFINVGIMLGY-VSNYAFS 205
+GR+L G GIGF + PI I + G+L + GI G V +
Sbjct: 508 VGRILLGFGIGFANQVCATLPI*NGFIQI*RSIGTLAFNCQS--QFGIPRGQCVELLLWL 681
Query: 206 GLSVHISWRVMLAVG-ILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEV 264
V W++ + G ++P++ I ++P++P ++ + E+A++ L + + V
Sbjct: 682 KSMVAWGWKIEVWEGAMVPALIITVGSLVLPDTPNSMIERGDREKAKAQLQRIRGIDN-V 858
Query: 265 EERLAEIQQAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYY 324
+E ++ A+ S + P WR LL R L + I FQQ++GI+ ++Y
Sbjct: 859 DEEFNDLVAAS---ESSSQVEHP-WRNLLQRK--YRPHLTMAVLIPFFQQLTGINVIMFY 1020
Query: 325 SPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFC 384
+P + + G +D + L++A + G+ V V+I +DK GR+ L + + M C
Sbjct: 1021APVLFSSIGFKDDAALMSAVIT-GVVNVVATCVSIYGVDKWGRRALFLEGGVQMLICQAV 1197
Query: 385 MGVTLSLF-----EKGPL---VIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQ 436
+ + G L + +LF+C V+ F+ GP+ W++ SEIFPL +R+
Sbjct: 1198VAAAIGAKFGTDGNPGDLPKWYAIVVVLFICIYVSAFAWSWGPLGWLVPSEIFPLEIRSA 1377
Query: 437 ASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIV-FVFTLVPETKGKSL 495
A ++ N + + L+A FL++ + FG FLF A L + FV+ +PETKG +
Sbjct: 1378AQSINVSVNMLFTFLIAQVFLTMLCHMKFG--LFLFFAFFVLIMTFFVYFFLPETKGIPI 1551
Query: 496 EQIEMMFEN--------EHGSQGKEMELG 516
E++ +++ EH G +E+G
Sbjct: 1552EEMGQVWQAHPFWSRFVEHDDYGNGVEMG 1638
>TC213053 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(27%)
Length = 440
Score = 143 bits (360), Expect = 2e-34
Identities = 77/146 (52%), Positives = 101/146 (68%), Gaps = 4/146 (2%)
Frame = +1
Query: 322 VYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTAC 381
V YSPEI AG+E + L ATVAVG KTVFILVA L+D+VGR+PLL+TS GM
Sbjct: 1 VLYSPEIFKKAGLESDGEQLLATVAVGFAKTVFILVATFLLDRVGRRPLLLTSVGGMVFS 180
Query: 382 LFCMGVTLSLFEKGPLV----IALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQA 437
L +G++L++ + V I L I V V+ FSVG GP+ WV +SEIFPLR+RAQ
Sbjct: 181 LLTLGLSLTVIDHSRAVLKWAIGLSIGMVLSYVSTFSVGAGPITWVYSSEIFPLRLRAQG 360
Query: 438 SALGAVANRVCSGLVAMSFLSVSDAI 463
+A+G V NRV SG+++M+FLS+S+ I
Sbjct: 361 AAMGVVVNRVTSGVISMTFLSLSNKI 438
>TC206001 similar to UP|Q8VZI5 (Q8VZI5) AT5g18840/F17K4_90, partial (66%)
Length = 951
Score = 134 bits (337), Expect = 1e-31
Identities = 96/317 (30%), Positives = 152/317 (47%), Gaps = 8/317 (2%)
Frame = +3
Query: 148 LMIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGL 207
L +GR G GIG + P+YIAEI+P RG L T ++ I G + ++ L
Sbjct: 39 LDMGRFFTGYGIGVISYVVPVYIAEIAPKNLRGGLATTNQLLIVTGGSVSFL-------L 197
Query: 208 SVHISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEER 267
I+WR + G++P + + L IPESPRWL R +E + L + ++ +
Sbjct: 198 GSVINWRELALAGLVPCICLLVGLCFIPESPRWLAKVGREKEFQLALSRLRGKHADISDE 377
Query: 268 LAEIQQAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPE 327
AEI E P + L ++ G+G+ QQ GI+ +Y+ E
Sbjct: 378 AAEILDYI-----ETLESLPKTKLLDLFQSKYVHSVVIGVGLMACQQSVGINGIGFYTAE 542
Query: 328 ILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGV 387
I +AAG+ S A T+A + F L+ +L+DK GR+PL++ S G L C
Sbjct: 543 IFVAAGL---SSGKAGTIAYACIQIPFTLLGAILMDKSGRRPLVMVSAAG--TFLGCFVA 707
Query: 388 TLSLFEKG--------PLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASA 439
+ F K P++ G+L +A FS+GLG V WV+ SEIFP+ ++ A +
Sbjct: 708 AFAFFLKDQSLLPEWVPILAFAGVLIY---IAAFSIGLGSVPWVIMSEIFPIHLKGTAGS 878
Query: 440 LGAVANRVCSGLVAMSF 456
L + + + +V+ +F
Sbjct: 879 LVVLVAWLGAWVVSYTF 929
>TC218395 similar to UP|Q9FIF2 (Q9FIF2) Sugar transporter-like protein,
partial (40%)
Length = 1013
Score = 128 bits (322), Expect = 6e-30
Identities = 78/243 (32%), Positives = 125/243 (51%), Gaps = 2/243 (0%)
Frame = +1
Query: 259 EDEKEVEERLAEIQQAAGFANSGKYEDKPVWRELLS--PPPALRRMLITGLGIQCFQQIS 316
E EK++EE L ++ Y D+ L P L+ +I G G+ FQQI+
Sbjct: 13 ESEKQIEETLVSLKSV--------YADQESEGNFLEVFQGPNLKAFIIGG-GLVLFQQIT 165
Query: 317 GIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTI 376
G + +YY+ IL +AG S +V +G+ K + +A++ +D +GR+PLLI
Sbjct: 166 GQPSVLYYAGPILQSAGFSAASDATKVSVVIGLFKLLMTWIAVLKVDDLGRRPLLIGGVS 345
Query: 377 GMTACLFCMGVTLSLFEKGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQ 436
G+ L + PLV A+G L + V + + GP+ W++ SE+FPLR R +
Sbjct: 346 GIALSLVLLSAYYKFLGGFPLV-AVGALLLY--VGCYQISFGPISWLMVSEVFPLRTRGK 516
Query: 437 ASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLE 496
+L + N + +V +F + + + F LF AI+ L+++F+ VPETKG SLE
Sbjct: 517 GISLAVLTNFASNAVVTFAFSPLKEFLGAENLFLLFGAIATLSLLFIIFSVPETKGMSLE 696
Query: 497 QIE 499
IE
Sbjct: 697 DIE 705
>AW705246
Length = 484
Score = 127 bits (319), Expect = 1e-29
Identities = 66/154 (42%), Positives = 99/154 (63%), Gaps = 4/154 (2%)
Frame = +2
Query: 344 TVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLV----I 399
+VAVG KTV I VA + +D+ GR+ LL+ + G+ L +G++L++ + +
Sbjct: 11 SVAVGFDKTVCIWVATIFLDRAGRRVLLLCNVSGLILSLMTLGLSLTVVDHSQTTLNCAV 190
Query: 400 ALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSV 459
L I V VA FS+ GP+ WV SEIFPLR+ A A+GA N+V SG++AM+FL +
Sbjct: 191 GLRIDSVLSYVATFSIWSGPITWVYISEIFPLRLWAHGVAIGAAVNKVTSGVIAMTFLYL 370
Query: 460 SDAISFGGTFFLFSAISALAIVFVFTLVPETKGK 493
AI+ GG FFLF+ ++A+A +F +TL+PE +GK
Sbjct: 371 QKAITIGGAFFLFAGVAAVAWIFHYTLIPEARGK 472
>TC229858 weakly similar to UP|Q8VHD6 (Q8VHD6) Glucose transporter 10 (Mus
musculus 13 days embryo heart cDNA, RIKEN full-length
enriched library, clone:D330001H08 product:solute
carrier family 2 (facilitated glucose transporter),
member 10, full insert sequence), partial (4%)
Length = 715
Score = 118 bits (295), Expect = 7e-27
Identities = 66/206 (32%), Positives = 106/206 (51%), Gaps = 2/206 (0%)
Frame = +3
Query: 299 LRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVA 358
+R + G G+ FQQ +GI+ +YYSP I+ AG L ++ V ++
Sbjct: 6 IRLAFLVGAGLLAFQQFTGINTVMYYSPTIVQMAGFHANELALLLSLIVAGMNAAGTILG 185
Query: 359 IVLIDKVGRKPLLITSTIGMTACLFCMGVTL--SLFEKGPLVIALGILFVCGNVAFFSVG 416
I LID GRK L ++S G+ L + L L ++ + + FFS G
Sbjct: 186 IYLIDHAGRKKLALSSLGGVIVSLVILAFAFYKQSSTSNELYGWLAVVGLALYIGFFSPG 365
Query: 417 LGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAIS 476
+GPV W L+SEI+P R + A V + +V+ +FLS+++ I G TF + I+
Sbjct: 366 MGPVPWTLSSEIYPEEYRGICGGMSATVCWVSNLIVSETFLSIAEGIGIGSTFLIIGVIA 545
Query: 477 ALAIVFVFTLVPETKGKSLEQIEMMF 502
+A VFV VPETKG + +++E+++
Sbjct: 546 VVAFVFVLVYVPETKGLTFDEVEVIW 623
>TC231818 weakly similar to UP|Q94CI6 (Q94CI6) Sugar-porter family protein 2,
partial (7%)
Length = 595
Score = 112 bits (280), Expect = 4e-25
Identities = 60/82 (73%), Positives = 70/82 (85%), Gaps = 2/82 (2%)
Frame = +2
Query: 451 LVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQG 510
LV +SFLSVS AI+ G FF+F+AIS+LAIVFV+ LVPETKGKSLEQIE+MF+NEH +G
Sbjct: 2 LVDLSFLSVSRAITVAGAFFVFAAISSLAIVFVYMLVPETKGKSLEQIEIMFKNEHEREG 181
Query: 511 KEMELGD--VEQLVQNKTGLTN 530
EMELGD VEQLVQ+KT LTN
Sbjct: 182 SEMELGDVEVEQLVQDKTVLTN 247
>TC203480 similar to UP|Q8LPQ8 (Q8LPQ8) AT4g35300/F23E12_140, partial (38%)
Length = 1040
Score = 110 bits (276), Expect = 1e-24
Identities = 78/231 (33%), Positives = 120/231 (51%), Gaps = 15/231 (6%)
Frame = +2
Query: 287 PVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVA 346
P W+ LL P ++ L+ G+GIQ QQ SGI+ +YY+P+IL AG+E +L + +
Sbjct: 167 PSWKALLEP--GVKHALVVGVGIQILQQFSGINGVLYYTPQILEEAGVE----VLLSDIG 328
Query: 347 VGITKTVFIL-------------VAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE 393
+G F++ VA+ L+D GR+ LL+T+ + L + V SL
Sbjct: 329 IGSESASFLISAFTTFLMLPCIGVAMKLMDVSGRRQLLLTTIPVLIVSLIIL-VIGSLVN 505
Query: 394 KGPLVIALGILFVCGNVAF--FSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGL 451
G + A I VC V F F +G GP+ +L SEIFP RVR A+ A+ + +
Sbjct: 506 FGNVAHA-AISTVCVVVYFCCFVMGYGPIPNILCSEIFPTRVRGLCIAICALVFWIGDII 682
Query: 452 VAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
+ S + ++ GG F +++ + ++ +FVF VPETKG LE I F
Sbjct: 683 ITYSLPVMLGSLGLGGVFAIYAVVCFISWIFVFLKVPETKGMPLEVISEFF 835
>TC218012 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporter 4, partial
(54%)
Length = 1142
Score = 110 bits (276), Expect = 1e-24
Identities = 77/279 (27%), Positives = 140/279 (49%), Gaps = 8/279 (2%)
Frame = +3
Query: 236 ESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWRELLSP 295
++P L+ + R+EE ++VL K + +E E+ +A+ A K+ +R LL
Sbjct: 3 DTPNSLIERGRLEEGKTVLKKIRGTDN-IELEFQELVEASRVAKEVKHP----FRNLLKR 167
Query: 296 PPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFI 355
R L+ + +Q FQQ +GI+A ++Y+P + G ++ + L +A + G +
Sbjct: 168 RN--RPQLVISIALQIFQQFTGINAIMFYAPVLFNTLGFKNDASLYSAVIT-GAVNVLST 338
Query: 356 LVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSL--------FEKGPLVIALGILFVC 407
+V+I +DK+GR+ LL+ + + M + + L + KG + L ++ VC
Sbjct: 339 VVSIYSVDKLGRRMLLLEAGVQMFLSQVVIAIILGIKVTDHSDDLSKG--IAILVVVMVC 512
Query: 408 GNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGG 467
V+ F+ GP+ W++ SE FPL R+ ++ N + + ++A +FLS+ F G
Sbjct: 513 TFVSSFAWSWGPLGWLIPSETFPLETRSAGQSVTVCVNLLFTFVIAQAFLSMLCHFKF-G 689
Query: 468 TFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEH 506
F FS + VFV L+PETK +E++ +H
Sbjct: 690 IFLFFSGWVLVMSVFVLFLLPETKNVPIEEMTERVWKQH 806
>TC208766 similar to UP|STA_RICCO (Q10710) Sugar carrier protein A, partial
(55%)
Length = 1202
Score = 110 bits (274), Expect = 2e-24
Identities = 67/288 (23%), Positives = 143/288 (49%), Gaps = 7/288 (2%)
Frame = +1
Query: 222 LPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSG 281
+P++ + +P++P L+ + E+ R +L K KEV+ ++ A+ A S
Sbjct: 4 VPALLMTVGGIFLPDTPNSLIERGLAEKGRKLLEKIR-GTKEVDAEFQDMVDASELAKSI 180
Query: 282 KYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLL 341
K+ + + P L+ + + FQ ++GI++ ++Y+P + + G + L+
Sbjct: 181 KHPFRNILERRYRPE------LVMAIFMPTFQILTGINSILFYAPVLFQSMGFGGDASLI 342
Query: 342 AATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSL-------FEK 394
++ + G+ + ++I +D++GR+ LL++ + M C + + L + K
Sbjct: 343 SSALTGGVLASS-TFISIATVDRLGRRVLLVSGGLQMITCQIIVAIILGVKFGADQELSK 519
Query: 395 GPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAM 454
G ++ ++ +C V F GP+ W + SEIFPL +R+ + N + + ++A
Sbjct: 520 GFSILV--VVVICLFVVAFGWSWGPLGWTVPSEIFPLEIRSAGQGITVAVNLLFTFIIAQ 693
Query: 455 SFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
+FL++ + F G F F+ + +FV+ +PETKG +E++ M+
Sbjct: 694 AFLALLCSFKF-GIFLFFAGWITIMTIFVYLFLPETKGIPIEEMSFMW 834
>TC208439 weakly similar to UP|O04078 (O04078) Monosaccharid transport
protein, partial (43%)
Length = 936
Score = 108 bits (269), Expect = 8e-24
Identities = 66/225 (29%), Positives = 116/225 (51%), Gaps = 10/225 (4%)
Frame = +1
Query: 300 RRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAI 359
R L+ + I FQQ +G++ +Y+P + G + L++A + +G K V LV+I
Sbjct: 1 RPQLVFAICIPFFQQFTGLNVITFYAPILFRTIGFGSGASLMSAVI-IGSFKPVSTLVSI 177
Query: 360 VLIDKVGRKPLLITSTIGMTACLFCMGVTLSL----------FEKGPLVIALGILFVCGN 409
+L+DK GR+ L + M C M + +++ K ++ +GI +C
Sbjct: 178 LLVDKFGRRTLFLEGGAQMLICQIIMTIAIAVTFGTNGNPGTLPKWYAIVVVGI--ICVY 351
Query: 410 VAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTF 469
V+ F+ GP+ W++ SEIFPL +R A ++ N + + +A F S+ + F G F
Sbjct: 352 VSGFAWSWGPLGWLIPSEIFPLEIRPAAQSITVGVNMISTFFIAQFFTSMLCHMKF-GLF 528
Query: 470 FLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGKEME 514
F + F++ L+PETKG LE++ M+++ +H GK +E
Sbjct: 529 IFFGCFVVIMTTFIYKLLPETKGIPLEEMSMVWQ-KHPIWGKFLE 660
>TC208908 weakly similar to UP|Q8LBI9 (Q8LBI9) Sugar transporter-like
protein, partial (28%)
Length = 753
Score = 104 bits (260), Expect = 9e-23
Identities = 57/164 (34%), Positives = 97/164 (58%), Gaps = 8/164 (4%)
Frame = +2
Query: 344 TVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEK--------G 395
T+A+ K + ++L+DK GR+PLL+ S +G C+ C LS +
Sbjct: 2 TIAIVAVKIPMTTIGVLLMDKSGRRPLLLVSAVG--TCVGCFLAALSFVLQDLHKWKGVS 175
Query: 396 PLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMS 455
P++ +G+L G+ +S+G+G + WV+ SEIFP+ V+ A +L + + +CS +++ S
Sbjct: 176 PILALVGVLVYVGS---YSIGMGAIPWVIMSEIFPINVKGSAGSLVTLVSWLCSWIISYS 346
Query: 456 FLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
F + + S GTF +FS+I ++FV LVPETKG++LE+I+
Sbjct: 347 F-NFLMSWSSAGTFLMFSSICGFTVLFVAKLVPETKGRTLEEIQ 475
>AI461030
Length = 271
Score = 101 bits (252), Expect = 7e-22
Identities = 50/88 (56%), Positives = 67/88 (75%)
Frame = +2
Query: 418 GPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISA 477
GP+C V+ EIFPLR+RAQASAL A +RV SG ++M FLSVS AI+ GTFF+F A+S
Sbjct: 2 GPICCVMAYEIFPLRLRAQASALVAAGSRVSSGAISM*FLSVSRAITEAGTFFVFGAVSC 181
Query: 478 LAIVFVFTLVPETKGKSLEQIEMMFENE 505
A+ F VPET+GK++E+IE++F +E
Sbjct: 182 CAVAFDHYCVPETRGKTVEEIEVLFRDE 265
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.140 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,861,353
Number of Sequences: 63676
Number of extensions: 277527
Number of successful extensions: 1558
Number of sequences better than 10.0: 114
Number of HSP's better than 10.0 without gapping: 1472
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1487
length of query: 530
length of database: 12,639,632
effective HSP length: 102
effective length of query: 428
effective length of database: 6,144,680
effective search space: 2629923040
effective search space used: 2629923040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126008.6


