

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
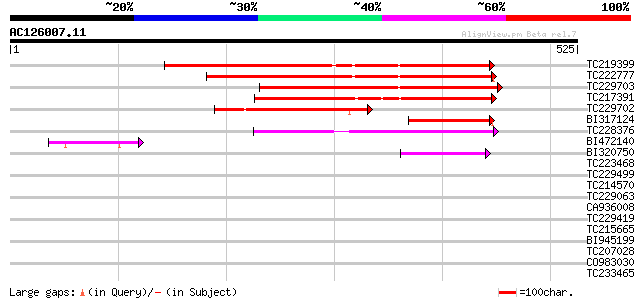
Query= AC126007.11 + phase: 0
(525 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC219399 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g3... 293 1e-79
TC222777 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g3... 251 4e-67
TC229703 230 1e-60
TC217391 similar to UP|Q6T283 (Q6T283) Predicted protein, partia... 199 3e-51
TC229702 150 1e-36
BI317124 95 9e-20
TC228376 similar to UP|Q9ZW00 (Q9ZW00) T25N20.3, partial (14%) 87 2e-17
BI472140 62 6e-10
BI320750 55 1e-07
TC223468 similar to GB|AAS92337.1|46402470|BT012421 At5g24330 {A... 41 0.001
TC229499 38 0.010
TC214570 homologue to PIR|T05766|T05766 peptidylprolyl isomerase... 37 0.016
TC229063 similar to GB|AAO64869.1|29028980|BT005934 At2g19260 {A... 37 0.022
CA936008 36 0.048
TC229419 UP|Q6PIN8 (Q6PIN8) TAO1 protein (Fragment), partial (5%) 34 0.14
TC215665 homologue to UP|Q42792 (Q42792) Asparagine synthetase ... 34 0.18
BI945199 34 0.18
TC207028 similar to UP|Q9LW95 (Q9LW95) KED, partial (13%) 33 0.24
CO983030 33 0.31
TC233465 similar to UP|Q9VAQ7 (Q9VAQ7) CG1420-PA, partial (3%) 31 1.5
>TC219399 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g36720
{Arabidopsis thaliana;} , partial (29%)
Length = 1270
Score = 293 bits (751), Expect = 1e-79
Identities = 149/306 (48%), Positives = 203/306 (65%)
Frame = +1
Query: 144 VESPPKEKWYCEYCRNKLQKDKNVEHKENVVTTQKIIESDPSEQIAKICTLSVKHKEVEH 203
+ S P+ WYC++C+N Q++K V H N V ++ DP EQIA C VK E +
Sbjct: 16 LSSIPRGDWYCQFCQNMFQREKFVAHNANAVAAGRVEGVDPIEQIANRCIRIVKDIEADL 195
Query: 204 SSCALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYD 263
SSCALC F+ F P T+++CDQCEK+YHVGCL+DH MA LK++P+ W C DC
Sbjct: 196 SSCALCRGVDFSRSGFGPRTIILCDQCEKEYHVGCLRDHKMAYLKELPEGNWLCCNDCTR 375
Query: 264 IHMKLKNFMARGDVLLSDSLLSLIKNKKEQKGLETEFGLDIKWKVFNRQLIVSKIITSSL 323
IH L+N + +G L +SLL +IK K+E+KGLE +D++W++ N ++ + T L
Sbjct: 376 IHSTLENLLVKGAERLPESLLGVIKKKQEEKGLEPI--IDVRWRLLNGKIASPE--TRPL 543
Query: 324 LSDVVTIFHEQFDSIVVTGTKIDLIPAMVKGRKIKDKYYFGGMYCAVLIVNQVVVSAGIF 383
L + V+IFHE F+ IV + DLIPAMV GR ++ + FGGMYCA+LIVN VVSAG+
Sbjct: 544 LLEAVSIFHECFNPIVDAASGRDLIPAMVYGRNVRGQ-EFGGMYCALLIVNSSVVSAGML 720
Query: 384 RVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKF 443
R+FG +VAEL L+AT +G+F+ L SCIE +L L V+ LVLPAA EAES+W DKF
Sbjct: 721 RIFGSDVAELPLVATSNGNHGKGYFQTLFSCIERLLAFLNVKNLVLPAAEEAESIWTDKF 900
Query: 444 GFTEPN 449
GF++ N
Sbjct: 901 GFSKMN 918
>TC222777 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g36720
{Arabidopsis thaliana;} , partial (26%)
Length = 1071
Score = 251 bits (642), Expect = 4e-67
Identities = 132/270 (48%), Positives = 179/270 (65%), Gaps = 2/270 (0%)
Frame = +1
Query: 183 DPSEQIAKICTLSVKHKEVEHSSCALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDH 242
DP EQIAK C VK E C LC F+ F P T++ICDQCEK+YHVGCL+DH
Sbjct: 4 DPIEQIAKRCIRIVKDIGAEMGGCVLCRSSDFSRSGFGPRTIIICDQCEKEYHVGCLRDH 183
Query: 243 NMANLKKVPKHYWFCGVDCYDIHMKLKNFMARGDVLLSDSLLSLIKNKKEQKGLETEFGL 302
MA LK++P+ WFC DC IH L+N + R L +SLL +IK K+ + LE +
Sbjct: 184 KMAYLKELPEGDWFCCNDCTRIHSTLENLLIRVAERLPESLLDVIKKKQVGRCLEPLNEI 363
Query: 303 DIKWKVFNRQLIVSKIITSSLLSDVVTIFHEQFDSIVVTGTKIDLIPAMVKGRKIKDKYY 362
D++WK+ N ++ + T LL + V++FHE FD IV DLIPAMV GR ++ +
Sbjct: 364 DVRWKLLNGKIASPE--TRPLLLEAVSMFHECFDPIVDPAAGRDLIPAMVYGRNLQTQ-D 534
Query: 363 FGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKEL 422
FGGMYCA+LIVN VVSAG+ R+FG+++AEL L+AT+ + + +G+F+ L +CIE +L L
Sbjct: 535 FGGMYCALLIVNSSVVSAGMVRIFGRDIAELPLVATRYKNRGKGYFQTLFACIERLLAFL 714
Query: 423 KVERLVLPAAHEAESMWIDKFGFT--EPNQ 450
V+ LVLPAA EA S+W +KFGF+ +PNQ
Sbjct: 715 NVKNLVLPAAEEAASIWTEKFGFSKMKPNQ 804
>TC229703
Length = 1172
Score = 230 bits (587), Expect = 1e-60
Identities = 118/225 (52%), Positives = 154/225 (68%)
Frame = +2
Query: 232 KDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIHMKLKNFMARGDVLLSDSLLSLIKNKK 291
K+YHVGCLKDHNM NL+++P WFC +C IH L + +A + + D LL+LIK K
Sbjct: 2 KEYHVGCLKDHNMENLEELPVGNWFCSGNCSQIHTALMDLVASKEKDVPDPLLNLIKKKH 181
Query: 292 EQKGLETEFGLDIKWKVFNRQLIVSKIITSSLLSDVVTIFHEQFDSIVVTGTKIDLIPAM 351
E+K L+ GLD+KW+V N +L + T LLS V IFHE+FD IV + + D IP M
Sbjct: 182 EEKSLDIGAGLDVKWRVINWKLDSDSVETRKLLSKAVAIFHERFDPIVDSTSGRDFIPTM 361
Query: 352 VKGRKIKDKYYFGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCL 411
+ GR I+ + F G+YCAVL VN +VSAG+FRVFG E+AEL L+AT A++Q QG+F+CL
Sbjct: 362 LFGRNIRGQ-DFSGIYCAVLTVNGDIVSAGVFRVFGLEIAELPLVATTADHQGQGYFQCL 538
Query: 412 LSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTEPNQGLGRRY 456
SCIE +L L V+ LVLPAA EAES+W KFGFT+ Q +Y
Sbjct: 539 FSCIETLLGSLNVKNLVLPAADEAESIWTGKFGFTKLPQDEINKY 673
>TC217391 similar to UP|Q6T283 (Q6T283) Predicted protein, partial (28%)
Length = 1127
Score = 199 bits (505), Expect = 3e-51
Identities = 108/225 (48%), Positives = 145/225 (64%), Gaps = 1/225 (0%)
Frame = +3
Query: 227 CDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIHMKLKNFMARGDVLLSDSLLSL 286
CDQCEK+YHVGCL+D + L+++PK WFC DC I+ L+N ++ G ++ S L
Sbjct: 15 CDQCEKEYHVGCLRDMGLCELEELPKDKWFCCDDCNRIYAALQNSVSAGAEIIPASFSEL 194
Query: 287 IKNKKEQKGLETEFGL-DIKWKVFNRQLIVSKIITSSLLSDVVTIFHEQFDSIVVTGTKI 345
I K E KGL T + DI+W++ + + + + LLS IF E FD IV +
Sbjct: 195 IIRKHEDKGLCTYGAMNDIQWRILSGKSRYPEHLP--LLSRAAAIFRECFDPIVAISGR- 365
Query: 346 DLIPAMVKGRKIKDKYYFGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQ 405
DLIP MV GR I + FGGMYC VLIVN VVVSAG+ R+FG+ VAEL L+AT +Q +
Sbjct: 366 DLIPVMVYGRNISGQE-FGGMYCIVLIVNYVVVSAGLLRIFGRNVAELPLVATSRAHQGK 542
Query: 406 GFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTEPNQ 450
G+F+ L SCIE +L L VE+LVLPAA +AES+W K GF + ++
Sbjct: 543 GYFQVLFSCIERLLSSLNVEKLVLPAAGDAESIWTKKLGFRKMSE 677
>TC229702
Length = 449
Score = 150 bits (380), Expect = 1e-36
Identities = 77/150 (51%), Positives = 97/150 (64%), Gaps = 3/150 (2%)
Frame = +2
Query: 190 KICTLSVKHKEVEHSSCALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKK 249
K C VK EV+H CALCS +F+ F P TV+ICDQCEK+YHVGCLK+HNM NL+K
Sbjct: 2 KRCIRVVKTVEVDHGGCALCSRPNFSKS-FGPRTVIICDQCEKEYHVGCLKEHNMENLEK 178
Query: 250 VPKHYWFCGVDCYDIHMKLKNFMARGDVLLSDSLLSLIKNKKEQKGLETEFGLDIKWKVF 309
+P+ WFC +C IH L + +A + + D LLSLIK K E+K LE GLD+KW+V
Sbjct: 179 LPEGNWFCSGNCSHIHTALTDLVASKEKDVPDPLLSLIKKKHEEKSLEIGAGLDVKWRVM 358
Query: 310 NRQL---IVSKIITSSLLSDVVTIFHEQFD 336
N +L + T LLS V IFHE+FD
Sbjct: 359 NWKLDSDSDDSVETRKLLSKAVAIFHERFD 448
>BI317124
Length = 419
Score = 94.7 bits (234), Expect = 9e-20
Identities = 49/86 (56%), Positives = 61/86 (69%), Gaps = 6/86 (6%)
Frame = +3
Query: 370 VLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVL 429
VLIVN VVVSAG+ R+FG+ VAEL L+AT +Q +G+F+ L SCIE +L L VE+LVL
Sbjct: 6 VLIVNSVVVSAGLLRIFGRNVAELPLVATSRAHQGKGYFQVLFSCIERLLSSLIVEKLVL 185
Query: 430 PAAHEAESMWIDKFGF------TEPN 449
PAA +AES+W K GF T PN
Sbjct: 186 PAAGDAESIWTKKLGFFCLSTLTHPN 263
>TC228376 similar to UP|Q9ZW00 (Q9ZW00) T25N20.3, partial (14%)
Length = 1340
Score = 87.0 bits (214), Expect = 2e-17
Identities = 63/228 (27%), Positives = 100/228 (43%), Gaps = 1/228 (0%)
Frame = +2
Query: 226 ICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIHMKLKNFMARGDVLLSDSLLS 285
IC+ CEK YH C K+ + FCG +C ++ LK ++ L + S
Sbjct: 47 ICNLCEKKYHDSCTKEMDNLPNNINTSSLSFCGKECKELSEHLKKYLGTKHELEAGFSWS 226
Query: 286 LIKNKKEQKGLETEFGLDIKWKVFNRQLIVSKIITSSLLSDVVTIFHEQFDSIVVTGTKI 345
LI E I ++ +S L+ +T+ E F ++ + I
Sbjct: 227 LIHRTDEDSEAACRG-------------ISQRVECNSKLAITLTVMDECFLPVIDRRSGI 367
Query: 346 DLIPAMVKGRKIK-DKYYFGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQK 404
+LI ++ + + G Y A L +++A R G ++AE+ I T+ Y++
Sbjct: 368 NLIRNVLYNSGSNFSRLNYSGFYTATLERGDEIIAAASIRFHGTKIAEMPFIGTRHIYRR 547
Query: 405 QGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTEPNQGL 452
QG + L S IE L LKVE+LV+PA E W FGFT ++ L
Sbjct: 548 QGMCRRLFSAIELALCSLKVEKLVIPAIAELTHTWTTVFGFTYLDESL 691
>BI472140
Length = 423
Score = 62.0 bits (149), Expect = 6e-10
Identities = 40/111 (36%), Positives = 56/111 (50%), Gaps = 23/111 (20%)
Frame = +2
Query: 37 KNKKESNLRSNESK---NLQPSPKHTTQTSNSHASPTTINYRDQCLHKLVFQENVLEDGA 93
+ KK+ R+ SK L+ +P + S + S I+ R Q LHKL+F+E+ L +GA
Sbjct: 83 QTKKQWKTRTKSSKLSVKLKTAPITSKCLSPQNKSQWRISKRYQRLHKLIFEEDGLPNGA 262
Query: 94 AVGYFVY--------------------EEVSPSKFEAHAGWASRRKPWPLI 124
V Y+ E+SPS+FE HAGWASRRKP+ I
Sbjct: 263 EVAYYARGQKLLEGIKTCSGIVCRCCNTEISPSQFEVHAGWASRRKPYAFI 415
>BI320750
Length = 428
Score = 54.7 bits (130), Expect = 1e-07
Identities = 29/83 (34%), Positives = 44/83 (52%)
Frame = +3
Query: 363 FGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKEL 422
+ G A+L ++SA R+ G ++AE+ I T+ Y++QG + LL+ +E L L
Sbjct: 21 YSGFVTAILERGDEIISAASIRIRGNQLAEMPFIGTRYMYRRQGMCRRLLNAVEWGLGSL 200
Query: 423 KVERLVLPAAHEAESMWIDKFGF 445
VE LV+PA W FGF
Sbjct: 201 NVELLVIPAISLLTETWSAVFGF 269
>TC223468 similar to GB|AAS92337.1|46402470|BT012421 At5g24330 {Arabidopsis
thaliana;} , partial (33%)
Length = 507
Score = 40.8 bits (94), Expect = 0.001
Identities = 20/53 (37%), Positives = 28/53 (52%)
Frame = +2
Query: 205 SCALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFC 257
SC C H SP +++CD+C++ YH+ CL+ L VPK WFC
Sbjct: 92 SCEECGGGH------SPSKLILCDKCDRGYHLFCLR----PILPSVPKGSWFC 220
>TC229499
Length = 915
Score = 38.1 bits (87), Expect = 0.010
Identities = 27/125 (21%), Positives = 51/125 (40%), Gaps = 26/125 (20%)
Frame = +1
Query: 159 NKLQKDKNVEHKENVVTTQKIIESDPSEQIAKICTLSVKHKEVEHSSCALCSERHFNNGE 218
N+L+++ N + ++T + SD KIC ++V ++V+ + C ++++
Sbjct: 130 NELEENSNCTYDGIQISTDRRNSSD-----CKICKMAVDGEKVKICGHSFCPSKYYHVSC 294
Query: 219 FSP---------W-----------------TVMICDQCEKDYHVGCLKDHNMANLKKVPK 252
S W +++CD C+ YHV C+K +PK
Sbjct: 295 LSSKQLKSYGHCWYCPSCICQVCLTDKDDNKIVLCDACDHAYHVYCMKPPQ----NSIPK 462
Query: 253 HYWFC 257
WFC
Sbjct: 463 GKWFC 477
>TC214570 homologue to PIR|T05766|T05766 peptidylprolyl isomerase M4E13.20 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(13%)
Length = 944
Score = 37.4 bits (85), Expect = 0.016
Identities = 21/60 (35%), Positives = 34/60 (56%), Gaps = 1/60 (1%)
Frame = +2
Query: 7 ELKDNIVVEAVAKDGDDVEKKMKNKKKKKNKNKKESN-LRSNESKNLQPSPKHTTQTSNS 65
E ++ + V+ V ++ +VE + K KKKKK KNK+ S S+E K + KH + +S
Sbjct: 422 EKQEKVKVKKVDEETVEVEVEKKEKKKKKKKNKENSEAASSDEEKTEKKKKKHKDKVEDS 601
>TC229063 similar to GB|AAO64869.1|29028980|BT005934 At2g19260 {Arabidopsis
thaliana;} , partial (10%)
Length = 1656
Score = 37.0 bits (84), Expect = 0.022
Identities = 17/53 (32%), Positives = 25/53 (47%)
Frame = +1
Query: 205 SCALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFC 257
SC +C + + +++CD CE YH+ C LKK+P WFC
Sbjct: 706 SCKICGDLD------NSLNMLLCDHCEDAYHLSCYN----PRLKKLPIDEWFC 834
>CA936008
Length = 421
Score = 35.8 bits (81), Expect = 0.048
Identities = 20/50 (40%), Positives = 26/50 (52%)
Frame = +2
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSNSHASPTTINY 74
+KK K KKKKK K KK+ + + K PK+ TQ N PT N+
Sbjct: 53 KKKKKKKKKKKKKKKKKKKKKKKKKKKGGGGPKN-TQKKNVFRWPTVKNF 199
Score = 30.0 bits (66), Expect = 2.6
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = +1
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESK 50
EKK K KKKKK K KK+ + + K
Sbjct: 46 EKKKKKKKKKKKKKKKKKKKKKKKKK 123
>TC229419 UP|Q6PIN8 (Q6PIN8) TAO1 protein (Fragment), partial (5%)
Length = 895
Score = 34.3 bits (77), Expect = 0.14
Identities = 15/34 (44%), Positives = 18/34 (52%)
Frame = +1
Query: 23 DVEKKMKNKKKKKNKNKKESNLRSNESKNLQPSP 56
D KK K KKKKK K KK+ + + K P P
Sbjct: 655 DFRKKKKKKKKKKKKKKKKKKKKKKKKKKKNPRP 756
>TC215665 homologue to UP|Q42792 (Q42792) Asparagine synthetase , complete
Length = 2492
Score = 33.9 bits (76), Expect = 0.18
Identities = 14/32 (43%), Positives = 21/32 (64%)
Frame = +3
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNLQPSP 56
+KK K KKKKK K KK++ + + K + P+P
Sbjct: 2172 KKKKKKKKKKKKKKKKKN*KKKKKKKKISPAP 2267
Score = 32.3 bits (72), Expect = 0.53
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = +1
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNLQPSP 56
+KK K KKKKK K KK+ + + K P P
Sbjct: 2173 KKKKKKKKKKKKKKKKKIKKKKKKKKKFPPPP 2268
Score = 31.2 bits (69), Expect = 1.2
Identities = 16/39 (41%), Positives = 20/39 (51%)
Frame = +1
Query: 18 AKDGDDVEKKMKNKKKKKNKNKKESNLRSNESKNLQPSP 56
AK +KK K KKKKK K KK+ + + K P P
Sbjct: 2149 AKKKKKKKKKKKKKKKKKKKKKKKKIKKKKKKKKKFPPP 2265
Score = 30.4 bits (67), Expect = 2.0
Identities = 16/41 (39%), Positives = 20/41 (48%)
Frame = +1
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSNS 65
+KK K KKKKK K KK + + K P P + NS
Sbjct: 2176 KKKKKKKKKKKKKKKKIKKKKKKKKKFPPPPPPQGGISENS 2298
>BI945199
Length = 256
Score = 33.9 bits (76), Expect = 0.18
Identities = 17/35 (48%), Positives = 21/35 (59%), Gaps = 2/35 (5%)
Frame = +3
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNL--QPSPK 57
+KK K KKKKK K KK+ + + KNL P PK
Sbjct: 90 KKKKKKKKKKKKKKKKKKKKKKKKKKNLGGGPGPK 194
Score = 30.0 bits (66), Expect = 2.6
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = +2
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESK 50
EKK K KKKKK K KK+ + + K
Sbjct: 80 EKKKKKKKKKKKKKKKKKKKKKKKKK 157
>TC207028 similar to UP|Q9LW95 (Q9LW95) KED, partial (13%)
Length = 490
Score = 33.5 bits (75), Expect = 0.24
Identities = 15/37 (40%), Positives = 25/37 (67%)
Frame = +3
Query: 14 VEAVAKDGDDVEKKMKNKKKKKNKNKKESNLRSNESK 50
V+ +DG++ EKK K+KK+KK K KK+ + ++ K
Sbjct: 15 VKDKGEDGEEKEKKEKDKKEKKKKEKKDKDEETDTLK 125
Score = 32.0 bits (71), Expect = 0.69
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = +3
Query: 17 VAKDGDDVEKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKH 58
V G+D E+K K +K KK K KKE + E+ L+ K+
Sbjct: 15 VKDKGEDGEEKEKKEKDKKEKKKKEKKDKDEETDTLKGKGKN 140
Score = 28.9 bits (63), Expect = 5.9
Identities = 18/60 (30%), Positives = 31/60 (51%)
Frame = +1
Query: 6 EELKDNIVVEAVAKDGDDVEKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSNS 65
+E D + + +G+D E NKKKKK+K +KE + E K+++ K ++ S
Sbjct: 205 DEETDTLKEKGKNDEGEDDEG---NKKKKKDKKEKEKD-HKKEKKDIEEGEKEDSKVEAS 372
>CO983030
Length = 505
Score = 33.1 bits (74), Expect = 0.31
Identities = 15/38 (39%), Positives = 21/38 (54%)
Frame = +3
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQT 62
+KK K KKKKKNK KK ++ + K + K +T
Sbjct: 252 KKKKKKKKKKKNKKKKXXKIKKTQKKKXXKT*KKKXKT 365
Score = 28.9 bits (63), Expect = 5.9
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = +3
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESK 50
+KK K KKKKK KNKK+ + +++
Sbjct: 246 KKKKKKKKKKKKKNKKKKXXKIKKTQ 323
>TC233465 similar to UP|Q9VAQ7 (Q9VAQ7) CG1420-PA, partial (3%)
Length = 719
Score = 30.8 bits (68), Expect = 1.5
Identities = 13/45 (28%), Positives = 23/45 (50%)
Frame = -3
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSNSHASP 69
+KK K KKKKK K KK+ + + + K + ++++P
Sbjct: 714 KKKKKKKKKKKKKKKKKKKKKKKKKNQKKKKKKKKNKKKKTNSTP 580
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,908,751
Number of Sequences: 63676
Number of extensions: 396868
Number of successful extensions: 3369
Number of sequences better than 10.0: 115
Number of HSP's better than 10.0 without gapping: 2996
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3151
length of query: 525
length of database: 12,639,632
effective HSP length: 102
effective length of query: 423
effective length of database: 6,144,680
effective search space: 2599199640
effective search space used: 2599199640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126007.11


