

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125481.7 + phase: 0 /pseudo
(489 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
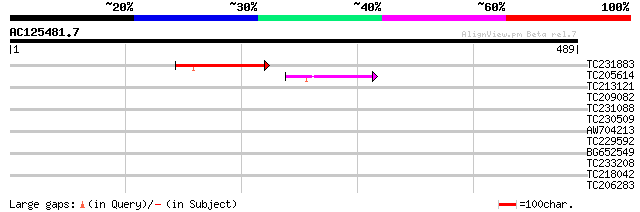
Score E
Sequences producing significant alignments: (bits) Value
TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment... 69 6e-12
TC205614 59 5e-09
TC213121 33 0.22
TC209082 homologue to GB|AAL15278.1|16323087|AY057647 AT3g01490/... 30 1.9
TC231088 weakly similar to UP|O65791 (O65791) Polycomb-like prot... 29 4.2
TC230509 weakly similar to UP|Q946J8 (Q946J8) Like heterochromat... 29 4.2
AW704213 similar to PIR|T06596|T06 lipoxygenase (EC 1.13.11.12) ... 29 4.2
TC229592 similar to UP|Q9M5Q3 (Q9M5Q3) S-ribonuclease binding pr... 29 4.2
BG652549 weakly similar to PIR|S62700|S62 photoassimilate-respon... 29 5.4
TC233208 similar to GB|AAO11557.1|27363276|BT002641 At4g13194/At... 28 7.1
TC218042 UP|Q81I99 (Q81I99) Dihydroorotase , partial (98%) 28 9.3
TC206283 similar to UP|O49553 (O49553) G10-like protein (AT4g211... 28 9.3
>TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment), partial
(17%)
Length = 1263
Score = 68.6 bits (166), Expect = 6e-12
Identities = 36/85 (42%), Positives = 52/85 (60%), Gaps = 4/85 (4%)
Frame = +1
Query: 144 EKEECLAKLMQRHV---PCGGARPFSPCMDRGRCTKYYRKNYKGTTTID-EGYPRYKRRD 199
E + L L+Q H+ PCG + SPCM G+C+++Y K + T +D GY Y+RR+
Sbjct: 1006 EDDPELYTLVQNHMVHGPCGILQSHSPCMKEGKCSRFYPKIFLPNTLLDLNGYLVYRRRN 1185
Query: 200 FGIHVDKQRVLLDNRYVVPYNPHLL 224
G + K V++DNRY+VPYN LL
Sbjct: 1186 DGRTILKNSVIVDNRYLVPYNAKLL 1260
>TC205614
Length = 742
Score = 58.9 bits (141), Expect = 5e-09
Identities = 32/82 (39%), Positives = 49/82 (59%), Gaps = 3/82 (3%)
Frame = +2
Query: 239 SNSIKYLFKYVNKVPD---KALMQLSVDGDNRDKSKPVDEIKQYYDCRYVSPCEAVWRIF 295
SN ++F Y++ V + + ++L D + + S+ Y DC+Y+SPCEA WRIF
Sbjct: 497 SNLFSFIFHYISMVENHN*RTSLKLK-DLASIEASQVSFPHMYYLDCQYISPCEACWRIF 673
Query: 296 AFDIHHK*PHVLKLLFHLHNEQ 317
+F I+ + P V +L FHL NEQ
Sbjct: 674 SFQIYGRIPIVERLYFHLPNEQ 739
>TC213121
Length = 713
Score = 33.5 bits (75), Expect = 0.22
Identities = 17/35 (48%), Positives = 21/35 (59%)
Frame = +3
Query: 282 CRYVSPCEAVWRIFAFDIHHK*PHVLKLLFHLHNE 316
C YVSP E +I AF +H + P V L FHL N+
Sbjct: 30 CIYVSPPETC*KISAFPMHGRAPTVECLYFHLENQ 134
>TC209082 homologue to GB|AAL15278.1|16323087|AY057647 AT3g01490/F4P13_4
{Arabidopsis thaliana;} , partial (29%)
Length = 632
Score = 30.4 bits (67), Expect = 1.9
Identities = 13/23 (56%), Positives = 17/23 (73%)
Frame = -2
Query: 429 YHGKW*VTFGPSIALVVCASLNH 451
YHGK+ + FGP I + V ASL+H
Sbjct: 274 YHGKYLIHFGPFIGIGVPASLHH 206
>TC231088 weakly similar to UP|O65791 (O65791) Polycomb-like protein, partial
(8%)
Length = 1203
Score = 29.3 bits (64), Expect = 4.2
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = +3
Query: 205 DKQRVLLDNRYVVPYNPHLLIRY 227
D V++DNRY+ YNP LLI +
Sbjct: 756 DGTEVMVDNRYLKAYNPLLLINF 824
>TC230509 weakly similar to UP|Q946J8 (Q946J8) Like heterochromatin protein
LHP1, partial (16%)
Length = 782
Score = 29.3 bits (64), Expect = 4.2
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = +2
Query: 205 DKQRVLLDNRYVVPYNPHLLIRY 227
D V++DNRY+ YNP LLI +
Sbjct: 362 DGTEVMVDNRYLKAYNPLLLINF 430
>AW704213 similar to PIR|T06596|T06 lipoxygenase (EC 1.13.11.12) 7 - soybean,
partial (22%)
Length = 589
Score = 29.3 bits (64), Expect = 4.2
Identities = 17/44 (38%), Positives = 21/44 (47%)
Frame = +3
Query: 226 RYGGHVNVECCNKSNSIKYLFKYVNKVPDKALMQLSVDGDNRDK 269
RYG + EC N S+S +VNK+P L DG N K
Sbjct: 381 RYGMPSSPECMNMSHSTTSQMHHVNKIPSSEL-----DGKNLSK 497
>TC229592 similar to UP|Q9M5Q3 (Q9M5Q3) S-ribonuclease binding protein SBP1
(Fragment), partial (91%)
Length = 1267
Score = 29.3 bits (64), Expect = 4.2
Identities = 13/53 (24%), Positives = 25/53 (46%)
Frame = -1
Query: 242 IKYLFKYVNKVPDKALMQLSVDGDNRDKSKPVDEIKQYYDCRYVSPCEAVWRI 294
+KY+ K KV +L N++ + +++ Y+ + PC +WRI
Sbjct: 1177 VKYILKTSYKVSWSLRHKLHERTHNKNMKLEISQVEVYFAVTFRPPCR*IWRI 1019
>BG652549 weakly similar to PIR|S62700|S62 photoassimilate-responsive protein
PAR-1c precursor - common tobacco, partial (69%)
Length = 390
Score = 28.9 bits (63), Expect = 5.4
Identities = 11/34 (32%), Positives = 19/34 (55%)
Frame = +3
Query: 116 ERNLKASDRPDIVCRVFKMKLDKLMDDFEKEECL 149
E+ +K + CR +++ DKL D E E+C+
Sbjct: 162 EKQVKRTGEEAYTCRTSEIEADKLKDHIETEQCI 263
>TC233208 similar to GB|AAO11557.1|27363276|BT002641 At4g13194/At4g13194
{Arabidopsis thaliana;} , partial (17%)
Length = 742
Score = 28.5 bits (62), Expect = 7.1
Identities = 14/28 (50%), Positives = 17/28 (60%)
Frame = +1
Query: 139 LMDDFEKEECLAKLMQRHVPCGGARPFS 166
L DDF K C A L+ +H+P A PFS
Sbjct: 124 LSDDFHKAFCFAWLVIQHLPSPLAWPFS 207
>TC218042 UP|Q81I99 (Q81I99) Dihydroorotase , partial (98%)
Length = 1356
Score = 28.1 bits (61), Expect = 9.3
Identities = 13/25 (52%), Positives = 15/25 (60%)
Frame = +3
Query: 173 RCTKYYRKNYKGTTTIDEGYPRYKR 197
RCT+Y R NY+G T Y RY R
Sbjct: 306 RCTRYRRANYRGQTFYST-YSRYDR 377
>TC206283 similar to UP|O49553 (O49553) G10-like protein (AT4g21110/F7J7_50),
complete
Length = 831
Score = 28.1 bits (61), Expect = 9.3
Identities = 14/44 (31%), Positives = 21/44 (46%), Gaps = 9/44 (20%)
Frame = +3
Query: 288 CEAVWRIFA---------FDIHHK*PHVLKLLFHLHNEQGNIDH 322
CE +W IF FD++H+ + K L+ +QG DH
Sbjct: 303 CETLWPIFKIAHQKSRYIFDLYHQRKEISKELYEFCLDQGYADH 434
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.342 0.150 0.514
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,613,513
Number of Sequences: 63676
Number of extensions: 322471
Number of successful extensions: 2087
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 2074
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2086
length of query: 489
length of database: 12,639,632
effective HSP length: 101
effective length of query: 388
effective length of database: 6,208,356
effective search space: 2408842128
effective search space used: 2408842128
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC125481.7


