

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125480.1 - phase: 0 /pseudo
(250 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
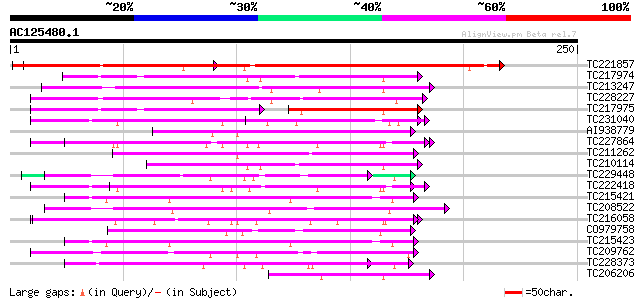
Score E
Sequences producing significant alignments: (bits) Value
TC221857 similar to UP|Q8LFJ3 (Q8LFJ3) Contains similarity to pe... 239 1e-63
TC217974 similar to UP|Q9LY28 (Q9LY28) Peroxisomal Ca-dependent ... 75 4e-14
TC213247 weakly similar to UP|SFC1_YEAST (P33303) Succinate/fuma... 74 6e-14
TC228227 similar to GB|AAL69531.1|18491125|AY074833 At2g47490/T3... 66 1e-11
TC217975 similar to UP|Q9LY28 (Q9LY28) Peroxisomal Ca-dependent ... 50 1e-10
TC231040 similar to UP|Q9MBE7 (Q9MBE7) SfUCPa, partial (67%) 63 1e-10
AI938779 63 1e-10
TC227864 uncoupling protein 1a [Glycine max] 60 9e-10
TC211262 similar to UP|Q9FM86 (Q9FM86) ADP/ATP translocase-like ... 59 2e-09
TC210114 similar to UP|Q9FI43 (Q9FI43) Calcium-binding transport... 59 2e-09
TC229448 similar to UP|Q6YVE7 (Q6YVE7) Mitochondrial aspartate-g... 58 4e-09
TC222418 UP|Q8W1A3 (Q8W1A3) Uncoupling protein 1b (Fragment), pa... 58 4e-09
TC215421 weakly similar to UP|DIC_MOUSE (Q9QZD8) Mitochondrial d... 57 8e-09
TC208522 similar to UP|Q6YVE7 (Q6YVE7) Mitochondrial aspartate-g... 56 1e-08
TC216058 similar to UP|MCAT_ARATH (Q93XM7) Mitochondrial carniti... 56 2e-08
CO979758 55 2e-08
TC215423 weakly similar to GB|AAH19631.1|18043006|BC019631 solut... 54 5e-08
TC209762 52 2e-07
TC228373 similar to GB|AAM19967.1|20466091|AY098957 AT5g01340/T1... 52 3e-07
TC206206 similar to PIR|T47703|T47703 Ca-dependent solute carrie... 50 1e-06
>TC221857 similar to UP|Q8LFJ3 (Q8LFJ3) Contains similarity to peroxisomal
membrane carrier protein, partial (93%)
Length = 945
Score = 239 bits (609), Expect = 1e-63
Identities = 127/231 (54%), Positives = 160/231 (68%), Gaps = 14/231 (6%)
Frame = +1
Query: 2 NELQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKE 61
++ QWE A GFA + +PLDVV+TR QV+DGR+ S +P Y NT HA+FTIAR E
Sbjct: 10 DQWQWENATAGAAAGFATVAVMHPLDVVRTRFQVNDGRV-SNFPSYKNTAHAVFTIARSE 186
Query: 62 GLKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK------------SGALMTV 109
GL+GLYAGF G+LG+TISWSL F+Y K R+AR++ K +GA+ V
Sbjct: 187 GLRGLYAGFLPGVLGSTISWSLYFFFYDRAKQRYARNREGKLSPGLHLASAAEAGAI--V 360
Query: 110 CLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP ++VKTRLQLQTPLH RPYSG+YDAFRTI REEGFSA YRGIVPG FL+S A
Sbjct: 361 SFFTNPVWLVKTRLQLQTPLHQTRPYSGVYDAFRTIMREEGFSALYRGIVPGLFLVSHGA 540
Query: 170 IQFIVYEQLRKTVVNLKTKGSKIQHQKPDQIL--VCYWTLDLFTSEISLVL 218
IQF YE+LRK +V+ K+KGS + +Q PD++L V Y L TS+++ VL
Sbjct: 541 IQFTAYEELRKVIVDFKSKGSTVDNQNPDKLLNSVDYAVLGA-TSKLAAVL 690
Score = 45.8 bits (107), Expect = 2e-05
Identities = 28/87 (32%), Positives = 40/87 (45%), Gaps = 1/87 (1%)
Frame = +1
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGL 66
+Y V T+ A + YP V++ RLQ PRY +T H + AR E ++G
Sbjct: 646 DYAVLGATSKLAAVLLTYPFQVIRARLQQRPSG--DGVPRYMDTLHVVKETARFESVRGF 819
Query: 67 YAGFPAGLL-GATISWSLLVFYYGIVK 92
Y G A LL A S + Y ++K
Sbjct: 820 YKGITANLLKNAPASSITFIVYENVLK 900
Score = 39.3 bits (90), Expect = 0.002
Identities = 47/180 (26%), Positives = 66/180 (36%), Gaps = 21/180 (11%)
Frame = +1
Query: 22 FQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLL----GA 77
F P+ +V+TRLQ+ Q Y+ A TI R+EG LY G GL GA
Sbjct: 367 FTNPVWLVKTRLQLQTP--LHQTRPYSGVYDAFRTIMREEGFSALYRGIVPGLFLVSHGA 540
Query: 78 TISWSLLVFYYGIVKDRHARSKVEKSG----------------ALMTVCLCANPAFVVKT 121
+ IV + S V+ + + L P V++
Sbjct: 541 IQFTAYEELRKVIVDFKSKGSTVDNQNPDKLLNSVDYAVLGATSKLAAVLLTYPFQVIRA 720
Query: 122 RLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQFIVYEQLRK 180
RLQ + Y + R E FY+GI + A +I FIVYE + K
Sbjct: 721 RLQQRPSGDGVPRYMDTLHVVKETARFESVRGFYKGITANLLKNAPASSITFIVYENVLK 900
>TC217974 similar to UP|Q9LY28 (Q9LY28) Peroxisomal Ca-dependent solute
carrier-like protein, partial (32%)
Length = 695
Score = 74.7 bits (182), Expect = 4e-14
Identities = 54/172 (31%), Positives = 87/172 (50%), Gaps = 13/172 (7%)
Frame = +2
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
YP+D+++TRLQ + P+ T I+ +EG + Y G LLG ++
Sbjct: 11 YPMDLIKTRLQTCPSE-GGKVPKLGTLTMNIWF---QEGPRAFYRGLVPSLLGMIPYAAI 178
Query: 84 LVFYYGIVKDRHARSKVEKS--GALMTV----------CLCANPAFVVKTRLQLQTPLHH 131
+ Y +KD R ++ S G L+ + C P V++TRLQ Q P +
Sbjct: 179 DLTAYDTMKDISKRYILQDSEPGPLVQLGCGTISGAVGATCVYPLQVIRTRLQAQ-PSNT 355
Query: 132 ARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRKTV 182
+ Y G++DAFR + EGF FY+G+ P ++ A+I ++VYE L+KT+
Sbjct: 356 SDAYKGMFDAFRRTFQLEGFIGFYKGLFPNLLKVVPAASITYVVYESLKKTL 511
Score = 29.6 bits (65), Expect = 1.4
Identities = 22/78 (28%), Positives = 30/78 (38%)
Frame = +2
Query: 15 TGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGL 74
+G T YPL V++TRLQ Y + F + EG G Y G L
Sbjct: 278 SGAVGATCVYPLQVIRTRLQAQPSNTSDAYKGMFDAFRRTFQL---EGFIGFYKGLFPNL 448
Query: 75 LGATISWSLLVFYYGIVK 92
L + S+ Y +K
Sbjct: 449 LKVVPAASITYVVYESLK 502
>TC213247 weakly similar to UP|SFC1_YEAST (P33303) Succinate/fumarate
mitochondrial transporter (Regulator of acetyl-CoA
synthetase activity), partial (11%)
Length = 829
Score = 73.9 bits (180), Expect = 6e-14
Identities = 55/186 (29%), Positives = 90/186 (47%), Gaps = 13/186 (6%)
Frame = +3
Query: 15 TGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGL 74
+G + YPLD+V+TRL ++ Y +HA TI R EG GLY G A L
Sbjct: 99 SGITSASATYPLDLVRTRLAAQRSTMY-----YRGISHAFSTICRDEGFLGLYKGLGATL 263
Query: 75 LGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCAN-----------PAFVVKTRL 123
LG S ++ Y ++ + + S A++ + C + P +V+ R+
Sbjct: 264 LGVGPSIAISFAVYEWLRSVWQSQRPDDSKAVVGLA-CGSLSGIASSTATFPLDLVRRRM 440
Query: 124 QLQTPLHHARPY-SGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRKT 181
QL+ AR Y +GL+ AF I + EG YRGI+P ++ ++ I F+ YE L+
Sbjct: 441 QLEGVGGRARVYNTGLFGAFGRIIQTEGVRGLYRGILPEYYKVVPGVGIVFMTYETLKML 620
Query: 182 VVNLKT 187
+ ++ +
Sbjct: 621 LSSISS 638
>TC228227 similar to GB|AAL69531.1|18491125|AY074833 At2g47490/T30B22.21
{Arabidopsis thaliana;} , partial (61%)
Length = 1098
Score = 66.2 bits (160), Expect = 1e-11
Identities = 60/193 (31%), Positives = 89/193 (46%), Gaps = 18/193 (9%)
Frame = +2
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A+ G A F PL VV+TRLQ R Y T A+ IA +EG++GLY+G
Sbjct: 92 IAASGAGAATTMFTNPLWVVKTRLQTQGIR--PGVVPYRGTLSALRRIAHEEGIRGLYSG 265
Query: 70 FPAGLLGATI------SWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCAN--------- 114
L G + ++ + FY D + +EK GA V + ++
Sbjct: 266 LVPALAGISHVAIQFPTYETIKFYLANQDD----TAMEKLGA-RDVAIASSVSKIFASTL 430
Query: 115 --PAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA-IQ 171
P VV++RLQ Q H + YSG+ D R + +EG S FYRG + AA I
Sbjct: 431 TYPHEVVRSRLQEQGH-HSEKRYSGVIDCIRKVFHQEGVSGFYRGCATNLLRTTPAAVIT 607
Query: 172 FIVYEQLRKTVVN 184
F +E + + +V+
Sbjct: 608 FTSFEMIHRFLVS 646
>TC217975 similar to UP|Q9LY28 (Q9LY28) Peroxisomal Ca-dependent solute
carrier-like protein, partial (44%)
Length = 1187
Score = 49.7 bits (117), Expect(2) = 1e-10
Identities = 25/61 (40%), Positives = 39/61 (62%), Gaps = 2/61 (3%)
Frame = +3
Query: 124 QLQT-PLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRKT 181
QLQ P + + Y G++DAFR + EGF FY+G+ P ++ A+I ++VYE L+KT
Sbjct: 675 QLQAQPSNTSDAYKGMFDAFRRTFQLEGFIGFYKGLFPNLLKVVPAASITYVVYESLKKT 854
Query: 182 V 182
+
Sbjct: 855 L 857
Score = 33.1 bits (74), Expect(2) = 1e-10
Identities = 29/103 (28%), Positives = 44/103 (42%)
Frame = +1
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
VA T G YP+D+++TRLQ + P+ T I+ +EG + Y G
Sbjct: 319 VAGGTAGAIAQAAIYPMDLIKTRLQTCPSE-GGKVPKLGTLTMNIWV---QEGPRAFYRG 486
Query: 70 FPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLC 112
LLG ++ + Y +KD R ++ SG V C
Sbjct: 487 LVPSLLGMIPYAAIDLTAYDTMKDISKRYILQDSGYSNKVQCC 615
Score = 32.7 bits (73), Expect = 0.16
Identities = 37/142 (26%), Positives = 60/142 (42%), Gaps = 15/142 (10%)
Frame = +1
Query: 53 AIFTIARKEGLKGLYAGFPAGLLGATISWSLLVFYYG------IVKDRHA-RSKVEKSGA 105
A+ I +++GL G + G GL +S + +Y ++ + H +S + +G
Sbjct: 142 AVTKIWKQDGLLGFFRG--NGLNVVKVSPESAIKFYAFEMLKKVIGEAHGNKSDIGTAGR 315
Query: 106 LMT-------VCLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGI 158
L+ P ++KTRLQ P G I +EG AFYRG+
Sbjct: 316 LVAGGTAGAIAQAAIYPMDLIKTRLQTCPSEGGKVPKLGTLTM--NIWVQEGPRAFYRGL 489
Query: 159 VPGFF-LISQAAIQFIVYEQLR 179
VP +I AAI Y+ ++
Sbjct: 490 VPSLLGMIPYAAIDLTAYDTMK 555
>TC231040 similar to UP|Q9MBE7 (Q9MBE7) SfUCPa, partial (67%)
Length = 980
Score = 62.8 bits (151), Expect = 1e-10
Identities = 55/185 (29%), Positives = 81/185 (43%), Gaps = 12/185 (6%)
Frame = +3
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKGLYA 68
+A+L TG IT P D+V+ RLQ +G+L S PR Y+ A TI R+EG+ L+
Sbjct: 84 LAALLTGALAITIANPTDLVKVRLQA-EGQLPSGVPRRYSGAIDAYLTILRQEGIGALWT 260
Query: 69 GFPAGLLGATISWSLLVFYYGIVK----------DRHARSKVEKSGALMTVCLCANPAFV 118
G + I + + Y VK D + GA + +P V
Sbjct: 261 GLGPNIARNAIINAAELASYDKVKRTILKIPGFMDNVYTHLLAGLGAGLFAVFIGSPVDV 440
Query: 119 VKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS-QAAIQFIVYEQ 177
VK+R+ + Y +D F EGF AFY+G +P F + I F+ EQ
Sbjct: 441 VKSRMMGDST------YKSTFDCFLKTLLNEGFLAFYKGFLPNFGRVGIWNVILFLTLEQ 602
Query: 178 LRKTV 182
++ V
Sbjct: 603 AKRAV 617
Score = 46.2 bits (108), Expect = 1e-05
Identities = 33/90 (36%), Positives = 50/90 (54%), Gaps = 9/90 (10%)
Frame = +3
Query: 105 ALMTVCLC---ANPAFVVKTRLQL--QTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIV 159
AL+T L ANP +VK RLQ Q P R YSG DA+ TI R+EG A + G+
Sbjct: 90 ALLTGALAITIANPTDLVKVRLQAEGQLPSGVPRRYSGAIDAYLTILRQEGIGALWTGLG 269
Query: 160 PGFFLISQAAI----QFIVYEQLRKTVVNL 185
P I++ AI + Y+++++T++ +
Sbjct: 270 PN---IARNAIINAAELASYDKVKRTILKI 350
>AI938779
Length = 417
Score = 62.8 bits (151), Expect = 1e-10
Identities = 38/127 (29%), Positives = 58/127 (44%), Gaps = 11/127 (8%)
Frame = +3
Query: 64 KGLYAGFPAGLLGATISWSLLVFYY----GIVKDRHARSK-------VEKSGALMTVCLC 112
+G+Y G ++ +W++ Y G+++ R + + +GA +
Sbjct: 3 RGMYRGLSPTIVALLPNWAVYFTSYEQLKGLLRSRDGCDELTTIGNIIAAAGAGAATAIS 182
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQF 172
NP +VVKTRLQ Q PY + A I EEG Y GIVP +S AIQF
Sbjct: 183 TNPLWVVKTRLQTQGMRPDVVPYKSVLSALTRITHEEGIRGLYSGIVPSLAGVSHVAIQF 362
Query: 173 IVYEQLR 179
YE+++
Sbjct: 363 PAYEKIK 383
Score = 35.8 bits (81), Expect = 0.019
Identities = 23/69 (33%), Positives = 33/69 (47%)
Frame = +3
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A+ G A PL VV+TRLQ R Y + A+ I +EG++GLY+G
Sbjct: 144 IAAAGAGAATAISTNPLWVVKTRLQTQGMR--PDVVPYKSVLSALTRITHEEGIRGLYSG 317
Query: 70 FPAGLLGAT 78
L G +
Sbjct: 318 IVPSLAGVS 344
>TC227864 uncoupling protein 1a [Glycine max]
Length = 1434
Score = 60.1 bits (144), Expect = 9e-10
Identities = 55/182 (30%), Positives = 82/182 (44%), Gaps = 21/182 (11%)
Frame = +1
Query: 25 PLDVVQTRLQVHDGRLFSQY---PRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISW 81
PLD + RLQ+ + P+Y + TIAR+EGL L+ G GL +
Sbjct: 310 PLDTAKVRLQLQKQAVAGDVVSLPKYKGMLGTVGTIAREEGLSALWKGIVPGLHRQCLYG 489
Query: 82 SLLV--------FYYGIVKDRHARSKVEKS--GALMT---VCLCANPAFVVKTRLQLQTP 128
L + FY G KD + K A T ANP +VK RLQ +
Sbjct: 490 GLRIGLYEPVKTFYVG--KDHVGDVPLSKKILAAFTTGAFAIAVANPTDLVKVRLQAEGK 663
Query: 129 LHHARP--YSGLYDAFRTIKREEGFSAFYRGIVPGFF---LISQAAIQFIVYEQLRKTVV 183
L P YSG +A+ TI R+EG A + G+ P +I+ A + Y+Q+++T++
Sbjct: 664 LPPGVPRRYSGSLNAYSTIVRQEGVGALWTGLGPNIARNGIIN--AAELASYDQVKQTIL 837
Query: 184 NL 185
+
Sbjct: 838 KI 843
Score = 57.4 bits (137), Expect = 6e-09
Identities = 55/190 (28%), Positives = 84/190 (43%), Gaps = 12/190 (6%)
Frame = +1
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKGLYA 68
+A+ TTG I P D+V+ RLQ +G+L PR Y+ + +A TI R+EG+ L+
Sbjct: 577 LAAFTTGAFAIAVANPTDLVKVRLQA-EGKLPPGVPRRYSGSLNAYSTIVRQEGVGALWT 753
Query: 69 GFPAGLLGATISWSLLVFYYGIVK----------DRHARSKVEKSGALMTVCLCANPAFV 118
G + I + + Y VK D + GA +P V
Sbjct: 754 GLGPNIARNGIINAAELASYDQVKQTILKIPGFTDNVVTHLLAGLGAGFFAVCIGSPVDV 933
Query: 119 VKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGF-FLISQAAIQFIVYEQ 177
VK+R+ + Y D F + +G AFY+G +P F L S I F+ EQ
Sbjct: 934 VKSRMMGDSS------YKNTLDCFIKTLKNDGPLAFYKGFLPNFGRLGSWNVIMFLTLEQ 1095
Query: 178 LRKTVVNLKT 187
+K V +L++
Sbjct: 1096TKKFVKSLES 1125
Score = 39.7 bits (91), Expect = 0.001
Identities = 23/56 (41%), Positives = 28/56 (49%), Gaps = 5/56 (8%)
Frame = +1
Query: 111 LCANPAFVVKTRLQLQTP-----LHHARPYSGLYDAFRTIKREEGFSAFYRGIVPG 161
+C P K RLQLQ + Y G+ TI REEG SA ++GIVPG
Sbjct: 298 VCTIPLDTAKVRLQLQKQAVAGDVVSLPKYKGMLGTVGTIAREEGLSALWKGIVPG 465
>TC211262 similar to UP|Q9FM86 (Q9FM86) ADP/ATP translocase-like protein,
partial (47%)
Length = 775
Score = 59.3 bits (142), Expect = 2e-09
Identities = 37/145 (25%), Positives = 67/145 (45%), Gaps = 10/145 (6%)
Frame = +3
Query: 46 RYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSK------ 99
++ H + TI K+G++G+Y G PA L G + L + +K+ +
Sbjct: 54 QFRGIYHFLATIFHKDGVRGIYKGLPASLHGMVVHRGLYFGGFDTMKEIMSEESKPELAL 233
Query: 100 ----VEKSGALMTVCLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFY 155
V + L + P V+ R+ +Q+ + Y+ D +R I R EG ++FY
Sbjct: 234 WKRWVVAQAVTTSAGLISYPLDTVRRRMMMQSGIEQP-VYNSTLDCWRKIYRTEGLASFY 410
Query: 156 RGIVPGFFLISQAAIQFIVYEQLRK 180
RG V F + AA ++Y++++K
Sbjct: 411 RGAVSNVFRSTGAAAILVLYDEVKK 485
Score = 39.3 bits (90), Expect = 0.002
Identities = 26/81 (32%), Positives = 41/81 (50%)
Frame = +3
Query: 12 SLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFP 71
++TT I+ YPLD V+ R+ + G + P YN+T I R EGL Y G
Sbjct: 258 AVTTSAGLIS--YPLDTVRRRMMMQSG---IEQPVYNSTLDCWRKIYRTEGLASFYRGAV 422
Query: 72 AGLLGATISWSLLVFYYGIVK 92
+ + +T + ++LV Y + K
Sbjct: 423 SNVFRSTGAAAILVLYDEVKK 485
>TC210114 similar to UP|Q9FI43 (Q9FI43) Calcium-binding transporter-like
protein, partial (28%)
Length = 753
Score = 59.3 bits (142), Expect = 2e-09
Identities = 40/135 (29%), Positives = 65/135 (47%), Gaps = 13/135 (9%)
Frame = +2
Query: 61 EGLKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKS--GALMTV--------- 109
EG + Y G LLG + + Y +KD R + S G L+ +
Sbjct: 35 EGPRAFYRGLVPSLLGMIPYAGIDLTAYDTLKDLSKRYILYDSDPGPLVQLGCGTVSGAL 214
Query: 110 -CLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQ 167
C P V++TRLQ Q P + Y G+ D F ++EGF FY+G++P ++
Sbjct: 215 GATCVYPLQVIRTRLQAQ-PANSTSAYKGMSDVFWKTLKDEGFRGFYKGLIPNLLKVVPA 391
Query: 168 AAIQFIVYEQLRKTV 182
A+I ++VYE ++K++
Sbjct: 392 ASITYMVYESMKKSL 436
Score = 33.5 bits (75), Expect = 0.094
Identities = 21/72 (29%), Positives = 30/72 (41%)
Frame = +2
Query: 21 TFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATIS 80
T YPL V++TRLQ S Y + + + EG +G Y G LL +
Sbjct: 221 TCVYPLQVIRTRLQAQPANSTS---AYKGMSDVFWKTLKDEGFRGFYKGLIPNLLKVVPA 391
Query: 81 WSLLVFYYGIVK 92
S+ Y +K
Sbjct: 392 ASITYMVYESMK 427
>TC229448 similar to UP|Q6YVE7 (Q6YVE7) Mitochondrial aspartate-glutamate
carrier protein-like, partial (96%)
Length = 1000
Score = 58.2 bits (139), Expect = 4e-09
Identities = 55/187 (29%), Positives = 75/187 (39%), Gaps = 13/187 (6%)
Frame = +1
Query: 6 WEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
+E +A T G T YP+D ++TRLQ G K LKG
Sbjct: 142 FEGVIAGGTAGVVVETALYPIDTIKTRLQAARG-------------------GEKLILKG 264
Query: 66 LYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGAL----------MTVCLCANP 115
LY+G L+G + +L V Y +K + R E A + L P
Sbjct: 265 LYSGLAGNLVGVLPASALFVGVYEPIKQKLLRIFPEHLSAFTHLTAGAIGGIAASLIRVP 444
Query: 116 AFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA---AIQF 172
V+K R+Q ++ A R I +EGF FY G G FL+ AIQF
Sbjct: 445 TEVIKQRMQ-------TGQFASASGAVRFIASKEGFKGFYAGY--GSFLLRDLPFDAIQF 597
Query: 173 IVYEQLR 179
+YEQ+R
Sbjct: 598 CIYEQIR 618
Score = 49.7 bits (117), Expect = 1e-06
Identities = 46/158 (29%), Positives = 70/158 (44%), Gaps = 13/158 (8%)
Frame = +1
Query: 16 GFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLL 75
G A + P +V++ R+Q ++ + + A+ IA KEG KG YAG+ + LL
Sbjct: 415 GIAASLIRVPTEVIKQRMQTG---------QFASASGAVRFIASKEGFKGFYAGYGSFLL 567
Query: 76 GATISWSLLVFY--------YGIVKDRHARSKVEK-----SGALMTVCLCANPAFVVKTR 122
+ + + F Y + R+ +GAL P V+KTR
Sbjct: 568 -RDLPFDAIQFCIYEQIRIGYMLAAQRNLNDPENAIIGAFAGALTGAI--TTPLDVIKTR 738
Query: 123 LQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVP 160
L +Q A Y G+ D +TI +EEG AF +GI P
Sbjct: 739 LMVQGS---ANQYKGIVDCVQTIIKEEGPRAFLKGIGP 843
Score = 27.7 bits (60), Expect = 5.1
Identities = 18/68 (26%), Positives = 32/68 (46%)
Frame = +1
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLL 84
PLDV++TRL V +Y + TI ++EG + G +L I S+
Sbjct: 715 PLDVIKTRLMVQ-----GSANQYKGIVDCVQTIIKEEGPRAFLKGIGPRVLWIGIGGSI- 876
Query: 85 VFYYGIVK 92
++G+++
Sbjct: 877 --FFGVLE 894
>TC222418 UP|Q8W1A3 (Q8W1A3) Uncoupling protein 1b (Fragment), partial (98%)
Length = 884
Score = 58.2 bits (139), Expect = 4e-09
Identities = 49/158 (31%), Positives = 76/158 (48%), Gaps = 17/158 (10%)
Frame = +3
Query: 45 PRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHA-------- 96
PRY + TIAR+EG L+ G GL ++ L + Y VK+ +
Sbjct: 33 PRYRGLLGTVGTIAREEGFSALWKGIVPGLHRQCLNGGLRIALYEPVKNFYVGADHVGDV 212
Query: 97 -RSKVEKSG---ALMTVCLCANPAFVVKTRLQLQTPLHHARP--YSGLYDAFRTIKREEG 150
SK +G M + + ANP +VK RLQ + L P YSG +A+ TI R+EG
Sbjct: 213 PLSKKILAGFTTGAMAIAV-ANPTDLVKVRLQAEGKLPPGVPRRYSGSLNAYSTIVRQEG 389
Query: 151 FSAFYRGIVPGFF---LISQAAIQFIVYEQLRKTVVNL 185
A + GI P +I+ A + Y+Q+++T++ +
Sbjct: 390 VGALWTGIGPNIARNGIIN--AAELASYDQVKQTILKI 497
Score = 53.9 bits (128), Expect = 7e-08
Identities = 52/182 (28%), Positives = 78/182 (42%), Gaps = 12/182 (6%)
Frame = +3
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKGLYA 68
+A TTG I P D+V+ RLQ +G+L PR Y+ + +A TI R+EG+ L+
Sbjct: 231 LAGFTTGAMAIAVANPTDLVKVRLQA-EGKLPPGVPRRYSGSLNAYSTIVRQEGVGALWT 407
Query: 69 GFPAGLLGATISWSLLVFYYGIVK----------DRHARSKVEKSGALMTVCLCANPAFV 118
G + I + + Y VK D + GA +P V
Sbjct: 408 GIGPNIARNGIINAAELASYDQVKQTILKIPGFTDNVVTHLLAGLGAGFFAVCVGSPVDV 587
Query: 119 VKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGF-FLISQAAIQFIVYEQ 177
VK+R+ + Y D F + +G AFY+G +P F L S I F+ EQ
Sbjct: 588 VKSRMMGDSS------YKSTLDCFVKTLKNDGPFAFYKGFIPNFGRLGSWNVIMFLTLEQ 749
Query: 178 LR 179
++
Sbjct: 750 VQ 755
>TC215421 weakly similar to UP|DIC_MOUSE (Q9QZD8) Mitochondrial dicarboxylate
carrier, partial (13%)
Length = 1292
Score = 57.0 bits (136), Expect = 8e-09
Identities = 48/172 (27%), Positives = 76/172 (43%), Gaps = 16/172 (9%)
Frame = +1
Query: 25 PLDVVQTRLQVHDGRLF-SQYPRYNNTTHAIFTIARKEGLKGLYAG----------FPAG 73
P DV R+Q DGRL +Q Y + AI +A++EG+ L+ G A
Sbjct: 622 PADVAMVRMQA-DGRLPPAQRRNYKSVVDAITRMAKQEGVTSLWRGSSLTVNRAMLVTAS 798
Query: 74 LLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCANPAFVVKTR-LQLQTPLHHA 132
L + + ++ G+++D A + +NP V+KTR + ++ A
Sbjct: 799 QLASYDQFKEMILENGVMRDGLGTHVTASFAAGFVAAVASNPIDVIKTRVMNMRVEPGEA 978
Query: 133 RPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQ----AAIQFIVYEQLRK 180
PY+G D R EG A Y+G +P IS+ + F+ EQ+RK
Sbjct: 979 PPYAGALDCALKTVRAEGPMALYKGFIP---TISRQGPFTVVLFVTLEQVRK 1125
>TC208522 similar to UP|Q6YVE7 (Q6YVE7) Mitochondrial aspartate-glutamate
carrier protein-like, partial (72%)
Length = 1068
Score = 56.2 bits (134), Expect = 1e-08
Identities = 53/190 (27%), Positives = 82/190 (42%), Gaps = 11/190 (5%)
Frame = +2
Query: 16 GFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGF----- 70
G A + P +VV+ R+Q+ ++ + A+ I EG KGL+AG+
Sbjct: 113 GIASSVVRVPTEVVKQRMQIG---------QFKSAPDAVRLIVANEGFKGLFAGYGSFLL 265
Query: 71 ---PAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCA--NPAFVVKTRLQL 125
P + I L + Y K + GA+ A P VVKTRL +
Sbjct: 266 RDLPFDAIELCIYEQLRIGYKLAAKRDPNDPENAMLGAVAGAVTGAVTTPLDVVKTRLMV 445
Query: 126 QTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS-QAAIQFIVYEQLRKTVVN 184
Q +H Y G+ D RTI +EEG A ++GI P I +I F V E+ +K +
Sbjct: 446 QGSQNH---YKGISDCVRTIVKEEGSHALFKGIGPRVLWIGIGGSIFFCVLEKTKKILAQ 616
Query: 185 LKTKGSKIQH 194
+ ++ Q+
Sbjct: 617 KRHSKAETQN 646
>TC216058 similar to UP|MCAT_ARATH (Q93XM7) Mitochondrial
carnitine/acylcarnitine carrier-like protein (A BOUT DE
SOUFFLE) (Carnitine/acylcarnitine translocase-like
protein) (CAC-like protein), partial (97%)
Length = 1441
Score = 55.8 bits (133), Expect = 2e-08
Identities = 54/189 (28%), Positives = 81/189 (42%), Gaps = 19/189 (10%)
Frame = +2
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGF 70
A G A + +P D ++ +LQ L Q P+Y+ A+ EG +GLY G
Sbjct: 407 AGTVGGAAQLICGHPFDTIKVKLQSQPAPLPGQLPKYSGAFDAVKQTIAAEGARGLYKGM 586
Query: 71 PAGLLGATISWSLLVFYYGIVKDRHARSK-----------VEKSGALMTVCLCANPAFVV 119
A L +++ ++F + RS V +GA + V + A P ++
Sbjct: 587 GAP-LATVAAFNAVLFTVRGQMETLVRSNPGAPLTVDQQVVCGAGAGVAVSILACPTELI 763
Query: 120 KTRLQLQTPLHH------ARPYSGLYDAFR-TIKREEGFSAFYRGIVPGFFL-ISQAAIQ 171
K RLQ Q+ L A Y G D R +K E G ++G+VP I AI
Sbjct: 764 KCRLQAQSALAGSETATVAVKYGGPMDVARHVLKSEGGMRGLFKGLVPTMGREIPGNAIM 943
Query: 172 FIVYEQLRK 180
F VYE L++
Sbjct: 944 FGVYEALKQ 970
Score = 45.1 bits (105), Expect = 3e-05
Identities = 46/191 (24%), Positives = 77/191 (40%), Gaps = 18/191 (9%)
Frame = +2
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYP----RYNNTTHAIFTIARKEG-LK 64
V G A P ++++ RLQ S+ +Y + + EG ++
Sbjct: 704 VCGAGAGVAVSILACPTELIKCRLQAQSALAGSETATVAVKYGGPMDVARHVLKSEGGMR 883
Query: 65 GLYAGFPAGLLGATISWSLLVF-YYGIVKDRHA---------RSKVEKSGALM--TVCLC 112
GL+ G +G I + ++F Y +K + A R + +G L +
Sbjct: 884 GLFKGL-VPTMGREIPGNAIMFGVYEALKQKFAGGTDTSGLSRGSLIVAGGLAGASFWFL 1060
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQ 171
P V+K+ +Q+ H +SG +DAFR I+ EGF Y+G P + A
Sbjct: 1061VYPTDVIKSVIQVDD--HRNPKFSGSFDAFRKIRATEGFKGLYKGFGPAMARSVPANAAC 1234
Query: 172 FIVYEQLRKTV 182
F+ YE R +
Sbjct: 1235FLAYEMTRSAL 1267
>CO979758
Length = 751
Score = 55.5 bits (132), Expect = 2e-08
Identities = 50/157 (31%), Positives = 67/157 (41%), Gaps = 21/157 (13%)
Frame = -1
Query: 44 YPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLLVFYYGIVKDR--------- 94
Y Y HA +I + +GLKGLYAG+ + L L+V +Y +KD
Sbjct: 709 YGYYTGMLHAGCSIWKAQGLKGLYAGYLSTLARDVPFAGLMVVFYEALKDAKDYVEQRWI 530
Query: 95 -----HARSKVE------KSGALMTVCLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFR 143
H + VE +G L P VVKTRLQ+Q Y+G DA
Sbjct: 529 SSPNWHVNNSVEGLVLGGLAGGLS--AYLTTPLDVVKTRLQVQ---GSTLRYNGWLDAIH 365
Query: 144 TIKREEGFSAFYRGIVPGF-FLISQAAIQFIVYEQLR 179
I EG +RG VP + I +A+ F+ E LR
Sbjct: 364 NIWATEGMKGMFRGSVPRITWYIPASALTFMAVEFLR 254
Score = 38.9 bits (89), Expect = 0.002
Identities = 22/45 (48%), Positives = 26/45 (56%)
Frame = -1
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
PLDVV+TRLQV L RYN AI I EG+KG++ G
Sbjct: 442 PLDVVKTRLQVQGSTL-----RYNGWLDAIHNIWATEGMKGMFRG 323
>TC215423 weakly similar to GB|AAH19631.1|18043006|BC019631 solute carrier
family 25 (mitochondrial carrier oxoglutarate carrier),
member 11 {Mus musculus;} , partial (25%)
Length = 773
Score = 54.3 bits (129), Expect = 5e-08
Identities = 47/172 (27%), Positives = 74/172 (42%), Gaps = 16/172 (9%)
Frame = +2
Query: 25 PLDVVQTRLQVHDGRLF-SQYPRYNNTTHAIFTIARKEGLKGLYAG----------FPAG 73
P DV R+Q DGRL +Q Y + AI +A++EG+ L+ G A
Sbjct: 35 PADVAMVRMQA-DGRLXXAQRRNYKSVVDAITRMAKQEGVTSLWRGSSLTVNRAMLVTAS 211
Query: 74 LLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCANPAFVVKTR-LQLQTPLHHA 132
L + + + G+++D A + +NP V+KTR + ++
Sbjct: 212 QLASYDQFKETILENGMMRDGLGTHVTASFAAGFVAAVASNPVDVIKTRVMNMRVEPGAT 391
Query: 133 RPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQ----AAIQFIVYEQLRK 180
PY+G D R EG A Y+G +P IS+ + F+ EQ+RK
Sbjct: 392 PPYAGALDCALKTVRAEGPMALYKGFIP---TISRQGPFTVVLFVTLEQVRK 538
>TC209762
Length = 950
Score = 52.4 bits (124), Expect = 2e-07
Identities = 52/195 (26%), Positives = 86/195 (43%), Gaps = 24/195 (12%)
Frame = +2
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
VA +T+ + P+DVV +L V +S + +Y+ + + R +G++GLY G
Sbjct: 20 VAGMTSSLFAQSVFVPIDVVSQKLMVQG---YSGHAQYSGGLDVVRQVLRTDGIRGLYRG 190
Query: 70 FPAGLLGATISWSLLVFY---------------YGIVKDRHARSK-----VEKSGALM-- 107
F GL T + + V++ +G D A S V+ +G ++
Sbjct: 191 F--GLSAITYAPASAVWWASYGSSQRFIWRFLDHGAKYDEVAPSLQKIMLVQATGGIIAG 364
Query: 108 -TVCLCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS 166
T P +KTRLQ+ H R S + + + E+G+ FYRG P FF +S
Sbjct: 365 ATSSCITTPLDTIKTRLQVMG--HENR--SSIKQVAKDLINEDGWRGFYRGFGPRFFSMS 532
Query: 167 QAAIQFIV-YEQLRK 180
I+ YE L++
Sbjct: 533 AWGTSMILTYEYLKR 577
>TC228373 similar to GB|AAM19967.1|20466091|AY098957 AT5g01340/T10O8_50
{Arabidopsis thaliana;} , partial (96%)
Length = 1424
Score = 51.6 bits (122), Expect = 3e-07
Identities = 52/169 (30%), Positives = 74/169 (43%), Gaps = 15/169 (8%)
Frame = +2
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLL 84
P +VV+ RLQ G L + +Y H I R+EG +GL+AG ++ + S +
Sbjct: 647 PFEVVKIRLQQQRG-LSPELLKYKGPVHCARMIIREEGFRGLWAGVAPTVMRNGKNQSAM 823
Query: 85 -----VFYYGIVKDRHARSKVEK------SGALMTVC--LCANPAFVVKTRLQLQTPLHH 131
F + K +V SG L +C P VVKTRL QT
Sbjct: 824 FTAKNAFDVLLWKKHEGDGRVLLPWQSMISGFLAGTAGPICTGPFDVVKTRLMAQTREGG 1003
Query: 132 A-RPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQFIVYEQL 178
Y G+ A RTI EEG A ++G++P I AI + V +Q+
Sbjct: 1004GVLKYKGMIHAIRTIYVEEGLLALWKGLLPRLMRIPPGQAIMWGVADQI 1150
Score = 49.7 bits (117), Expect = 1e-06
Identities = 39/148 (26%), Positives = 61/148 (40%), Gaps = 12/148 (8%)
Frame = +2
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLL 84
P+DV++TRLQ+ + Y H TI+R EG++ L+ G T+ ++L
Sbjct: 362 PIDVIKTRLQL------DRSGNYKGILHCGATISRTEGVRALWKGLTPFATHLTLKYALR 523
Query: 85 VFYYGIVKDRHARSKVEK-----------SGALMTVCLCANPAFVVKTRLQLQTPLH-HA 132
+ +++ + K ++ + P VVK RLQ Q L
Sbjct: 524 MGSNAVLQSAFKDPETGKLSGYGRILSGFGAGVLEAIIIVTPFEVVKIRLQQQRGLSPEL 703
Query: 133 RPYSGLYDAFRTIKREEGFSAFYRGIVP 160
Y G R I REEGF + G+ P
Sbjct: 704 LKYKGPVHCARMIIREEGFRGLWAGVAP 787
Score = 37.0 bits (84), Expect = 0.008
Identities = 29/96 (30%), Positives = 41/96 (42%), Gaps = 4/96 (4%)
Frame = +2
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
L W+ ++ G A P DVV+TRL R +Y HAI TI +EGL
Sbjct: 890 LPWQSMISGFLAGTAGPICTGPFDVVKTRLMAQT-REGGGVLKYKGMIHAIRTIYVEEGL 1066
Query: 64 KGLYAGFPAGLL----GATISWSLLVFYYGIVKDRH 95
L+ G L+ G I W + G+ + R+
Sbjct: 1067LALWKGLLPRLMRIPPGQAIMWGVADQIIGLYERRY 1174
Score = 36.2 bits (82), Expect = 0.014
Identities = 19/49 (38%), Positives = 25/49 (50%)
Frame = +2
Query: 112 CANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVP 160
C P V+KTRLQL + Y G+ TI R EG A ++G+ P
Sbjct: 353 CLQPIDVIKTRLQLD----RSGNYKGILHCGATISRTEGVRALWKGLTP 487
>TC206206 similar to PIR|T47703|T47703 Ca-dependent solute carrier-like
protein - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (38%)
Length = 772
Score = 49.7 bits (117), Expect = 1e-06
Identities = 29/75 (38%), Positives = 43/75 (56%), Gaps = 2/75 (2%)
Frame = +3
Query: 115 PAFVVKTRLQLQTPLHHARPYS-GLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQF 172
P +V+ R QL+ AR Y+ GLY FR I R EGF YRGI+P ++ ++ I F
Sbjct: 312 PLDLVRRRKQLEGAGGRARVYTTGLYGVFRHIIRTEGFRGLYRGILPEYYKVVPGVGICF 491
Query: 173 IVYEQLRKTVVNLKT 187
+ YE L+ + ++ T
Sbjct: 492 MTYETLKMLLADIAT 536
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.143 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,273,197
Number of Sequences: 63676
Number of extensions: 150098
Number of successful extensions: 1186
Number of sequences better than 10.0: 108
Number of HSP's better than 10.0 without gapping: 1050
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1128
length of query: 250
length of database: 12,639,632
effective HSP length: 95
effective length of query: 155
effective length of database: 6,590,412
effective search space: 1021513860
effective search space used: 1021513860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC125480.1


