

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125480.3 + phase: 0
(476 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
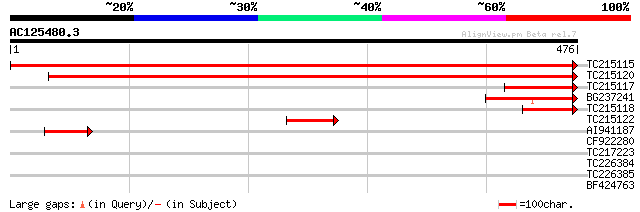
Score E
Sequences producing significant alignments: (bits) Value
TC215115 homologue to PIR|T02313|T02313 endoplasmic reticulum in... 920 0.0
TC215120 homologue to PIR|T02313|T02313 endoplasmic reticulum in... 855 0.0
TC215117 homologue to PIR|T02313|T02313 endoplasmic reticulum in... 124 9e-29
BG237241 99 5e-21
TC215118 homologue to PIR|T02313|T02313 endoplasmic reticulum in... 93 2e-19
TC215122 weakly similar to UP|Q9LZ05 (Q9LZ05) Protein kinase-lik... 88 7e-18
AI941187 45 9e-05
CF922280 31 1.1
TC217223 similar to UP|Q9SMJ7 (Q9SMJ7) MAP kinase protein (Fragm... 28 6.9
TC226384 homologue to UP|Q7Y067 (Q7Y067) Plasma membrane H+-ATPa... 28 6.9
TC226385 homologue to UP|Q7Y067 (Q7Y067) Plasma membrane H+-ATPa... 28 6.9
BF424763 28 9.0
>TC215115 homologue to PIR|T02313|T02313 endoplasmic reticulum insertion
protein F13P17.9 - Arabidopsis thaliana {Arabidopsis
thaliana;} , complete
Length = 1786
Score = 920 bits (2378), Expect = 0.0
Identities = 469/476 (98%), Positives = 473/476 (98%)
Frame = +2
Query: 1 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG
Sbjct: 131 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 310
Query: 61 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 120
ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK
Sbjct: 311 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 490
Query: 121 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS 180
LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILII+QL FA IIVICLDELLQKGYGLGS
Sbjct: 491 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIILQLCFAAIIVICLDELLQKGYGLGS 670
Query: 181 GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ 240
GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ
Sbjct: 671 GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ 850
Query: 241 NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL 300
NLPNVTNLLAT+LIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL
Sbjct: 851 NLPNVTNLLATILIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL 1030
Query: 301 VSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANP 360
VSNLYFISQLLHRKYSGNFIV+LLGKWKESEYGGG S+PVGGIAYYITAPSSLADMAANP
Sbjct: 1031VSNLYFISQLLHRKYSGNFIVDLLGKWKESEYGGGQSVPVGGIAYYITAPSSLADMAANP 1210
Query: 361 FHALFYLVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP 420
FHALFYLVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP
Sbjct: 1211FHALFYLVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP 1390
Query: 421 TAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 476
TAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF
Sbjct: 1391TAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 1558
>TC215120 homologue to PIR|T02313|T02313 endoplasmic reticulum insertion
protein F13P17.9 - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (93%)
Length = 1566
Score = 855 bits (2209), Expect = 0.0
Identities = 436/444 (98%), Positives = 441/444 (99%)
Frame = +3
Query: 33 KVIYTVISLFIFLVCSQLPLYGIHSTTGADPFYWMRVILASNRGTVMELGITPIVTSGLV 92
+VIYTVISLFIFLVCSQLPLYGIHSTTGADPFYWMRVILASNRGTVMELGITPIVTSGLV
Sbjct: 3 EVIYTVISLFIFLVCSQLPLYGIHSTTGADPFYWMRVILASNRGTVMELGITPIVTSGLV 182
Query: 93 MQLLAGSKIIEVDNNVREDRALLNGAQKLLGILIAVGEAVAYVLSGMYGSVGQLGVGNAI 152
MQLLAGSKIIEVDNNVREDRALLNGAQKLLGILIAVGEAVAYVLSGMYGSVGQLGVGNAI
Sbjct: 183 MQLLAGSKIIEVDNNVREDRALLNGAQKLLGILIAVGEAVAYVLSGMYGSVGQLGVGNAI 362
Query: 153 LIIVQLFFAGIIVICLDELLQKGYGLGSGISLFIATNICENIIWKAFSPTTINSGRGAEF 212
LII+QL FAGIIVICLDELLQKGYGLGSGISLFIATNICENIIWKAFSPTTINSGRGAEF
Sbjct: 363 LIIIQLCFAGIIVICLDELLQKGYGLGSGISLFIATNICENIIWKAFSPTTINSGRGAEF 542
Query: 213 EGAVIALFHLLITRTDKVRALREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRS 272
EGAVIALFHLLITRTDKVRALREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRS
Sbjct: 543 EGAVIALFHLLITRTDKVRALREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRS 722
Query: 273 KNARGQQGSYPIKLFYTSNMPIILQSALVSNLYFISQLLHRKYSGNFIVNLLGKWKESEY 332
KNARGQQGSYPIKLFYTSNMPIILQSALVSNLYFISQLLHRKYSGNF V+LLGKWKESEY
Sbjct: 723 KNARGQQGSYPIKLFYTSNMPIILQSALVSNLYFISQLLHRKYSGNFFVDLLGKWKESEY 902
Query: 333 GGGHSIPVGGIAYYITAPSSLADMAANPFHALFYLVFMLSACALFSKTWIEVSGSSARDV 392
GGG S+PVGGIAYYITAPSSLADMAANPFHALFYLVFMLSACALFSKTWIEVSGSSA+DV
Sbjct: 903 GGGQSVPVGGIAYYITAPSSLADMAANPFHALFYLVFMLSACALFSKTWIEVSGSSAKDV 1082
Query: 393 AKQLKEQQMVMPGHRESNLQKELNRYIPTAAAFGGICIGALTVLADFMGAIGSGTGILLA 452
AKQLKEQQMVMPGHRESNLQKELNRYIPTAAAFGGICIGALTVLADFMGAIGSGTGILLA
Sbjct: 1083AKQLKEQQMVMPGHRESNLQKELNRYIPTAAAFGGICIGALTVLADFMGAIGSGTGILLA 1262
Query: 453 VTIIYQYFETFEKERASELGFFGF 476
VTIIYQYFETFEKERASELGFFGF
Sbjct: 1263VTIIYQYFETFEKERASELGFFGF 1334
>TC215117 homologue to PIR|T02313|T02313 endoplasmic reticulum insertion
protein F13P17.9 - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (13%)
Length = 424
Score = 124 bits (311), Expect = 9e-29
Identities = 61/61 (100%), Positives = 61/61 (100%)
Frame = +1
Query: 416 NRYIPTAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFG 475
NRYIPTAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFG
Sbjct: 1 NRYIPTAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFG 180
Query: 476 F 476
F
Sbjct: 181 F 183
>BG237241
Length = 447
Score = 98.6 bits (244), Expect = 5e-21
Identities = 55/80 (68%), Positives = 57/80 (70%), Gaps = 3/80 (3%)
Frame = +2
Query: 400 QMVMPGHRESNLQKELNRYIPTAAAFGGICIGALTVLA---DFMGAIGSGTGILLAVTII 456
QMVMPGHRESNLQKELNRYIPTAAA + + G GILLAVTII
Sbjct: 2 QMVMPGHRESNLQKELNRYIPTAAAXWRHXVSGASDXCWQNFXWGPSVQEXGILLAVTII 181
Query: 457 YQYFETFEKERASELGFFGF 476
YQYFETFEKERASELGFFGF
Sbjct: 182 YQYFETFEKERASELGFFGF 241
>TC215118 homologue to PIR|T02313|T02313 endoplasmic reticulum insertion
protein F13P17.9 - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (10%)
Length = 577
Score = 93.2 bits (230), Expect = 2e-19
Identities = 46/46 (100%), Positives = 46/46 (100%)
Frame = +2
Query: 431 GALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 476
GALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF
Sbjct: 2 GALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 139
>TC215122 weakly similar to UP|Q9LZ05 (Q9LZ05) Protein kinase-like, partial
(22%)
Length = 747
Score = 88.2 bits (217), Expect = 7e-18
Identities = 44/44 (100%), Positives = 44/44 (100%)
Frame = +3
Query: 233 LREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNAR 276
LREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNAR
Sbjct: 3 LREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNAR 134
>AI941187
Length = 121
Score = 44.7 bits (104), Expect = 9e-05
Identities = 22/40 (55%), Positives = 26/40 (65%)
Frame = +2
Query: 30 FREKVIYTVISLFIFLVCSQLPLYGIHSTTGADPFYWMRV 69
F +KV T I ++ VCSQ LYG HSTTGAD FY M +
Sbjct: 2 FTDKVTDTAIGEYMSHVCSQPTLYGTHSTTGADAFYSMHL 121
>CF922280
Length = 436
Score = 31.2 bits (69), Expect = 1.1
Identities = 17/60 (28%), Positives = 28/60 (46%)
Frame = -3
Query: 291 NMPIILQSALVSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAP 350
++P + + +L+F G + N LG WK+ +GGG P GG ++ AP
Sbjct: 203 HLPFFFKGKMGVSLFFFL------LGGGLLFNPLGVWKKIWFGGGGGPPEGGGFFF*GAP 42
>TC217223 similar to UP|Q9SMJ7 (Q9SMJ7) MAP kinase protein (Fragment),
partial (10%)
Length = 723
Score = 28.5 bits (62), Expect = 6.9
Identities = 9/25 (36%), Positives = 17/25 (68%)
Frame = -2
Query: 323 LLGKWKESEYGGGHSIPVGGIAYYI 347
+L +WK+ ++ HS PVG + +Y+
Sbjct: 455 VLHQWKQQDHLSDHSFPVGSMGWYL 381
>TC226384 homologue to UP|Q7Y067 (Q7Y067) Plasma membrane H+-ATPase, partial
(15%)
Length = 754
Score = 28.5 bits (62), Expect = 6.9
Identities = 21/52 (40%), Positives = 28/52 (53%)
Frame = +2
Query: 31 REKVIYTVISLFIFLVCSQLPLYGIHSTTGADPFYWMRVILASNRGTVMELG 82
R+ VIY +SLFIFL C +G++ A W +V+LAS M LG
Sbjct: 530 RKPVIYLSLSLFIFLSC-LFRCFGLNFCFDA----WGKVVLASILDFHMLLG 670
>TC226385 homologue to UP|Q7Y067 (Q7Y067) Plasma membrane H+-ATPase, partial
(25%)
Length = 1109
Score = 28.5 bits (62), Expect = 6.9
Identities = 21/52 (40%), Positives = 28/52 (53%)
Frame = +1
Query: 31 REKVIYTVISLFIFLVCSQLPLYGIHSTTGADPFYWMRVILASNRGTVMELG 82
R+ VIY +SLFIFL C +G++ A W +V+LAS M LG
Sbjct: 832 RKPVIYLSLSLFIFLSC-LFRCFGLNFCFDA----WGKVVLASILDFHMLLG 972
>BF424763
Length = 413
Score = 28.1 bits (61), Expect = 9.0
Identities = 14/56 (25%), Positives = 28/56 (50%)
Frame = -3
Query: 292 MPIILQSALVSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYI 347
M IIL+ ++ I +H +S + ++ +L + +GG H I G + Y++
Sbjct: 285 MKIILKDLVMVERRVIPVAIHVLHSLSVLIQILNQRDPMPWGGRHGILSGSLEYFL 118
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.143 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,883,640
Number of Sequences: 63676
Number of extensions: 254161
Number of successful extensions: 1569
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1553
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1568
length of query: 476
length of database: 12,639,632
effective HSP length: 101
effective length of query: 375
effective length of database: 6,208,356
effective search space: 2328133500
effective search space used: 2328133500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC125480.3


