

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
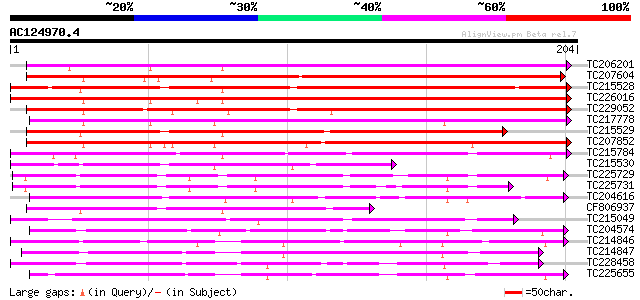
Query= AC124970.4 + phase: 0 /pseudo
(204 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC206201 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible trypsi... 181 3e-46
TC207604 similar to UP|P93378 (P93378) Tumor-related protein, pa... 180 3e-46
TC215528 weakly similar to UP|P93378 (P93378) Tumor-related prot... 179 6e-46
TC226016 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible trypsi... 168 2e-42
TC229052 weakly similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible... 164 3e-41
TC217778 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible trypsi... 162 1e-40
TC215529 weakly similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible... 161 2e-40
TC207852 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible trypsi... 161 2e-40
TC215784 weakly similar to UP|P93378 (P93378) Tumor-related prot... 129 7e-31
TC215530 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible trypsi... 86 2e-17
TC225729 similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor precur... 75 2e-14
TC225731 similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor precur... 71 3e-13
TC204616 similar to UP|O17170 (O17170) Serpentine receptor, clas... 70 5e-13
CF806937 69 1e-12
TC215049 UP|Q9LLX2 (Q9LLX2) Trypsin inhibitor, complete 67 7e-12
TC204574 similar to UP|KTI1_SOYBN (P25272) Kunitz-type trypsin i... 63 1e-10
TC214846 UP|KTI1_SOYBN (P25272) Kunitz-type trypsin inhibitor KT... 61 3e-10
TC214847 similar to UP|Q9XIS8 (Q9XIS8) Trypsin inhibitor p20, pa... 57 4e-09
TC228458 similar to UP|Q8W3K5 (Q8W3K5) Mcp20, partial (31%) 57 6e-09
TC225655 UP|Q8W3K5 (Q8W3K5) Mcp20, complete 57 6e-09
>TC206201 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible
trypsin-inhibitor-like protein, partial (65%)
Length = 880
Score = 181 bits (458), Expect = 3e-46
Identities = 96/208 (46%), Positives = 124/208 (59%), Gaps = 12/208 (5%)
Frame = +2
Query: 7 LLFLLLTLFTTKPL---QGAEQPEEVRDTSGNLVRNSINYFILPSS-------IQCGTRC 56
L FLL TTKPL PE V DTSG +R NY I+P+ + C T
Sbjct: 47 LAFLLFFALTTKPLLLGAAGAAPEPVIDTSGKKLRADANYHIIPAVPFTICGFVSCFTGG 226
Query: 57 EMALLNTNKTCPLDVVEEE--EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVT 114
++L + +++CPLDV+ E+ E + F P N KKGVIRVSTDLN+ S T
Sbjct: 227 GLSLDSIDESCPLDVIIEKANEGLPLRFSPVNTKKGVIRVSTDLNIFFSDSDERCPHHST 406
Query: 115 VWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCK 174
VW +D+ D + Q +VTTGGV GNPG T+ NWFKI+++E YKLV+CP VC C +CK
Sbjct: 407 VWMLDQFDASIGQTYVTTGGVVGNPGEHTILNWFKIQKYEDAYKLVYCPRVCPSCHHLCK 586
Query: 175 DIGIFLDENRNTRFVLSDFPFGVKFQRA 202
DIG+F+D NR LSD PF +KF+ A
Sbjct: 587 DIGMFVDANRRMHLALSDDPFKIKFKEA 670
>TC207604 similar to UP|P93378 (P93378) Tumor-related protein, partial (44%)
Length = 816
Score = 180 bits (457), Expect = 3e-46
Identities = 98/207 (47%), Positives = 131/207 (62%), Gaps = 13/207 (6%)
Frame = +1
Query: 7 LLFLLLTLFTTKPLQGAEQ--PEEVRDTSGNLVRNSINYFILP-SSIQC--------GTR 55
L FLL+ +F+++ L G P +V DT G VR ++Y+I P + C G+
Sbjct: 25 LAFLLIFVFSSQFLLGGADASPRQVIDTEGKKVRAGVDYYIRPVPTTPCDGRGPCVVGSG 204
Query: 56 CEMALLNTNKTCPLDV--VEEEEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
+ ++N TCPL V VE + +F P N KKGVIRVSTDLN+ S TN S S
Sbjct: 205 YVLIARSSNHTCPLSVAVVEGFRGLAVTFTPVNPKKGVIRVSTDLNIKTSL-TNTSCSES 381
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVC 173
TVWK+D D +T Q FVTTGGV GNPG++T+DNWFKIE ++ YKLVFCPTVC C+ +C
Sbjct: 382 TVWKLDAFDDSTGQWFVTTGGVLGNPGKDTIDNWFKIEEYDDDYKLVFCPTVCNFCKPLC 561
Query: 174 KDIGIFLDENRNTRFVLSDFPFGVKFQ 200
+++G+F D N N R L+D P+ V+FQ
Sbjct: 562 RNVGVFRDSNGNQRVALTDEPYKVRFQ 642
>TC215528 weakly similar to UP|P93378 (P93378) Tumor-related protein, partial
(57%)
Length = 763
Score = 179 bits (455), Expect = 6e-46
Identities = 102/206 (49%), Positives = 128/206 (61%), Gaps = 4/206 (1%)
Frame = +1
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAE--QPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEM 58
MK+ L L++ L +TK L GA PE+V DTSG +VR +Y+I+P+S G +
Sbjct: 34 MKMTLVTLVLIVAL-STKALLGAAGPAPEQVLDTSGKIVRARSSYYIVPASPDLGG---L 201
Query: 59 ALLNTNKTCPLDVVEEE--EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVW 116
+ +T CPLDVV + + F P NF KGVIRVSTDLN+ FP S TVW
Sbjct: 202 DMASTGADCPLDVVAVDGYQGQPLIFTPVNFNKGVIRVSTDLNIY--FPVATSCPQTTVW 375
Query: 117 KVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDI 176
K+ D +TSQ FVTTGG GNPG +TV NWFKIE++E YKLV+CP+VC +C C DI
Sbjct: 376 KLKDYDYSTSQWFVTTGGDFGNPGSQTVANWFKIEKYEDAYKLVYCPSVCNDCSYPCSDI 555
Query: 177 GIFLDENRNTRFVLSDFPFGVKFQRA 202
GI+ DE R LS P+ VKFQRA
Sbjct: 556 GIYQDE-YGKRLALSSEPYKVKFQRA 630
>TC226016 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible
trypsin-inhibitor-like protein, partial (85%)
Length = 853
Score = 168 bits (425), Expect = 2e-42
Identities = 94/209 (44%), Positives = 128/209 (60%), Gaps = 7/209 (3%)
Frame = +3
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQ--PEEVRDTSGNLVRNSINYFILPSS--IQCGTRC 56
MK L F L+ +++PL G + PE+V DT G +R NY+I+PS + T
Sbjct: 45 MKSTMLLAFALVLALSSQPLLGGAEASPEQVVDTLGKKLRVGTNYYIVPSLPYTKIRTTR 224
Query: 57 EMALLNTNKT-CPLDVVEEE--EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
+ L + K CPLDVV + +F P N KKGVIRVSTDLN+ S T+C
Sbjct: 225 GLGLASVGKPYCPLDVVVVNGYHGLPVTFSPVNPKKGVIRVSTDLNIKFSARTSCPRQYS 404
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVC 173
TVWK+D D + Q FVTTGGV GNP ET+ NWFKIE+++ YKLV+CP+V + + +C
Sbjct: 405 TVWKLDDFDFSKRQWFVTTGGVVGNPSLETIHNWFKIEKYDGAYKLVYCPSVVKCPKHLC 584
Query: 174 KDIGIFLDENRNTRFVLSDFPFGVKFQRA 202
K++G+F+DE N R L+D P V+FQ+A
Sbjct: 585 KNVGLFVDEKGNKRLALTDVPLKVQFQQA 671
>TC229052 weakly similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible
trypsin-inhibitor-like protein, partial (75%)
Length = 799
Score = 164 bits (414), Expect = 3e-41
Identities = 91/213 (42%), Positives = 136/213 (63%), Gaps = 17/213 (7%)
Frame = +1
Query: 7 LLFLLLTLFTTKPLQGAEQ--PEEVRDTSGNLVRNSINYFILPSSIQCGTRCE------- 57
L+F+LL ++PL GA PE V DTSG ++ ++Y+++P+ ++ TRC
Sbjct: 61 LVFVLLFGLVSQPLLGAVDALPEAVIDTSGTELQPGLSYYVVPA-MRSFTRCGKFECLNA 237
Query: 58 --MALLNTNKTCPLDVVEEEEA--MQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
++L + ++CPLDVV E+ + + SF P + + V+RVSTDLN++ F T+ ++ S
Sbjct: 238 EGLSLASIGESCPLDVVVEQRSFGLPLSFSPLDTNESVVRVSTDLNIM--FCTDRTSYSC 411
Query: 114 T----VWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCREC 169
VWK+D DV+ + FVTTGG GNP ET+ NWFKIE+ +S Y++V+CP+VC
Sbjct: 412 AEYSPVWKLDHFDVSKGKWFVTTGGSMGNPSWETIRNWFKIEKCDSAYRIVYCPSVCPSS 591
Query: 170 EVVCKDIGIFLDENRNTRFVLSDFPFGVKFQRA 202
+ +CKD+G+F+DEN R LSD PF VKFQ A
Sbjct: 592 KHMCKDVGVFVDENGYRRLALSDVPFKVKFQLA 690
>TC217778 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible
trypsin-inhibitor-like protein, partial (63%)
Length = 842
Score = 162 bits (410), Expect = 1e-40
Identities = 88/203 (43%), Positives = 122/203 (59%), Gaps = 8/203 (3%)
Frame = +3
Query: 8 LFLLLTLFTTKPLQGAEQ--PEEVRDTSGNLVRNSINYFILPSS--IQCGTRCEMALLNT 63
L L+L +++ L G + PE+V DTSG ++R +NY IL S C + + L
Sbjct: 63 LVLILLALSSQSLLGEVETSPEQVLDTSGKVLREGVNYNILISMPYTSCRSPQGLGLSKI 242
Query: 64 NKTCPLDVV--EEEEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKV 121
+CPLDVV + + F+P N KKGVIRV+TDLN++ TVWKVD
Sbjct: 243 GNSCPLDVVVVDINHRLPLRFIPVNPKKGVIRVATDLNIMFPDRNVTCPHHSTVWKVDNF 422
Query: 122 DVATSQRFVTTGGVQGNPGRETVDNWFKIERFES--GYKLVFCPTVCRECEVVCKDIGIF 179
V+ R VTTGGV G PGRET+ NWFKIE+++ YKLV+CP+VC C+ CK++G+
Sbjct: 423 HVSKGHRLVTTGGVVGYPGRETIGNWFKIEKYDGAYNYKLVYCPSVCPSCKHECKNVGMV 602
Query: 180 LDENRNTRFVLSDFPFGVKFQRA 202
+D+N N R LSD P+ +F +A
Sbjct: 603 VDQNGNQRLALSDVPYQFRFFKA 671
>TC215529 weakly similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible
trypsin-inhibitor-like protein, partial (46%)
Length = 551
Score = 161 bits (408), Expect = 2e-40
Identities = 84/177 (47%), Positives = 112/177 (62%), Gaps = 4/177 (2%)
Frame = +3
Query: 7 LLFLLLTLFTTKPLQGAE--QPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTN 64
+ LLL +TK L GA PE+V DTSG +VR +Y+I+P+S G +A+ +T
Sbjct: 36 IALLLLVALSTKALLGAAGPAPEQVLDTSGKMVRARTSYYIVPASPDVGG---LAMASTG 206
Query: 65 KTCPLDVVEEE--EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVD 122
+ CPLDVV + + F P N KGVIRVSTDLN+ T+C + WK+ D
Sbjct: 207 EDCPLDVVAVDGYQGQPLIFTPVNVNKGVIRVSTDLNIYFPIDTSCPLTKA--WKLKDYD 380
Query: 123 VATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIF 179
+TSQ FVTTGG GNPG +T+ NWFKIE++E YKLV+CP+VC++C C DIGI+
Sbjct: 381 YSTSQWFVTTGGDFGNPGSQTLANWFKIEKYEDAYKLVYCPSVCKDCSYPCSDIGIY 551
>TC207852 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible
trypsin-inhibitor-like protein, partial (80%)
Length = 893
Score = 161 bits (407), Expect = 2e-40
Identities = 94/211 (44%), Positives = 133/211 (62%), Gaps = 15/211 (7%)
Frame = +3
Query: 7 LLFLLLTLFTTKPLQGAEQ--PEEVRDTSGNLVRNSINYFILPSS--IQCGT--RCE--- 57
L F+LL +++PL A PE+V DTSG +R ++Y+I+P+ +CG RC
Sbjct: 69 LAFVLLFALSSQPLLAAADASPEQVVDTSGKKLRAGLSYYIVPAVPLTRCGRYERCMGGG 248
Query: 58 -MALLNTNKTCPLDVV--EEEEAMQFSFVPFNFKKGVIRVSTDLNVIHSFP-TNCSTSSV 113
++L + ++CPLDVV + F P + KKGV+RVSTDLN++ S T+C+ S
Sbjct: 249 GLSLASIGESCPLDVVVVPRSHGLPLQFSPVDPKKGVVRVSTDLNIMFSTDHTSCAEYS- 425
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTV--CRECEV 171
VWK+D DV+ + FV+TGG GNP ET+ NWFKIE+ + YK+V+CP+V +
Sbjct: 426 PVWKLDHFDVSKGKWFVSTGGSMGNPSWETIRNWFKIEKCDGAYKIVYCPSVFPSSSSKH 605
Query: 172 VCKDIGIFLDENRNTRFVLSDFPFGVKFQRA 202
+CKDIG+F+DEN R LS+ PF VKFQRA
Sbjct: 606 MCKDIGVFVDENGFRRLALSNVPFKVKFQRA 698
>TC215784 weakly similar to UP|P93378 (P93378) Tumor-related protein, partial
(57%)
Length = 870
Score = 129 bits (325), Expect = 7e-31
Identities = 87/211 (41%), Positives = 119/211 (56%), Gaps = 9/211 (4%)
Frame = +2
Query: 1 MKLHFPLLFLLLTL-FTTKPLQG--AEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCE 57
MK PL F +L L FT +P G A PE V DTSG +R + Y+ILP G
Sbjct: 32 MKKVSPLAFSILFLAFTIEPFIGIAAAAPEAVLDTSGQKLRTGVKYYILPVFRGKGGGLT 211
Query: 58 MALLNTNKTCPLDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
++ + N TCPL VV+E+ + +F P+N K GVI STDLN I S+ +
Sbjct: 212 VSS-SGNNTCPLFVVQEKLEVSKGTPVTFTPYNAKSGVILTSTDLN-IKSYGKTTTCDKP 385
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVC 173
VWK+ KV T F++TGGV+GNPG TV NWFKIE+ E Y L FCP+ + +C
Sbjct: 386 PVWKLLKV--LTGVWFLSTGGVEGNPGVNTVVNWFKIEKAEKDYVLSFCPSF---AQTLC 550
Query: 174 KDIGIFLDENRNTRFVLSDF--PFGVKFQRA 202
+++G+++ ++ N LSD F V F+RA
Sbjct: 551 RELGLYVGDDGNKHLSLSDKVPSFKVMFKRA 643
>TC215530 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible
trypsin-inhibitor-like protein, partial (19%)
Length = 427
Score = 85.5 bits (210), Expect = 2e-17
Identities = 61/142 (42%), Positives = 77/142 (53%), Gaps = 3/142 (2%)
Frame = +1
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
MK+ L L+ L T K L G E V D G VR Y+I+P+S G +A
Sbjct: 22 MKMTLVTLVLVFALIT-KALAGPAS-EPVLDALGKKVRADSIYYIVPASSDIGG---LAS 186
Query: 61 LNTNKTCPLDVV--EEEEAMQFSFVPFNFKKGVIRVSTDLNV-IHSFPTNCSTSSVTVWK 117
T+ CPLDVV + + + SF P N KKG+IRVSTDLN+ S+ C TVWK
Sbjct: 187 ARTDVDCPLDVVAVDGDLGLPLSFTPVNDKKGIIRVSTDLNIYFTSYTIFC--PQTTVWK 360
Query: 118 VDKVDVATSQRFVTTGGVQGNP 139
+ D +TSQ FVTTGG G+P
Sbjct: 361 LKDYDDSTSQWFVTTGGELGHP 426
>TC225729 similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor precursor, partial
(82%)
Length = 732
Score = 75.1 bits (183), Expect = 2e-14
Identities = 69/210 (32%), Positives = 97/210 (45%), Gaps = 10/210 (4%)
Frame = +3
Query: 2 KLH-FPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
K H PL F LL F TKPL AE PE V D GN + + Y++ P G + L
Sbjct: 75 KFHSLPLCFFLLLAFNTKPLLAAE-PEPVVDKQGNPLEPGVGYYVWPLWADEG---GLTL 242
Query: 61 LNT-NKTCPLDVVEEEEAMQFSFVPFNF-KKGVIRVSTDLNVIHSFPTNCSTSSVTVWKV 118
T NKTCPL V+ + F P +F G+ V T ++ FP + TVW++
Sbjct: 243 GQTRNKTCPLYVIRDP---SFIGTPVSFLAPGLDHVPTLTDLTIDFPVVTVCNQPTVWRL 413
Query: 119 DKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG-----YKLVFCPTVCRECEVVC 173
+K V + FV+T G + + + FKIER E Y FCP+V +C
Sbjct: 414 NK--VGSGFWFVSTSGDPND-----ITSKFKIERLEGDHAYEIYSFKFCPSV---PGALC 563
Query: 174 KDIGIFLDENRNTRFVLSD--FPFGVKFQR 201
+G F D + + D P+ V+FQ+
Sbjct: 564 APVGTFEDADGTKVMAVGDDIEPYYVRFQK 653
>TC225731 similar to UP|Q8RVX2 (Q8RVX2) Protease inhibitor precursor, partial
(76%)
Length = 583
Score = 71.2 bits (173), Expect = 3e-13
Identities = 65/188 (34%), Positives = 89/188 (46%), Gaps = 8/188 (4%)
Frame = +1
Query: 2 KLH-FPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
K H PL FLLL T+PL AE PE V D GN + + Y++ P G + L
Sbjct: 61 KFHSLPLCFLLLLAINTQPLLAAE-PEPVVDKQGNPLVPGVGYYVWPLWADNG---GLTL 228
Query: 61 LNT-NKTCPLDVVEEEEAMQFSFVPFNF-KKGVIRVSTDLNVIHSFPTNCSTSSVTVWKV 118
T NKTCPLDV+ + F P F G+ + T ++ FP + TVW++
Sbjct: 229 GQTRNKTCPLDVIRDP---SFIGSPVRFHASGLNHIPTLTDLTIDFPVVTVCNQPTVWRL 399
Query: 119 DKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG-----YKLVFCPTVCRECEVVC 173
K + FV+T +GNP + + FKIER E Y FCP+V V+C
Sbjct: 400 SK--EGSGFWFVST---RGNP--QDLITKFKIERLEGDHAYEIYSFKFCPSV---PGVLC 549
Query: 174 KDIGIFLD 181
+G F+D
Sbjct: 550 APVGTFVD 573
>TC204616 similar to UP|O17170 (O17170) Serpentine receptor, class i protein
29, partial (5%)
Length = 904
Score = 70.5 bits (171), Expect = 5e-13
Identities = 62/204 (30%), Positives = 96/204 (46%), Gaps = 10/204 (4%)
Frame = +2
Query: 8 LFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTC 67
L+ LTL + + + V DT+G V N Y+I P+ G R L+N N +C
Sbjct: 74 LYASLTLTVWLFMATFSRAQYVIDTNGEPVDNDDEYYIRPAITDNGGR--FTLINRNGSC 247
Query: 68 PLDVVEEEE------AMQFSFVPFNFKKGVIRVSTDLNV-IHSFPTNCSTSSVTVWKVDK 120
PL V E ++F+ N + IRV++DL + T C S T W+V +
Sbjct: 248 PLYVGLENTDTPLGYPVKFTHFALNVQDEDIRVNSDLRIEFVEVSTTCVQS--TEWRVGE 421
Query: 121 VDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG--YKLVFCP-TVCRECEVVCKDIG 177
D + +R + T G+ N G ++ N+F+I +S Y + +CP +C +C VC G
Sbjct: 422 NDTRSGRRLIIT-GLDDNFG--SIGNYFRIVETQSVGIYNIEWCPMEICSDCGFVCSTGG 592
Query: 178 IFLDENRNTRFVLSDFPFGVKFQR 201
I ++ R F L P V FQ+
Sbjct: 593 ILREDGR-IFFALDGTPLPVVFQK 661
>CF806937
Length = 375
Score = 69.3 bits (168), Expect = 1e-12
Identities = 49/129 (37%), Positives = 64/129 (48%), Gaps = 4/129 (3%)
Frame = +2
Query: 7 LLFLLLTLFTTKPLQGAE--QPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTN 64
+ LLL +TK L GA P +V DTS N+VR Y+I P+S +A+ T
Sbjct: 2 IALLLLVALSTKALLGAXGPAPXQVLDTSXNMVRAXTTYYIXPASPXVR---GLAMAXTX 172
Query: 65 KTCPLDVVEEE--EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVD 122
+ CPLDVV + + F P N K VIRVSTDLN+ T C + + D
Sbjct: 173 ENCPLDVVPVDGYQGHPLIFTPVNVNKRVIRVSTDLNIYFPIDTXCPLTKXC--NLXDYD 346
Query: 123 VATSQRFVT 131
+TS FVT
Sbjct: 347 NSTSHWFVT 373
>TC215049 UP|Q9LLX2 (Q9LLX2) Trypsin inhibitor, complete
Length = 936
Score = 66.6 bits (161), Expect = 7e-12
Identities = 57/188 (30%), Positives = 83/188 (43%), Gaps = 5/188 (2%)
Frame = +2
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
+ L F LF L L +E E+V D SGN + Y+I+PS+
Sbjct: 65 LSLSFLPLFAFLAL--------SEDVEQVVDISGNPIFPGGTYYIMPSTWGAAGGGLKLG 220
Query: 61 LNTNKTCPLDVVEEEEAMQFSFVPFNFK-----KGVIRVSTDLNVIHSFPTNCSTSSVTV 115
N CP+ V+++ + F P F G+I T L + + C+ SS V
Sbjct: 221 RTGNSNCPVTVLQDYSEI-FRGTPVKFSIPGISPGIIFTGTPLEIEFAEKPYCAESSKWV 397
Query: 116 WKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKD 175
VD + V GG +G+PG++T F I++++ GYKLVFC T C D
Sbjct: 398 AFVDN---EIQKACVGIGGPEGHPGQQTFSGTFSIQKYKFGYKLVFCIT----GSGTCLD 556
Query: 176 IGIFLDEN 183
IG F +N
Sbjct: 557 IGRFDAKN 580
>TC204574 similar to UP|KTI1_SOYBN (P25272) Kunitz-type trypsin inhibitor
KTI1 precursor, partial (34%)
Length = 898
Score = 62.8 bits (151), Expect = 1e-10
Identities = 58/205 (28%), Positives = 90/205 (43%), Gaps = 11/205 (5%)
Frame = +3
Query: 8 LFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTC 67
+FLL + P A+ +V D G+ +RN Y++LP+ G E A L + TC
Sbjct: 78 IFLLCAFTSYLPSATAQ---DVLDVDGDPIRNGFIYYVLPAIRGNGGGIERAALGKD-TC 245
Query: 68 PLDVVEE----EEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDV 123
P+ VV+ + ++ F + I + L + S+P C + W + K D+
Sbjct: 246 PITVVQSPNPNSKGLEIKFESA-YPAYYINETLILQIKFSYPQQCERKNPW-WAISK-DI 416
Query: 124 ATSQRFVTTGGVQGNPGRETVDNWFKIERFESG-----YKLVFCPTVCRECEVVCKDIGI 178
+ + G G T WFKI++ YKLVFC + E C D+GI
Sbjct: 417 SEGPPAIKLSGFHG-----TELGWFKIQKASKSCDSNDYKLVFC----QYDETWCLDVGI 569
Query: 179 FLDENRNTRFVL--SDFPFGVKFQR 201
++D N R VL + PF V F +
Sbjct: 570 YVDRQGNRRLVLAVTGEPFLVHFHK 644
>TC214846 UP|KTI1_SOYBN (P25272) Kunitz-type trypsin inhibitor KTI1
precursor, complete
Length = 1027
Score = 61.2 bits (147), Expect = 3e-10
Identities = 65/212 (30%), Positives = 95/212 (44%), Gaps = 11/212 (5%)
Frame = +1
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
MK L+ FT L A + V DT + ++N Y++LP + G + +
Sbjct: 31 MKSTIFFALFLVCAFTISYLPSATA-QFVLDTDDDPLQNGGTYYMLP--VMRGKGGGIEV 201
Query: 61 LNTNKT-CPLDVVEEEEAMQFSFVPFNFKKGVIRVSTD----LNVIHSFPTNCSTSSVTV 115
+T K CPL VV+ P KG+ V T L + +P + S V
Sbjct: 202 DSTGKEICPLTVVQS---------PNELDKGIGLVFTSPLHALFIAEGYPLSIKFGSFAV 354
Query: 116 WKVDKVDVATSQRFVTTGGVQGNP--GRETVDNWFKIERFE---SGYKLVFCPTVCRECE 170
+ + T V G+Q R+TVD WF IER + YKLVFCP + +
Sbjct: 355 ITLC-AGMPTEWAIVEREGLQAVKLAARDTVDGWFNIERVSREYNDYKLVFCPQQAEDNK 531
Query: 171 VVCKDIGIFLDENRNTRFVLS-DFPFGVKFQR 201
C+DIGI +D++ R VLS + P V+FQ+
Sbjct: 532 --CEDIGIQIDDDGIRRLVLSKNKPLVVQFQK 621
>TC214847 similar to UP|Q9XIS8 (Q9XIS8) Trypsin inhibitor p20, partial (75%)
Length = 811
Score = 57.4 bits (137), Expect = 4e-09
Identities = 55/195 (28%), Positives = 82/195 (41%), Gaps = 7/195 (3%)
Frame = +3
Query: 5 FPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTN 64
F LFL+ + P A+ + D +G+ VRN Y+ILP G E A N
Sbjct: 75 FFALFLVCAFTSYLPSATADT---IFDINGDFVRNGGTYYILPVIRGDGGGIEFAATG-N 242
Query: 65 KTCPLDVVEEEEAMQFSFVPFNFKKGVIRVSTD----LNVIHSFPTNCSTSSVTVWKVDK 120
+TCPL VV+ P KG+ + + L++ N + V +
Sbjct: 243 ETCPLTVVQS---------PLEVSKGLPLIISSPFEILSIQEGLILNIGFTFVPPCALIP 395
Query: 121 VDVATSQRFVTTGGVQGNPGRETVDNWFKIERFE---SGYKLVFCPTVCRECEVVCKDIG 177
+ T + V+ V WFKIER + YKLVFC T + C DIG
Sbjct: 396 SEWTTVKGLPEGLAVKLTGYENKVPGWFKIERVSLEFNDYKLVFCATE----DSTCVDIG 563
Query: 178 IFLDENRNTRFVLSD 192
+++D N R V+++
Sbjct: 564 VYIDGEGNRRLVVTE 608
>TC228458 similar to UP|Q8W3K5 (Q8W3K5) Mcp20, partial (31%)
Length = 786
Score = 57.0 bits (136), Expect = 6e-09
Identities = 60/200 (30%), Positives = 83/200 (41%), Gaps = 8/200 (4%)
Frame = +3
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
MK LFLL L + P A +V DT GN V N YF+LP+ I G E A
Sbjct: 24 MKSTLFALFLLCALTSYLPSATAG---DVVDTDGNPVENGGTYFVLPTIILNGGGIEYAT 194
Query: 61 LNTNKTCPLDVVEEEEAMQFSFVPFNFKKGV----IRVSTDLNVIHSFPTNCSTSSVTVW 116
N+TCP+ V + + F P I LN+ +F + CS +S W
Sbjct: 195 FG-NETCPVTVAQSRDQFCKGF-PITISSPARIRHISEGLSLNIGFTFASPCSPAS--EW 362
Query: 117 KVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFES----GYKLVFCPTVCRECEVV 172
+ K D TG + TV F ++R + GY ++FCP C V
Sbjct: 363 TIVK-DQPEGLAVKLTG------FKNTVPGVFTLKRVPADEIIGYNILFCPLDNNPCGYV 521
Query: 173 CKDIGIFLDENRNTRFVLSD 192
+ D+ RN R V+S+
Sbjct: 522 ----AVHFDQFRNRRLVVSE 569
>TC225655 UP|Q8W3K5 (Q8W3K5) Mcp20, complete
Length = 960
Score = 57.0 bits (136), Expect = 6e-09
Identities = 61/212 (28%), Positives = 90/212 (41%), Gaps = 18/212 (8%)
Frame = +1
Query: 8 LFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTC 67
LFLL L T Q + + V DT GN +RN Y++LP G E A T +TC
Sbjct: 97 LFLLCAL--TSSYQPSATADIVFDTEGNPIRNGGTYYVLPVIRGKGGGIEFAKTET-ETC 267
Query: 68 PLDVVEEEEAMQFSFVPFNFKKGV------------IRVSTDLNVIHSFPTNCSTSSVTV 115
PL VV+ PF KG+ I L++ ++ C+ S+ +
Sbjct: 268 PLTVVQS---------PFEVSKGLPLIISSPFKILDITEGLILSLSFTYVPPCA-STPSR 417
Query: 116 WKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG---YKLVFCPTVCRECEVV 172
W V + TG + T+D WF+I+R S YKLVFC + +
Sbjct: 418 WTVILKGLPEELHVKLTG------YKNTIDGWFRIQRASSESNYYKLVFCTS---NDDSS 570
Query: 173 CKDIGIFLDENRNTRFVLS---DFPFGVKFQR 201
C DI +D N +++ + P V+FQ+
Sbjct: 571 CGDIVAPIDREGNRPLIVTHDQNHPLLVQFQK 666
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.139 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,515,843
Number of Sequences: 63676
Number of extensions: 148842
Number of successful extensions: 1073
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 1015
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1019
length of query: 204
length of database: 12,639,632
effective HSP length: 93
effective length of query: 111
effective length of database: 6,717,764
effective search space: 745671804
effective search space used: 745671804
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC124970.4


