

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126008.14 + phase: 0 /pseudo/partial
(731 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
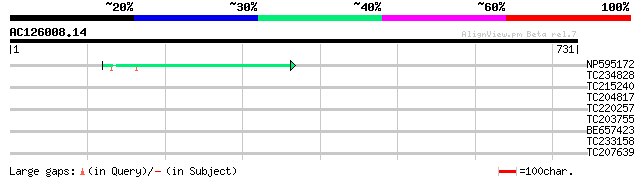
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 47 4e-05
TC234828 42 0.001
TC215240 homologue to UP|RK28_ARATH (O22795) 50S ribosomal prote... 34 0.20
TC204817 similar to UP|MYO3_LYCES (P54928) Inositol-1(or 4)-mono... 33 0.59
TC220257 similar to GB|AAB62842.1|2252843|IG005I10 A_IG005I10.24... 31 2.2
TC203755 weakly similar to UP|Q9LG50 (Q9LG50) ESTs AU078742(C118... 30 3.8
BE657423 30 3.8
TC233158 similar to UP|Q9LRC7 (Q9LRC7) BZIP transcriptional acti... 29 6.5
TC207639 similar to UP|O48592 (O48592) GT2 protein, partial (24%) 27 6.8
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 46.6 bits (109), Expect = 4e-05
Identities = 74/270 (27%), Positives = 105/270 (38%), Gaps = 21/270 (7%)
Frame = +2
Query: 120 KSFLRFFQRIL----PSYRWRGKLSLQLIWYQARVLFRLRRIRCLH-------------- 161
K+ R +Q+IL PSY L L+ + YQ R RI H
Sbjct: 1643 KTL*RIYQQILILNWPSY---STLMLKCLQYQHHCHLRGNRIMLYH*NKGQAPLK*DHTD 1813
Query: 162 ---QSWVN*RISWKSCLRSSLLDQVFHRGVLQCCWLRRRMEA*GCVWIIGS*TR*RLRIG 218
+ + +K C + +QV C LRR+ME II * + RI
Sbjct: 1814 ILTHKRIRLKR*YKKCWFKASFNQVTVHFPYPYCLLRRKMEVGDSAQIIEL*MQSL*RIA 1993
Query: 219 IHFQGLMI*WIS*LVLRCLVRLI*DQDIIRLE*RRKIYPKLRSERGMDTMSIL*CHLEYL 278
Q M ++ + L I* QD I+ +I KL E M M+ *CHL L
Sbjct: 1994 FQCQQWMNC*MNYMELNIFQNWI*GQDTIKFWYSLRIERKLLLEHIMAIMNG**CHLVLL 2173
Query: 279 MRQVSLWNI*IEYFILIWTVLWWCS*MTFWCIRSLRKSMLSI*GLCCKS*KITGCVPSCL 338
M Q * +YF L+ L+ C MT I L + + S LC + * + C
Sbjct: 2174 MHQPHSNA**TKYFSLLSENLFLCFLMTS*YIVLLGRIISSTWNLCYRP*SNISYLLDCP 2353
Query: 339 SVSFG*RK*VSLVM*FLRVVFLWILPRLMQ 368
+V +K ++ +L F W + R Q
Sbjct: 2354 NVHLEIQKLITWATRYLG*EFPWRIQRYKQ 2443
>TC234828
Length = 857
Score = 42.0 bits (97), Expect = 0.001
Identities = 24/74 (32%), Positives = 39/74 (52%)
Frame = +1
Query: 114 VICR*CKSFLRFFQRILPSYRWRGKLSLQLIWYQARVLFRLRRIRCLHQSWVN*RISWKS 173
V+ R C++F +FF+ + + R K + L W R+ RLR I CL + *R +++
Sbjct: 625 VVYRWCQNFQKFFRMMFVNCHLREKWNSLLTWCLGRIQCRLRLIECLRWN*QR*RHKYRT 804
Query: 174 CLRSSLLDQVFHRG 187
++L Q HRG
Sbjct: 805 F*ANNLFVQAHHRG 846
>TC215240 homologue to UP|RK28_ARATH (O22795) 50S ribosomal protein L28,
chloroplast precursor, partial (60%)
Length = 778
Score = 34.3 bits (77), Expect = 0.20
Identities = 18/43 (41%), Positives = 23/43 (52%), Gaps = 2/43 (4%)
Frame = -2
Query: 362 ILPRLMQCCSGS--LLSLFLRLEAFLVWPVIIGGSLRDSLSWR 402
+LP C GS +SL L+ L WP +I RD+LSWR
Sbjct: 333 LLPTKFSCTEGSQTAVSLSCGLKMILCWPCLISFQ*RDTLSWR 205
>TC204817 similar to UP|MYO3_LYCES (P54928) Inositol-1(or 4)-monophosphatase
3 (IMPase 3) (IMP 3) (Inositol monophosphatase 3) ,
partial (94%)
Length = 1176
Score = 32.7 bits (73), Expect = 0.59
Identities = 17/51 (33%), Positives = 25/51 (48%)
Frame = +1
Query: 276 EYLMRQVSLWNI*IEYFILIWTVLWWCS*MTFWCIRSLRKSMLSI*GLCCK 326
E+L+R SLWN + ++ T LWW + WC R I* + C+
Sbjct: 721 EWLLRSESLWNCMWKAGCIL*TWLWWSLGCSRWCCHC*RSWRCCI*SVRCR 873
>TC220257 similar to GB|AAB62842.1|2252843|IG005I10 A_IG005I10.24
{Arabidopsis thaliana;} , partial (4%)
Length = 573
Score = 30.8 bits (68), Expect = 2.2
Identities = 15/34 (44%), Positives = 25/34 (73%), Gaps = 1/34 (2%)
Frame = +2
Query: 434 *LQC*FCRI-RRNHLLCIVMLLRWVLVVCLCRIV 466
*+ C CR R++HLL +++LL W+L++ +C IV
Sbjct: 221 *MLCHLCR*GRQSHLLKLLLLLLWILLLGVCLIV 322
>TC203755 weakly similar to UP|Q9LG50 (Q9LG50) ESTs AU078742(C11888), partial
(5%)
Length = 1460
Score = 30.0 bits (66), Expect = 3.8
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = +1
Query: 634 LDWVRMVF*CFVIEFVCRMCLSLRGKFWMKVIEVV*VFIQE 674
L+W+R+V +CR C G+FW K+ + VFI+E
Sbjct: 913 LEWLRIVG----CPELCRKCQPHVGEFWSKISHIKEVFIEE 1023
>BE657423
Length = 737
Score = 30.0 bits (66), Expect = 3.8
Identities = 13/34 (38%), Positives = 19/34 (55%)
Frame = -2
Query: 276 EYLMRQVSLWNI*IEYFILIWTVLWWCS*MTFWC 309
E+L+R SLWN + ++ T LWW + WC
Sbjct: 448 EWLLRSESLWNCMWKAGCIL*TWLWWSLGCSRWC 347
>TC233158 similar to UP|Q9LRC7 (Q9LRC7) BZIP transcriptional activator RSG,
partial (27%)
Length = 874
Score = 29.3 bits (64), Expect = 6.5
Identities = 10/26 (38%), Positives = 18/26 (68%)
Frame = +3
Query: 136 RGKLSLQLIWYQARVLFRLRRIRCLH 161
RG+ Q+ W + +LFRL+++ C+H
Sbjct: 69 RGRFVTQVSWRRRCILFRLKQLTCVH 146
>TC207639 similar to UP|O48592 (O48592) GT2 protein, partial (24%)
Length = 1689
Score = 27.3 bits (59), Expect(2) = 6.8
Identities = 11/40 (27%), Positives = 21/40 (52%), Gaps = 4/40 (10%)
Frame = -1
Query: 290 EYFILIWTVLWW----CS*MTFWCIRSLRKSMLSI*GLCC 325
++ + +W++LWW C+ + W +R + GLCC
Sbjct: 324 QWSLWLWSLLWWWLYYCNFLWRWLLREI--------GLCC 229
Score = 20.0 bits (40), Expect(2) = 6.8
Identities = 13/45 (28%), Positives = 18/45 (39%)
Frame = -2
Query: 357 VVFLWILPRLMQCCSGSLLSLFLRLEAFLVWPVIIGGSLRDSLSW 401
VV W P CC ++ + FL L W L ++SW
Sbjct: 233 VVEFWC*PD--NCCKKAITAAFLAAAVDLSWTRSSCSLLILAISW 105
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.376 0.169 0.715
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,114,208
Number of Sequences: 63676
Number of extensions: 815526
Number of successful extensions: 15847
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 5008
Number of HSP's successfully gapped in prelim test: 677
Number of HSP's that attempted gapping in prelim test: 10329
Number of HSP's gapped (non-prelim): 6648
length of query: 731
length of database: 12,639,632
effective HSP length: 104
effective length of query: 627
effective length of database: 6,017,328
effective search space: 3772864656
effective search space used: 3772864656
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.5 bits)
S2: 63 (28.9 bits)
Medicago: description of AC126008.14


