

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124967.9 - phase: 0
(122 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
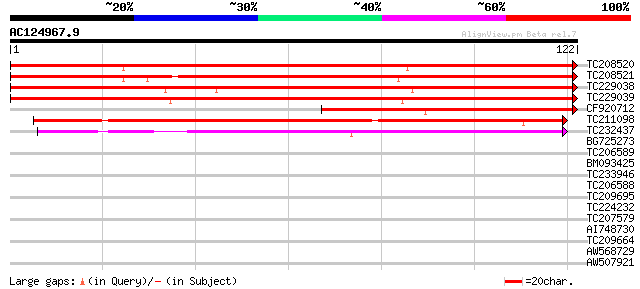
Score E
Sequences producing significant alignments: (bits) Value
TC208520 similar to UP|Q6ZKQ7 (Q6ZKQ7) Auxin-induced protein-rel... 196 1e-51
TC208521 similar to UP|Q6ZKQ7 (Q6ZKQ7) Auxin-induced protein-rel... 194 5e-51
TC229038 similar to UP|Q6ZKQ7 (Q6ZKQ7) Auxin-induced protein-rel... 120 7e-29
TC229039 113 1e-26
CF920712 86 3e-18
TC211098 similar to UP|Q9LSI2 (Q9LSI2) Gb|AAF27147.1, partial (27%) 76 2e-15
TC232437 similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, partial ... 39 3e-04
BG725273 similar to PIR|F86331|F863 F6F9.11 protein - Arabidopsi... 35 0.004
TC206589 similar to UP|Q6NMZ3 (Q6NMZ3) At4g34750, partial (39%) 35 0.005
BM093425 homologue to GP|24943206|gb auxin-regulated protein {Ph... 34 0.011
TC233946 UP|Q7R3F4 (Q7R3F4) GLP_158_77883_78482, partial (5%) 33 0.018
TC206588 weakly similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, p... 32 0.041
TC209695 similar to UP|Q9LTV3 (Q9LTV3) Auxin-regulated protein-l... 29 0.35
TC224232 homologue to UP|Q8H9B5 (Q8H9B5) UDP-glucose:sterol 3-O-... 27 1.3
TC207579 similar to GB|AAN15531.1|23198008|BT000212 expressed pr... 27 1.7
AI748730 weakly similar to GP|10178137|dbj leucoanthocyanidin di... 26 2.9
TC209664 UP|Q6T2Z7 (Q6T2Z7) Cyclin d2, partial (69%) 26 2.9
AW568729 26 2.9
AW507921 26 2.9
>TC208520 similar to UP|Q6ZKQ7 (Q6ZKQ7) Auxin-induced protein-related-like
protein, partial (26%)
Length = 726
Score = 196 bits (499), Expect = 1e-51
Identities = 98/127 (77%), Positives = 111/127 (87%), Gaps = 5/127 (3%)
Frame = +2
Query: 1 MAKGGKLMKLKSALKKWNSFGNG-KQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRV 59
MA+GGKLMKLKS LKKWNSFGNG K SRH S+ +D SSSRSDLH V+VGKSRRLYRV
Sbjct: 11 MARGGKLMKLKSVLKKWNSFGNGSKHSRHHSSSAVADDESSSRSDLHAVYVGKSRRLYRV 190
Query: 60 TSDVVDNPVFRELVERSRETEQQND----TVNVVACEVVLFEHLLWMLENADPQPESLDE 115
+SDVVD+PVFRELVERSR+++QQ + T+NVVACEVVLFEHLLWML+NADPQPESL+E
Sbjct: 191 SSDVVDHPVFRELVERSRDSDQQQNEDTTTINVVACEVVLFEHLLWMLDNADPQPESLNE 370
Query: 116 LVDFYAC 122
LVDFY C
Sbjct: 371 LVDFYGC 391
>TC208521 similar to UP|Q6ZKQ7 (Q6ZKQ7) Auxin-induced protein-related-like
protein, partial (50%)
Length = 774
Score = 194 bits (493), Expect = 5e-51
Identities = 103/128 (80%), Positives = 113/128 (87%), Gaps = 6/128 (4%)
Frame = +3
Query: 1 MAKGGKLMKLKSALKKWNSFGNG-KQSRH-SISAVADEDSSSSRSDLHTVFVGKSRRLYR 58
MA+GGKL KLKS LKKWNSFGN K SRH SISAVAD D SSSRSDLH V+VGKSRRLYR
Sbjct: 99 MARGGKLTKLKSVLKKWNSFGNNSKHSRHHSISAVAD-DESSSRSDLHAVYVGKSRRLYR 275
Query: 59 VTSDVVDNPVFRELVERSRETEQQ----NDTVNVVACEVVLFEHLLWMLENADPQPESLD 114
V SDVVD+PVFRELVERSR+++QQ + T+NVVACEVVLFEHLLWML+NADPQPESL+
Sbjct: 276 VASDVVDHPVFRELVERSRDSDQQQQSEDTTINVVACEVVLFEHLLWMLDNADPQPESLN 455
Query: 115 ELVDFYAC 122
ELVDFYAC
Sbjct: 456 ELVDFYAC 479
>TC229038 similar to UP|Q6ZKQ7 (Q6ZKQ7) Auxin-induced protein-related-like
protein, partial (50%)
Length = 589
Score = 120 bits (302), Expect = 7e-29
Identities = 70/132 (53%), Positives = 91/132 (68%), Gaps = 10/132 (7%)
Frame = +2
Query: 1 MAKGGKLMKLKSALKKWNSFGNGKQSRHSISA---VADEDSSSSRS-----DLHTVFVGK 52
MAK GKL KLKSA+K+ S ++ S+S+ V+ SSSS+ LH V+VGK
Sbjct: 71 MAKVGKLTKLKSAIKRLPSLTKLSRNSSSVSSNKRVSGGTSSSSKEHEQEQQLHAVYVGK 250
Query: 53 SRRLYRVTSDVVDNPVFRELVERSRETEQQNDT--VNVVACEVVLFEHLLWMLENADPQP 110
SRR Y V S+V+D+PVF+ELV+RS T +D V VV+CEVVLFEHLLWMLE+ + Q
Sbjct: 251 SRRRYLVNSEVIDHPVFQELVDRSCSTSSHDDEDGVVVVSCEVVLFEHLLWMLESEETQL 430
Query: 111 ESLDELVDFYAC 122
S+DELV+FY+C
Sbjct: 431 GSMDELVEFYSC 466
>TC229039
Length = 795
Score = 113 bits (283), Expect = 1e-26
Identities = 63/139 (45%), Positives = 88/139 (62%), Gaps = 17/139 (12%)
Frame = +1
Query: 1 MAKGGKLMKLKSALKKWNSFGNGKQSRHSISAV-----------ADEDSSSSRSDLHTVF 49
MAK GKL KLKSA+K+W S ++ S+S+ + ++ + L V+
Sbjct: 124 MAKVGKLTKLKSAIKRWPSLTRLSRNSSSVSSNKGVSGKGTSLNSSKEHEHEQEQLRAVY 303
Query: 50 VGKSRRLYRVTSDVVDNPVFRELVERSRETEQQN------DTVNVVACEVVLFEHLLWML 103
VGKSRR Y V S+V+D+PVF+ELV+RS + + D V VV+CEVVLFEHLLWML
Sbjct: 304 VGKSRRRYLVNSEVIDHPVFQELVDRSCSSNSSSSHHHDDDGVVVVSCEVVLFEHLLWML 483
Query: 104 ENADPQPESLDELVDFYAC 122
E+ + + S+DELV+FY+C
Sbjct: 484 ESEETKLGSMDELVEFYSC 540
>CF920712
Length = 365
Score = 85.5 bits (210), Expect = 3e-18
Identities = 41/56 (73%), Positives = 46/56 (81%), Gaps = 1/56 (1%)
Frame = -3
Query: 68 VFRELVERSRETEQQNDTVNV-VACEVVLFEHLLWMLENADPQPESLDELVDFYAC 122
+FRELV RSR+ ++ D + VACEVVLFEHLLWML NADPQPESLDEL DFYAC
Sbjct: 363 LFRELVCRSRDGSKEEDEATINVACEVVLFEHLLWMLHNADPQPESLDELADFYAC 196
>TC211098 similar to UP|Q9LSI2 (Q9LSI2) Gb|AAF27147.1, partial (27%)
Length = 578
Score = 76.3 bits (186), Expect = 2e-15
Identities = 43/117 (36%), Positives = 72/117 (60%), Gaps = 2/117 (1%)
Frame = +1
Query: 6 KLMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVD 65
K L L + +SF N +S+ + + + + TVFVG +R+ Y ++S ++
Sbjct: 88 KCKNLSKHLGRSSSF-NSLRSKFAKEDLWEGNGMQEGEHCETVFVGSTRKRYVISSKYLN 264
Query: 66 NPVFRELVERSRETEQQNDTVNVVACEVVLFEHLLWMLENADPQ--PESLDELVDFY 120
+P+ + L+ S++ + +++V VV CEVVLF+HLLWMLENADP+ +SL+EL + Y
Sbjct: 265 HPLLKALINNSKQ-KGSDESVLVVNCEVVLFDHLLWMLENADPKFGSDSLEELAELY 432
>TC232437 similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, partial (71%)
Length = 587
Score = 39.3 bits (90), Expect = 3e-04
Identities = 30/115 (26%), Positives = 53/115 (46%), Gaps = 1/115 (0%)
Frame = +1
Query: 7 LMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDN 66
+++L+ L++W S + S H I S + V VG + + + V + +++
Sbjct: 52 IVRLRQMLRRWRS--KARTSAHRIP-------SDVPAGHVAVCVGNNSKRFVVRTTYLNH 204
Query: 67 PVFREL-VERSRETEQQNDTVNVVACEVVLFEHLLWMLENADPQPESLDELVDFY 120
PVF+ L VE E N + C+ +FE LL + ++D L +DFY
Sbjct: 205 PVFKRLLVEAEEEYGFSNHGPLAIPCDEAIFEQLLRFVSHSDDCHVPLRNNLDFY 369
>BG725273 similar to PIR|F86331|F863 F6F9.11 protein - Arabidopsis thaliana,
partial (56%)
Length = 395
Score = 35.4 bits (80), Expect = 0.004
Identities = 27/102 (26%), Positives = 47/102 (45%), Gaps = 1/102 (0%)
Frame = +2
Query: 7 LMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDN 66
+++L+ L++W S + S VA V VG + R + V + +++
Sbjct: 56 IVRLRQMLRRWRSKAHRIPSDVPAVHVA-------------VCVGTNSRRFVVRATYLNH 196
Query: 67 PVFREL-VERSRETEQQNDTVNVVACEVVLFEHLLWMLENAD 107
PVF++L VE E N + + C+ LFE LL + +D
Sbjct: 197 PVFKKLLVEAEEEYGFSNHGLLAIPCDEALFEQLLRFISRSD 322
>TC206589 similar to UP|Q6NMZ3 (Q6NMZ3) At4g34750, partial (39%)
Length = 686
Score = 35.0 bits (79), Expect = 0.005
Identities = 25/73 (34%), Positives = 37/73 (50%), Gaps = 1/73 (1%)
Frame = +3
Query: 48 VFVGKSRRLYRVTSDVVDNPVFRE-LVERSRETEQQNDTVNVVACEVVLFEHLLWMLENA 106
V VG SRR + V + +++P+F+ LV+ E N + C+ LFEHLL ++
Sbjct: 180 VCVGPSRRRFIVRATHLNHPIFKMLLVKAEEEYGFCNHGPLAIPCDESLFEHLLRVVARP 359
Query: 107 DPQPESLDELVDF 119
P P L DF
Sbjct: 360 VPLP-GFSSLEDF 395
>BM093425 homologue to GP|24943206|gb auxin-regulated protein {Phaseolus
vulgaris}, partial (73%)
Length = 420
Score = 33.9 bits (76), Expect = 0.011
Identities = 23/95 (24%), Positives = 46/95 (48%), Gaps = 8/95 (8%)
Frame = +2
Query: 34 ADEDSSSSRSDLH-------TVFVGKSRRLYRVTSDVVDNPVFRELVERSRETEQQNDTV 86
+DED S H V+VG R + + + + + +F+ L+E++ E + +
Sbjct: 116 SDEDGCYSPQPPHDVPKGYLAVYVGPQLRRFIIPTSYLSHSLFKALLEKAAEEFGFDQSG 295
Query: 87 NV-VACEVVLFEHLLWMLENADPQPESLDELVDFY 120
+ + CE+ F++LL +EN D L L+ ++
Sbjct: 296 GLTIPCEIETFKYLLNCIENHDDSSSKLIMLLYYH 400
>TC233946 UP|Q7R3F4 (Q7R3F4) GLP_158_77883_78482, partial (5%)
Length = 677
Score = 33.1 bits (74), Expect = 0.018
Identities = 23/95 (24%), Positives = 46/95 (48%), Gaps = 1/95 (1%)
Frame = +2
Query: 18 NSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVERS- 76
NS G+G R + + + + V+VG R + + + ++P+F+ L++ +
Sbjct: 128 NSGGSGSYGRRL--STSKKKKKKAPQGCICVYVGAERERFVIKVKIANHPLFKALLDAAE 301
Query: 77 RETEQQNDTVNVVACEVVLFEHLLWMLENADPQPE 111
RE +N+ + C+V LF L +EN+ + E
Sbjct: 302 REYGYRNNGPLWLPCDVDLFSEALKDMENSIQEEE 406
>TC206588 weakly similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, partial (44%)
Length = 633
Score = 32.0 bits (71), Expect = 0.041
Identities = 24/73 (32%), Positives = 36/73 (48%), Gaps = 1/73 (1%)
Frame = +1
Query: 48 VFVGKSRRLYRVTSDVVDNPVFRE-LVERSRETEQQNDTVNVVACEVVLFEHLLWMLENA 106
V VG SRR + V + +++P+F+ LV+ E N + C+ LFE LL ++
Sbjct: 214 VCVGPSRRRFIVRATHLNHPIFKMLLVKAEEEYGFCNHGPLAIPCDESLFEELLRVVSRP 393
Query: 107 DPQPESLDELVDF 119
P P L DF
Sbjct: 394 VPVP-GFSTLEDF 429
>TC209695 similar to UP|Q9LTV3 (Q9LTV3) Auxin-regulated protein-like
(At3g12830), partial (70%)
Length = 844
Score = 28.9 bits (63), Expect = 0.35
Identities = 20/73 (27%), Positives = 39/73 (53%), Gaps = 1/73 (1%)
Frame = +3
Query: 48 VFVGKSRRLYRVTSDVVDNPVFRELV-ERSRETEQQNDTVNVVACEVVLFEHLLWMLENA 106
++VG + V ++++++PVF +L+ E ++E + V + C V +FE +L L
Sbjct: 423 IYVGDEMERFVVCAELLNHPVFVKLLNESAQEYGYEQKGVLRLPCRVFVFERVLDAL-RL 599
Query: 107 DPQPESLDELVDF 119
+ ELV+F
Sbjct: 600 GLNARDIAELVNF 638
>TC224232 homologue to UP|Q8H9B5 (Q8H9B5) UDP-glucose:sterol
3-O-glucosyltransferase, partial (4%)
Length = 507
Score = 26.9 bits (58), Expect = 1.3
Identities = 20/64 (31%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Frame = +2
Query: 21 GNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRL-YRVTSDVVDNPVFRELVERSRET 79
GNGK S S+VA+ SSSS S + + K L ++ D ++ + +ERS+
Sbjct: 233 GNGKDMEDSTSSVANGTSSSSASGILGKGLSKVTTLPADISQDKSESSSSKFKMERSKTE 412
Query: 80 EQQN 83
Q++
Sbjct: 413 RQRH 424
>TC207579 similar to GB|AAN15531.1|23198008|BT000212 expressed protein
{Arabidopsis thaliana;} , partial (57%)
Length = 1249
Score = 26.6 bits (57), Expect = 1.7
Identities = 13/22 (59%), Positives = 17/22 (77%)
Frame = -2
Query: 29 SISAVADEDSSSSRSDLHTVFV 50
++SAV E SSSSRSD +VF+
Sbjct: 438 AVSAVPSEVSSSSRSDFISVFL 373
>AI748730 weakly similar to GP|10178137|dbj leucoanthocyanidin
dioxygenase-like protein {Arabidopsis thaliana}, partial
(21%)
Length = 405
Score = 25.8 bits (55), Expect = 2.9
Identities = 11/25 (44%), Positives = 17/25 (68%), Gaps = 1/25 (4%)
Frame = -2
Query: 94 VLFEHLLWMLENADPQ-PESLDELV 117
+LF HL W++E PQ P +L +L+
Sbjct: 377 ILFLHLKWLVEELTPQFPRTLHQLM 303
>TC209664 UP|Q6T2Z7 (Q6T2Z7) Cyclin d2, partial (69%)
Length = 903
Score = 25.8 bits (55), Expect = 2.9
Identities = 10/18 (55%), Positives = 12/18 (66%)
Frame = -2
Query: 93 VVLFEHLLWMLENADPQP 110
V L HL+W+LE PQP
Sbjct: 140 VELLSHLIWLLEIHTPQP 87
>AW568729
Length = 221
Score = 25.8 bits (55), Expect = 2.9
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = +2
Query: 81 QQNDTVNVVACEVVLFEHLLWMLEN 105
Q V V + +LFEH+LW++ N
Sbjct: 104 QSERRVIFVDVDDILFEHMLWLMHN 178
>AW507921
Length = 445
Score = 25.8 bits (55), Expect = 2.9
Identities = 10/18 (55%), Positives = 12/18 (66%)
Frame = -3
Query: 93 VVLFEHLLWMLENADPQP 110
V L HL+W+LE PQP
Sbjct: 137 VELLSHLIWLLEINTPQP 84
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.130 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,987,543
Number of Sequences: 63676
Number of extensions: 51762
Number of successful extensions: 304
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 291
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 291
length of query: 122
length of database: 12,639,632
effective HSP length: 98
effective length of query: 24
effective length of database: 6,399,384
effective search space: 153585216
effective search space used: 153585216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC124967.9


