

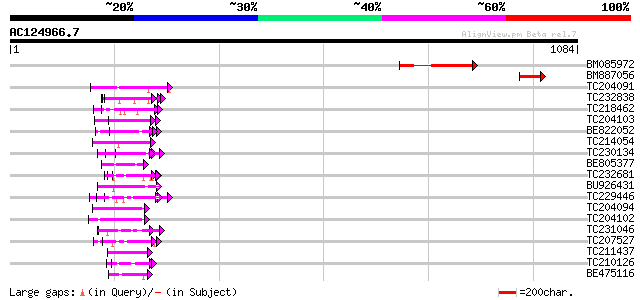
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124966.7 + phase: 0 /pseudo
(1084 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BM085972 166 7e-41
BM887056 86 9e-17
TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 65 2e-10
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 59 9e-09
TC218462 57 3e-08
TC204103 similar to UP|Q75QC2 (Q75QC2) Glutamate-rich protein, p... 52 1e-06
BE822052 similar to GP|4557063|gb|A expressed protein {Arabidops... 50 5e-06
TC214054 similar to UP|Q75QC2 (Q75QC2) Glutamate-rich protein, p... 50 7e-06
TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Pro... 49 9e-06
BE805377 similar to GP|21648010|gb| peptidase M23/M37 family {C... 49 2e-05
TC232681 weakly similar to GB|AAP21377.1|30102918|BT006569 At1g4... 49 2e-05
BU926431 48 2e-05
TC229446 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 47 3e-05
TC204094 weakly similar to UP|Q75QC2 (Q75QC2) Glutamate-rich pro... 47 3e-05
TC204102 similar to UP|Q75QC2 (Q75QC2) Glutamate-rich protein, p... 46 8e-05
TC231046 46 8e-05
TC207527 weakly similar to UP|O08661 (O08661) Ryanodine receptor... 45 1e-04
TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%) 45 2e-04
TC210126 similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C2... 45 2e-04
BE475116 45 2e-04
>BM085972
Length = 414
Score = 166 bits (419), Expect = 7e-41
Identities = 77/150 (51%), Positives = 99/150 (65%)
Frame = +1
Query: 745 ALQKLDEGCGIEEAKAICEPEIIRQLFTWQKQLKIYLAPFLHGRRYTSFGRHFTKIDKLK 804
AL+KL+ GC I++ KA C+P+ ++QLF W
Sbjct: 58 ALRKLESGCSIQDVKAFCDPDDLKQLFKW------------------------------- 144
Query: 805 EIVDRLHWYVQSGDTVLDFCCGANDFSCLMKSKLEQTGKLCSFKNYDLFQAKNDFNFEKR 864
+IVD+LHWYVQ+ DT++DFCCGANDFS LMK KLE+ GK CS++NYDL KNDF+FE+R
Sbjct: 145 KIVDKLHWYVQNSDTIVDFCCGANDFSILMKKKLEENGKKCSYRNYDLLPTKNDFSFERR 324
Query: 865 DWMSVQAEELPHGSNLIIGLNPPFGVRGSL 894
DWM+VQ ELP GS LI+GLNPPFG + +L
Sbjct: 325 DWMTVQPTELPTGSQLIMGLNPPFGHKAAL 414
>BM887056
Length = 447
Score = 85.9 bits (211), Expect = 9e-17
Identities = 35/48 (72%), Positives = 39/48 (80%)
Frame = +1
Query: 976 GKSFYLPGSVDTRDKQLEDWNLKPPPLYLWSRPDWTAWHMQIARMHRH 1023
GKSFYLPGSVD D+Q++ WN+KPPPLYLWSRPDWT H IAR H H
Sbjct: 1 GKSFYLPGSVDANDRQIDQWNVKPPPLYLWSRPDWTDKHKAIARKHGH 144
>TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(70%)
Length = 2075
Score = 64.7 bits (156), Expect = 2e-10
Identities = 49/168 (29%), Positives = 82/168 (48%), Gaps = 11/168 (6%)
Frame = +2
Query: 154 LISEAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEED 213
L+ + R + + T LK ENP QE + VE E E+ E E + VE EE+
Sbjct: 14 LVMPPMGRKRKVAPKSSLSTEPPLKQQQPKPENPPQEYEEVEEEVEEEEVEEE-VEEEEE 190
Query: 214 PEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVE------SAE 267
E+E++ E EEE+ E+END+V+ ++ ++E+D+ S +P ++ ++ +
Sbjct: 191 EEEEDEEEEEEEEEEEEENDVVKQQQQQQQDEDDIPISKLLEPLGKDQLLNLLCDAAAKH 370
Query: 268 EDPEEENDK-SDGEGVLNLDEEQDIGYDTVCAICDNG----GEILPCE 310
D EE K +DG+ V +G+DT + GEI C+
Sbjct: 371 RDVEERIRKAADGDPVHRKIFVHGLGWDTTAGTLISAFRQYGEIEDCK 514
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 59.3 bits (142), Expect = 9e-09
Identities = 38/126 (30%), Positives = 64/126 (50%), Gaps = 9/126 (7%)
Frame = +2
Query: 176 KLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMV 235
K+K E E E EN++ EED EE + E +D E + D E +++D +ND
Sbjct: 218 KMKATKELSEQVE-ENEVKVLVEEDDEEYESVDEVVDDAEDDEDEEEDDDDDEGDDNDDE 394
Query: 236 E---------SAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLD 286
E +ED EEE D+ E D +++ND + +++D +E+ + + +G
Sbjct: 395 EDDAPDGGDDDDDEDDEEEGDVQRGGEPD-DDDNDSDDDSDDDEDEDEEDEEEQG----- 556
Query: 287 EEQDIG 292
EE+D+G
Sbjct: 557 EEEDLG 574
Score = 53.9 bits (128), Expect = 4e-07
Identities = 33/126 (26%), Positives = 65/126 (51%), Gaps = 6/126 (4%)
Frame = +2
Query: 178 KVIVESEENPEQENDIVESEEEDPEEENDIV------ESEEDPEQENDIVESEEEDLEKE 231
+V+ ++E++ ++E D + E +D ++E D + +ED E+E D+ E D + +
Sbjct: 314 EVVDDAEDDEDEEEDDDDDEGDDNDDEEDDAPDGGDDDDDEDDEEEGDVQRGGEPD-DDD 490
Query: 232 NDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDI 291
ND + +++D +E+ + E E+ + + + E EEE SD E N +EE++
Sbjct: 491 NDSDDDSDDDEDEDEEDEEEQGEEEDLGTEYLIRPLETAEEEEASSDFEPEENGEEEEED 670
Query: 292 GYDTVC 297
D C
Sbjct: 671 VDDEDC 688
Score = 48.5 bits (114), Expect = 2e-05
Identities = 33/107 (30%), Positives = 56/107 (51%), Gaps = 8/107 (7%)
Frame = +2
Query: 182 ESEENPEQENDIVESEE--EDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAE 239
+ +E+ E+E D+ E +D + +D + +ED ++E++ + EEEDL E ++ E
Sbjct: 425 DDDEDDEEEGDVQRGGEPDDDDNDSDDDSDDDEDEDEEDEEEQGEEEDLGTEY-LIRPLE 601
Query: 240 EDPEEENDMVESAEEDPEEENDMV-----ESAEEDPE-EENDKSDGE 280
EEE EE+ EEE + V E AE P+ + +DK D +
Sbjct: 602 TAEEEEASSDFEPEENGEEEEEDVDDEDCEKAEAPPKRKRSDKDDSD 742
Score = 40.4 bits (93), Expect = 0.004
Identities = 24/99 (24%), Positives = 51/99 (51%), Gaps = 4/99 (4%)
Frame = +2
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQE---NDIVESEEEDLEKENDMVESA 238
E +++ +D + +E++ EE+ + EED E + +EEE+ + + E+
Sbjct: 473 EPDDDDNDSDDDSDDDEDEDEEDEEEQGEEEDLGTEYLIRPLETAEEEEASSDFEPEENG 652
Query: 239 EEDPEEENDM-VESAEEDPEEENDMVESAEEDPEEENDK 276
EE+ E+ +D E AE P+ + + +++D E+D+
Sbjct: 653 EEEEEDVDDEDCEKAEAPPKRKRSDKDDSDDDDGGEDDE 769
>TC218462
Length = 1117
Score = 57.4 bits (137), Expect = 3e-08
Identities = 36/123 (29%), Positives = 69/123 (55%), Gaps = 6/123 (4%)
Frame = +1
Query: 176 KLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQE-----NDIVESEEEDLEK 230
KLK I SEENP ++N + ++ ++ + + +++E+P E ND +E +++
Sbjct: 199 KLKSI--SEENPSEQNGV----DQQAKDNSVVSKTDENPRSEITSGENDQKVTENVGVDE 360
Query: 231 E-NDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQ 289
E N+M E+ E EE+N+M E+ E + E+ DM E+ + E+ N +G+ + +++Q
Sbjct: 361 EKNEMKEAKETVNEEDNEMKEAEETESEDNTDMKEAKGTESEDNNGMKEGKVTKDEEKKQ 540
Query: 290 DIG 292
G
Sbjct: 541 QAG 549
Score = 45.8 bits (107), Expect = 1e-04
Identities = 38/151 (25%), Positives = 70/151 (46%), Gaps = 24/151 (15%)
Frame = +1
Query: 160 KRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVE----SEEDPE 215
K+D D + K++ KK + V S N + +++ EE+ P+ + SEE+P
Sbjct: 67 KKDFDAEREKELEDYKKSRPPVGS--NHQNSSNV---EEDYPKGLIIAFKLKSISEENPS 231
Query: 216 QENDIVESEEEDLEKENDMVESAEEDP--------------------EEENDMVESAEED 255
++N + + K+N +V +E+P EE+N+M E+ E
Sbjct: 232 EQNGV-----DQQAKDNSVVSKTDENPRSEITSGENDQKVTENVGVDEEKNEMKEAKETV 396
Query: 256 PEEENDMVESAEEDPEEENDKSDGEGVLNLD 286
EE+N+M E+ E + E+ D + +G + D
Sbjct: 397 NEEDNEMKEAEETESEDNTDMKEAKGTESED 489
Score = 32.3 bits (72), Expect = 1.2
Identities = 27/117 (23%), Positives = 52/117 (44%)
Frame = +1
Query: 122 EESRASILVKVLKEFSSFDITPSENDVLCHMALISEAVKRDTDLTKSKDVHTTKKLKVIV 181
++++ + +V E +IT END ++E V D + + K+ T V
Sbjct: 250 QQAKDNSVVSKTDENPRSEITSGEND-----QKVTENVGVDEEKNEMKEAKET------V 396
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESA 238
E+N +E + ESE+ +E ESE++ + V +EE ++ + +A
Sbjct: 397 NEEDNEMKEAEETESEDNTDMKEAKGTESEDNNGMKEGKVTKDEEKKQQAGEKFSAA 567
>TC204103 similar to UP|Q75QC2 (Q75QC2) Glutamate-rich protein, partial (38%)
Length = 845
Score = 52.0 bits (123), Expect = 1e-06
Identities = 40/131 (30%), Positives = 66/131 (49%), Gaps = 6/131 (4%)
Frame = +3
Query: 163 TDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEED--PE-QEND 219
T++TK ++ + + +EE E+ + EEE P E N+ E E+ PE +EN
Sbjct: 180 TEVTKIEETTPEQPATQVTTTEEPKEETTKEAKEEEEAPVETNEATEVTEEVKPEVEENP 359
Query: 220 IVE-SEEEDLEKENDMVESA--EEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDK 276
E +EEE++++E ESA EE +EEN E+ EE E+ + E+
Sbjct: 360 APERTEEEEVKEETKETESAPVEEQKQEENKPAETVEETKTEQVSV--------EKTEA* 515
Query: 277 SDGEGVLNLDE 287
+ G G++ +DE
Sbjct: 516 AHGYGIIEVDE 548
Score = 43.9 bits (102), Expect = 4e-04
Identities = 34/96 (35%), Positives = 51/96 (52%), Gaps = 6/96 (6%)
Frame = +3
Query: 189 QENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKE-NDMVESAEE-DPE-EE 245
+ ++ + EE PE+ V + E+P++E EEE+ E N+ E EE PE EE
Sbjct: 174 EATEVTKIEETTPEQPATQVTTTEEPKEETTKEAKEEEEAPVETNEATEVTEEVKPEVEE 353
Query: 246 NDMVE-SAEEDPEEENDMVESA--EEDPEEENDKSD 278
N E + EE+ +EE ESA EE +EEN ++
Sbjct: 354 NPAPERTEEEEVKEETKETESAPVEEQKQEENKPAE 461
Score = 38.9 bits (89), Expect = 0.012
Identities = 32/98 (32%), Positives = 49/98 (49%), Gaps = 4/98 (4%)
Frame = +3
Query: 203 EENDIVESEE-DPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEE-DPE-EE 259
E ++ + EE PEQ V + EE E+ + EE P E N+ E EE PE EE
Sbjct: 174 EATEVTKIEETTPEQPATQVTTTEEPKEETTKEAKEEEEAPVETNEATEVTEEVKPEVEE 353
Query: 260 NDMVE-SAEEDPEEENDKSDGEGVLNLDEEQDIGYDTV 296
N E + EE+ +EE +++ V +E++ +TV
Sbjct: 354 NPAPERTEEEEVKEETKETESAPVEEQKQEENKPAETV 467
>BE822052 similar to GP|4557063|gb|A expressed protein {Arabidopsis
thaliana}, partial (28%)
Length = 668
Score = 50.1 bits (118), Expect = 5e-06
Identities = 35/114 (30%), Positives = 60/114 (51%), Gaps = 1/114 (0%)
Frame = -1
Query: 164 DLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIV-ESEEDPEQENDIVE 222
DL K K HT + K +S+E+ ++E+D +++ D E + D E EE+ + E+D
Sbjct: 569 DLCKGK--HTYRLNKDASDSDEDEDEEDDHDTNDQNDEEGDEDFSGEGEEEADPEDD--- 405
Query: 223 SEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDK 276
E ND +S +ED +++ND + E++ E+E + E EE P+ K
Sbjct: 404 PEANGAGGSNDDDDS-DEDDDDDNDGDDDGEDEDEKEGEDEEDEEEVPQPPTKK 246
Score = 45.4 bits (106), Expect = 1e-04
Identities = 29/99 (29%), Positives = 48/99 (48%)
Frame = -1
Query: 191 NDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVE 250
N +ED +EE+D D +ND E +ED E + E+DPE
Sbjct: 536 NKDASDSDEDEDEEDD-----HDTNDQND--EEGDEDFSGEGEEEADPEDDPEANGAGGS 378
Query: 251 SAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQ 289
+ ++D +E++D ++D E+E++K EG DEE+
Sbjct: 377 NDDDDSDEDDDDDNDGDDDGEDEDEK---EGEDEEDEEE 270
Score = 44.3 bits (103), Expect = 3e-04
Identities = 24/92 (26%), Positives = 49/92 (53%), Gaps = 1/92 (1%)
Frame = -1
Query: 192 DIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVES 251
D +S+E++ EE++ + D E + D EE+ + E+D + +++D
Sbjct: 530 DASDSDEDEDEEDDHDTNDQNDEEGDEDFSGEGEEEADPEDDPEANGAGGSNDDDD---- 363
Query: 252 AEEDPEEENDMVESAEEDPEEE-NDKSDGEGV 282
++ED +++ND + E++ E+E D+ D E V
Sbjct: 362 SDEDDDDDNDGDDDGEDEDEKEGEDEEDEEEV 267
Score = 38.5 bits (88), Expect = 0.016
Identities = 25/83 (30%), Positives = 40/83 (48%), Gaps = 5/83 (6%)
Frame = -1
Query: 217 ENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEE-----ENDMVESAEEDPE 271
++D E EE+D + + E +ED E + E+DPE ND +S E+D +
Sbjct: 521 DSDEDEDEEDDHDTNDQNDEEGDEDFSGEGEEEADPEDDPEANGAGGSNDDDDSDEDDDD 342
Query: 272 EENDKSDGEGVLNLDEEQDIGYD 294
+ + DGE DE++ G D
Sbjct: 341 DNDGDDDGE-----DEDEKEGED 288
>TC214054 similar to UP|Q75QC2 (Q75QC2) Glutamate-rich protein, partial (68%)
Length = 1306
Score = 49.7 bits (117), Expect = 7e-06
Identities = 37/131 (28%), Positives = 61/131 (46%), Gaps = 11/131 (8%)
Frame = +1
Query: 159 VKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEEN-----DIVE---S 210
V + + T + T++ V + E P E ES E P+EE + VE +
Sbjct: 157 VSKTQETTPVTEAPATEQPAAEVPATEQPAAEAPAPESTTEAPKEETTEAPTETVEKTTT 336
Query: 211 EEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDP 270
E PE+ ++ EE + KE + + AEE EE+ + V+ E+P+E + +A P
Sbjct: 337 EVAPEEPKEVPVETEEVVAKETEEEKPAEEKSEEKTEEVKEEAEEPKETTETESAAAAPP 516
Query: 271 ---EEENDKSD 278
EEEN ++
Sbjct: 517 ATTEEENKPAE 549
Score = 29.3 bits (64), Expect = 9.8
Identities = 17/59 (28%), Positives = 30/59 (50%), Gaps = 4/59 (6%)
Frame = +1
Query: 183 SEENPEQENDIVESEEEDPEEENDIVESEEDP----EQENDIVESEEEDLEKENDMVES 237
+EE E++ + V+ E E+P+E + + P E+EN ES E +E + E+
Sbjct: 418 AEEKSEEKTEEVKEEAEEPKETTETESAAAAPPATTEEENKPAESVETPVEVPVEKTEA 594
>TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23),
partial (8%)
Length = 581
Score = 49.3 bits (116), Expect = 9e-06
Identities = 28/108 (25%), Positives = 63/108 (57%)
Frame = +2
Query: 169 KDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDL 228
K HT+++ K ++E++ + ++D+ + E+ D ++E+D S +D +E D + E +
Sbjct: 236 KGKHTSEENKDASDTEDD-DDDDDVNDGEDGDDDDEDDEDFSGDDGGEEADSDDDPEANG 412
Query: 229 EKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDK 276
E+D + ++D +++ND + E+D +EE++ + EE P+ + K
Sbjct: 413 GGESDDDDEDDDDDDDDNDEDDGDEDDDDEEDE--DEDEETPQPPSKK 550
Score = 47.8 bits (112), Expect = 3e-05
Identities = 25/98 (25%), Positives = 53/98 (53%)
Frame = +2
Query: 183 SEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDP 242
SEEN + S+ ED ++++D+ + E+ + + D + +D +E D + E +
Sbjct: 251 SEENKDA------SDTEDDDDDDDVNDGEDGDDDDEDDEDFSGDDGGEEADSDDDPEANG 412
Query: 243 EEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGE 280
E+D + ++D +++ND + E+D +EE++ D E
Sbjct: 413 GGESDDDDEDDDDDDDDNDEDDGDEDDDDEEDEDEDEE 526
Score = 45.8 bits (107), Expect = 1e-04
Identities = 25/93 (26%), Positives = 52/93 (55%)
Frame = +2
Query: 203 EENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDM 262
EEN ED + ++D+ + E+ D + E+D + + +D EE D + E + E+D
Sbjct: 254 EENKDASDTEDDDDDDDVNDGEDGDDDDEDDE-DFSGDDGGEEADSDDDPEANGGGESDD 430
Query: 263 VESAEEDPEEENDKSDGEGVLNLDEEQDIGYDT 295
+ ++D +++ND+ DG+ + +E++D +T
Sbjct: 431 DDEDDDDDDDDNDEDDGDEDDDDEEDEDEDEET 529
>BE805377 similar to GP|21648010|gb| peptidase M23/M37 family {Chlorobium
tepidum TLS}, partial (4%)
Length = 323
Score = 48.5 bits (114), Expect = 2e-05
Identities = 29/92 (31%), Positives = 53/92 (57%), Gaps = 1/92 (1%)
Frame = +1
Query: 175 KKLKVIVE-SEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKEND 233
KK + + E +EEN E E + +++ + +EE + V EE PE++ +++ESE +++ E +
Sbjct: 58 KKDEAVKEVAEENTETEKKVADADTK--KEETEKVSIEESPEEKKEVIESETKEIVAETE 231
Query: 234 MVESAEEDPEEENDMVESAEEDPEEENDMVES 265
+ + E EE VE+ E+ PE E E+
Sbjct: 232 VSKKTNE--TEEIKSVETEEKKPEVEKSESET 321
Score = 38.1 bits (87), Expect = 0.021
Identities = 24/84 (28%), Positives = 44/84 (51%), Gaps = 1/84 (1%)
Frame = +1
Query: 155 ISEAVKRDTDLTKS-KDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEED 213
+ E + +T+ K D T K+ V EE+PE++ +++ESE ++ E ++ + +
Sbjct: 73 VKEVAEENTETEKKVADADTKKEETEKVSIEESPEEKKEVIESETKEIVAETEVSKKTNE 252
Query: 214 PEQENDIVESEEEDLEKENDMVES 237
E E VE+EE+ E E E+
Sbjct: 253 TE-EIKSVETEEKKPEVEKSESET 321
>TC232681 weakly similar to GB|AAP21377.1|30102918|BT006569 At1g47970
{Arabidopsis thaliana;} , partial (39%)
Length = 867
Score = 48.5 bits (114), Expect = 2e-05
Identities = 31/107 (28%), Positives = 54/107 (49%), Gaps = 8/107 (7%)
Frame = +1
Query: 182 ESEENPEQENDIVES--------EEEDPEEENDIVESEEDPEQENDIVESEEEDLEKEND 233
+ ++N ++E+D + ++ED EEE+D+ E + +ND + E+ED E E +
Sbjct: 214 DDDDNEDEEDDDEDDAPDGGDDDDDEDEEEESDVQRGGEPDDDDNDD-DDEDEDEEDEEE 390
Query: 234 MVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGE 280
E A+ E + +AEE+ + E E+ EE+ D DGE
Sbjct: 391 QGEEADLGTEYLIRPLVTAEEEEASSDFEPEENGEEEEEDVDDEDGE 531
Score = 47.4 bits (111), Expect = 3e-05
Identities = 32/114 (28%), Positives = 59/114 (51%), Gaps = 8/114 (7%)
Frame = +1
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKE---NDMVESA 238
+ +E+ E+E+D+ E D +++ND + +ED E E + + EE DL E +V +
Sbjct: 280 DDDEDEEEESDVQRGGEPD-DDDNDDDDEDEDEEDEEE--QGEEADLGTEYLIRPLVTAE 450
Query: 239 EEDPEEENDMVESAEEDPEEENDMVESAEEDP-----EEENDKSDGEGVLNLDE 287
EE+ + + E+ EE+ E+ +D E P +++D D +G + DE
Sbjct: 451 EEEASSDFEPEENGEEEEEDVDDEDGEKSEAPPKRKRSDKDDSDDDDGGEDDDE 612
Score = 45.8 bits (107), Expect = 1e-04
Identities = 28/106 (26%), Positives = 49/106 (45%), Gaps = 12/106 (11%)
Frame = +1
Query: 188 EQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEEND 247
+ +N+ E ++ED + + +ED E+E+D+ E D + +D + +ED E+E +
Sbjct: 217 DDDNEDEEDDDEDDAPDGGDDDDDEDEEEESDVQRGGEPDDDDNDD--DDEDEDEEDEEE 390
Query: 248 MVESAE------------EDPEEENDMVESAEEDPEEENDKSDGEG 281
E A+ + EE + E E EEE D D +G
Sbjct: 391 QGEEADLGTEYLIRPLVTAEEEEASSDFEPEENGEEEEEDVDDEDG 528
Score = 44.3 bits (103), Expect = 3e-04
Identities = 30/104 (28%), Positives = 53/104 (50%), Gaps = 10/104 (9%)
Frame = +1
Query: 197 EEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDP 256
+++D E+E D + E+D D + ++ED E+E+D+ E D ++ +D + +ED
Sbjct: 214 DDDDNEDEED--DDEDDAPDGGD--DDDDEDEEEESDVQRGGEPDDDDNDD--DDEDEDE 375
Query: 257 EEENDMVESAE----------EDPEEENDKSDGEGVLNLDEEQD 290
E+E + E A+ EEE SD E N +EE++
Sbjct: 376 EDEEEQGEEADLGTEYLIRPLVTAEEEEASSDFEPEENGEEEEE 507
Score = 36.2 bits (82), Expect = 0.080
Identities = 19/66 (28%), Positives = 39/66 (58%), Gaps = 2/66 (3%)
Frame = +1
Query: 215 EQENDIVESEEEDLEKEND-MVESAEEDPEEENDMVESAE-EDPEEENDMVESAEEDPEE 272
+ ++D + E++D + D + +ED EEE+D+ E +D + ++D + EED EE
Sbjct: 211 DDDDDNEDEEDDDEDDAPDGGDDDDDEDEEEESDVQRGGEPDDDDNDDDDEDEDEEDEEE 390
Query: 273 ENDKSD 278
+ +++D
Sbjct: 391 QGEEAD 408
>BU926431
Length = 457
Score = 48.1 bits (113), Expect = 2e-05
Identities = 41/130 (31%), Positives = 65/130 (49%), Gaps = 8/130 (6%)
Frame = +3
Query: 169 KDVHTTKKLKVIVESEENPEQENDIVES----EEEDPEEENDIVESEE--DPEQENDIVE 222
KD+H K +V + E+E+D VE + + EEE + ESEE D
Sbjct: 21 KDLHPGKD-EVNQNEKHEEEEEDDNVEDAYKHDHNEHEEEGNEHESEEKGDKRGVRGGGS 197
Query: 223 SEEEDLEKENDMVE--SAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGE 280
+EED K N++ + +D +END +SA + +E E +E+ EEE+D+ + E
Sbjct: 198 EQEEDENKSNEVEDQRGGGDDEIDENDQEQSAVDTDHDE----EFTDEEKEEESDEKEKE 365
Query: 281 GVLNLDEEQD 290
+ DEE+D
Sbjct: 366 N--SRDEEKD 389
Score = 37.7 bits (86), Expect = 0.028
Identities = 24/82 (29%), Positives = 37/82 (44%), Gaps = 2/82 (2%)
Frame = +3
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIV--ESEEEDLEKENDMVESAE 239
E +EN E + +D +END +S D + + + E EEE EKE + E
Sbjct: 204 EEDENKSNEVEDQRGGGDDEIDENDQEQSAVDTDHDEEFTDEEKEEESDEKEKENSRDEE 383
Query: 240 EDPEEENDMVESAEEDPEEEND 261
+D EN A E+ + +D
Sbjct: 384 KDGSVENHNSHEAREENYKGDD 449
>TC229446 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(20%)
Length = 850
Score = 47.4 bits (111), Expect = 3e-05
Identities = 43/139 (30%), Positives = 62/139 (43%), Gaps = 4/139 (2%)
Frame = +3
Query: 153 ALISEAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPE----EENDIV 208
A+ + R +D +K + ++ + VE ++ +Q N E EDP +E
Sbjct: 81 AMAKKRKLRSSDPEPTKPIEPQQQDQEPVELDQQQQQPNP--EQTLEDPNTMAVDEPKQE 254
Query: 209 ESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEE 268
E EED E +N ESEEE+ E E E E EE+ D A + E N E E
Sbjct: 255 EEEEDEEPQN--AESEEEEEEAEEQQEERPPETLEEQQDPETLAAANHAEVNGGNEGNNE 428
Query: 269 DPEEENDKSDGEGVLNLDE 287
+ EEE+ + E V L E
Sbjct: 429 EEEEEDLTLEEEPVEKLLE 485
Score = 44.3 bits (103), Expect = 3e-04
Identities = 39/145 (26%), Positives = 61/145 (41%), Gaps = 16/145 (11%)
Frame = +3
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPE-------------QENDIVESEEEDL 228
E +N E E + E+EE+ E + +E ++DPE E + E EEEDL
Sbjct: 270 EEPQNAESEEEEEEAEEQQEERPPETLEEQQDPETLAAANHAEVNGGNEGNNEEEEEEDL 449
Query: 229 EKENDMVESAEEDPEEE---NDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNL 285
E + VE E +E + + ++ + PE + + A+ DP G G
Sbjct: 450 TLEEEPVEKLLEPFTKEQLHSLVTQAVDMFPEFVDSVRRLADVDPAHRKIFVHGLG---- 617
Query: 286 DEEQDIGYDTVCAICDNGGEILPCE 310
D DT+ A+ GEI C+
Sbjct: 618 ---WDATADTLTAVFGKYGEIEDCK 683
Score = 43.1 bits (100), Expect = 7e-04
Identities = 36/127 (28%), Positives = 61/127 (47%), Gaps = 2/127 (1%)
Frame = +3
Query: 167 KSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEE--ENDIVESEEDPEQENDIVESE 224
+S D TK ++ + +E E + + ++ +PE+ E+ + ++P+QE E
Sbjct: 105 RSSDPEPTKPIEPQQQDQEPVELDQ---QQQQPNPEQTLEDPNTMAVDEPKQEE-----E 260
Query: 225 EEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLN 284
EED E +N ES EE+ E E E E EE+ D A + E N ++G
Sbjct: 261 EEDEEPQN--AESEEEEEEAEEQQEERPPETLEEQQDPETLAAANHAEVNGGNEGNN--E 428
Query: 285 LDEEQDI 291
+EE+D+
Sbjct: 429 EEEEEDL 449
>TC204094 weakly similar to UP|Q75QC2 (Q75QC2) Glutamate-rich protein,
partial (43%)
Length = 616
Score = 47.4 bits (111), Expect = 3e-05
Identities = 36/113 (31%), Positives = 56/113 (48%), Gaps = 4/113 (3%)
Frame = +1
Query: 159 VKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQEN 218
V + T T +++ TT+ K+ + E P E E+ E+ +EE + +E E
Sbjct: 181 VAQQTPTTVTENETTTEVTKIEETTPEQPATEVTAPETVNEEAKEEAAVETNEATEVTEE 360
Query: 219 DIVES--EEEDLEKENDMVESA--EEDPEEENDMVESAEEDPEEENDMVESAE 267
E EEE++++E ESA EE+ +EEN E+ EE P+ E VE E
Sbjct: 361 GKAEKTEEEEEVKEETKETESAPVEEEKQEENKPAETVEE-PKTEQVSVEKTE 516
>TC204102 similar to UP|Q75QC2 (Q75QC2) Glutamate-rich protein, partial (23%)
Length = 777
Score = 46.2 bits (108), Expect = 8e-05
Identities = 38/120 (31%), Positives = 60/120 (49%), Gaps = 4/120 (3%)
Frame = +1
Query: 152 MALISEAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESE 211
MA + A + T +T+++ TT+ K+ + E P E E+ E+ +EE + +E
Sbjct: 76 MANVEVAQQTXTCVTENET--TTEVTKIEETTSEQPATEVGAPETVNEEAKEEAAVETNE 249
Query: 212 EDPEQENDIVES--EEEDLEKENDMVESA--EEDPEEENDMVESAEEDPEEENDMVESAE 267
E E EEE++++E ESA EE+ +EEN E+ EE P+ E VE E
Sbjct: 250 ATEVTEEGKAEKTEEEEEVKEETKETESAPVEEEKQEENKPAETVEE-PKTEQVSVEKTE 426
>TC231046
Length = 959
Score = 46.2 bits (108), Expect = 8e-05
Identities = 46/143 (32%), Positives = 72/143 (50%), Gaps = 14/143 (9%)
Frame = +1
Query: 168 SKDVHTTKKLK--VIVESEE--------NPEQENDIVESEEEDPEEENDIVESEEDPEQE 217
S++ H + +K VIVE+E+ P +E IVESE E+P +E + +P +E
Sbjct: 64 SEETHINEPVKXXVIVEAEKPSEETHINEPAKEIMIVESEAENPSKETLV----NEPAKE 231
Query: 218 NDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVE--SAEEDPEE--E 273
+ES+ E +EN+ +S+E EE VE A E P EE +E +E+P E E
Sbjct: 232 TVKLESDAEKPSEENN--QSSESLKEE---FVE-AMEAPSEETQSIEPIKLKEEPIEPSE 393
Query: 274 NDKSDGEGVLNLDEEQDIGYDTV 296
K GE N E + + +++
Sbjct: 394 EGKPSGESEKNASETKTLIVESI 462
Score = 43.9 bits (102), Expect = 4e-04
Identities = 35/107 (32%), Positives = 54/107 (49%)
Frame = +1
Query: 171 VHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEK 230
++ K VIVESE E E+ +P + IVE+E+ P +E I E +E +
Sbjct: 10 INXXXKXXVIVESEAKKPSE----ETHINEPVKXXVIVEAEK-PSEETHINEPAKEIM-- 168
Query: 231 ENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKS 277
+VES E+P + E+ +P +E +ES E P EEN++S
Sbjct: 169 ---IVESEAENPSK-----ETLVNEPAKETVKLESDAEKPSEENNQS 285
>TC207527 weakly similar to UP|O08661 (O08661) Ryanodine receptor type II
(Fragment), partial (4%)
Length = 981
Score = 45.4 bits (106), Expect = 1e-04
Identities = 36/143 (25%), Positives = 66/143 (45%), Gaps = 12/143 (8%)
Frame = +1
Query: 160 KRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDP---EEENDIVESEEDPEQ 216
KR D + KD KK V + ++P +++ + +P ++E +VE+E +
Sbjct: 478 KRQRDDEQEKD--DVKKSPVKRQKGDDPSVKSEPTNMDTSNPTQVDDEKAVVENENSSNK 651
Query: 217 ENDIVESEEEDLEKENDMVESAEEDPEEENDMVE-SAEEDPEEENDMVESAEEDPEEEN- 274
E+D+ +E +D E EEDPEE +M S + + +N+ + + D + EN
Sbjct: 652 EDDV------KMEDGSDEEEDPEEDPEEYEEMENGSPQHEASHDNNAEQEVKADTKSENI 813
Query: 275 -------DKSDGEGVLNLDEEQD 290
D++ E + DE Q+
Sbjct: 814 TTNNKTTDETSKEEIKVKDEVQE 882
Score = 42.7 bits (99), Expect = 9e-04
Identities = 30/110 (27%), Positives = 53/110 (47%), Gaps = 6/110 (5%)
Frame = +1
Query: 178 KVIVESEENPEQENDIV----ESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKEND 233
K +VE+E + +E+D+ EEEDPEE + EE E EN S + + +N+
Sbjct: 616 KAVVENENSSNKEDDVKMEDGSDEEEDPEE-----DPEEYEEMENG---SPQHEASHDNN 771
Query: 234 MVESAEEDPEEENDMV--ESAEEDPEEENDMVESAEEDPEEENDKSDGEG 281
+ + D + EN ++ +E +EE + + +E + K + EG
Sbjct: 772 AEQEVKADTKSENITTNNKTTDETSKEEIKVKDEVQESKADAQVKEEKEG 921
Score = 38.5 bits (88), Expect = 0.016
Identities = 30/130 (23%), Positives = 58/130 (44%), Gaps = 10/130 (7%)
Frame = +1
Query: 176 KLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDI----------VESEE 225
++K +++ + Q +D E E++D ++ + +DP +++ V+ E+
Sbjct: 445 RIKFVIKRNQKKRQRDD--EQEKDDVKKSPVKRQKGDDPSVKSEPTNMDTSNPTQVDDEK 618
Query: 226 EDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNL 285
+E EN + + E+ +D E EEDPEE +M EN E +
Sbjct: 619 AVVENENSSNKEDDVKMEDGSDEEEDPEEDPEEYEEM----------ENGSPQHEASHDN 768
Query: 286 DEEQDIGYDT 295
+ EQ++ DT
Sbjct: 769 NAEQEVKADT 798
>TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%)
Length = 857
Score = 45.1 bits (105), Expect = 2e-04
Identities = 25/86 (29%), Positives = 45/86 (52%), Gaps = 1/86 (1%)
Frame = +1
Query: 188 EQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEEND 247
+ +N E EE+ E++ D + +ED + E E +E + E++ +D +E++D
Sbjct: 349 QNKNSETEDEEDGDEDDEDGPDEDEDGDDEEFSGEEGDEGGDPEDEANGEGSDDGDEDDD 528
Query: 248 MVESA-EEDPEEENDMVESAEEDPEE 272
++ EED ++ D E EED EE
Sbjct: 529 GDDNGDEEDDDDGEDEEEDEEEDEEE 606
Score = 42.0 bits (97), Expect = 0.001
Identities = 26/90 (28%), Positives = 48/90 (52%), Gaps = 2/90 (2%)
Frame = +1
Query: 202 EEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEE--DPEEENDMVESAEEDPEEE 259
+ +N E EED +++++ E+ED + E E +E DPE+E + S + D +++
Sbjct: 349 QNKNSETEDEEDGDEDDEDGPDEDEDGDDEEFSGEEGDEGGDPEDEANGEGSDDGDEDDD 528
Query: 260 NDMVESAEEDPEEENDKSDGEGVLNLDEEQ 289
D E+D + E+++ D E DEE+
Sbjct: 529 GDDNGDEEDDDDGEDEEEDEEE----DEEE 606
Score = 40.8 bits (94), Expect = 0.003
Identities = 29/97 (29%), Positives = 45/97 (45%), Gaps = 2/97 (2%)
Frame = +1
Query: 164 DLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEE--DPEQENDIV 221
DL K + K + E+ E + D + +E+ +EE E +E DPE E +
Sbjct: 319 DLFGRKRAYLQNKNSETEDEEDGDEDDEDGPDEDEDGDDEEFSGEEGDEGGDPEDEANGE 498
Query: 222 ESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEE 258
S++ D + + D EED ++ D E EED EE
Sbjct: 499 GSDDGDEDDDGDD-NGDEEDDDDGEDEEEDEEEDEEE 606
Score = 34.3 bits (77), Expect = 0.31
Identities = 32/139 (23%), Positives = 61/139 (43%)
Frame = +1
Query: 152 MALISEAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESE 211
M + +AV DT L ++ + L S +P ++ E + P E D+ +
Sbjct: 175 MPSLLQAVVADTVLAAAQSIALIGYLLTNGTSSVSP----NLTEVDGRFPLE--DLFGRK 336
Query: 212 EDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPE 271
Q + +EED +++++ +ED ++E E +E + E++ +D +
Sbjct: 337 RAYLQNKNSETEDEEDGDEDDEDGPDEDEDGDDEEFSGEEGDEGGDPEDEANGEGSDDGD 516
Query: 272 EENDKSDGEGVLNLDEEQD 290
E++D D N DEE D
Sbjct: 517 EDDDGDD-----NGDEEDD 558
Score = 30.8 bits (68), Expect = 3.4
Identities = 22/85 (25%), Positives = 42/85 (48%), Gaps = 6/85 (7%)
Frame = +1
Query: 212 EDPEQENDIVESEEEDLE------KENDMVESAEEDPEEENDMVESAEEDPEEENDMVES 265
++ E + E +ED E ++ D E + E+ +E D + A + ++ D +
Sbjct: 349 QNKNSETEDEEDGDEDDEDGPDEDEDGDDEEFSGEEGDEGGDPEDEANGEGSDDGDEDDD 528
Query: 266 AEEDPEEENDKSDGEGVLNLDEEQD 290
+++ +EE+D DGE DEE+D
Sbjct: 529 GDDNGDEEDD-DDGE-----DEEED 585
>TC210126 similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23), partial
(7%)
Length = 735
Score = 44.7 bits (104), Expect = 2e-04
Identities = 28/87 (32%), Positives = 47/87 (53%), Gaps = 2/87 (2%)
Frame = +3
Query: 196 SEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEED 255
SE ED +++ D E +ED ++D E EE E+ ++ E DPE++ + +D
Sbjct: 330 SETEDDDKDGD--EDDEDGHDQDDDGEDEEFSGEEGDE-----EGDPEDDPEANGGGSDD 488
Query: 256 PEEENDMVESAE--EDPEEENDKSDGE 280
+++ND E + ED EEE D+ + E
Sbjct: 489 DDDDNDNEEDDDDGEDEEEEEDEEEDE 569
Score = 43.9 bits (102), Expect = 4e-04
Identities = 29/94 (30%), Positives = 50/94 (52%), Gaps = 2/94 (2%)
Frame = +3
Query: 185 ENPEQENDIVESEE--EDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDP 242
+N E E+D + +E ED +++D E EE +E D EE D E + + +D
Sbjct: 324 KNSETEDDDKDGDEDDEDGHDQDDDGEDEEFSGEEGD----EEGDPEDDPEANGGGSDDD 491
Query: 243 EEENDMVESAEEDPEEENDMVESAEEDPEEENDK 276
+++ND + E+D + E++ E EE+ EEE +
Sbjct: 492 DDDND---NEEDDDDGEDEEEEEDEEEDEEETSQ 584
Score = 41.6 bits (96), Expect = 0.002
Identities = 29/94 (30%), Positives = 48/94 (50%)
Frame = +3
Query: 197 EEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDP 256
E ++ E E+D + +ED E +D +++D E E E + E+ +EE D E+DP
Sbjct: 318 ENKNSETEDDDKDGDEDDEDGHD----QDDDGEDE----EFSGEEGDEEGD----PEDDP 461
Query: 257 EEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
E + ++D + E D DGE DEE++
Sbjct: 462 EANGGGSDDDDDDNDNEEDDDDGE-----DEEEE 548
Score = 37.0 bits (84), Expect = 0.047
Identities = 23/98 (23%), Positives = 48/98 (48%), Gaps = 3/98 (3%)
Frame = +3
Query: 164 DLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEE---NDIVESEEDPEQENDI 220
DL K + K + +++ +++++ +++D E+E + + E DPE + +
Sbjct: 288 DLFGRKHAYLENKNSETEDDDKDGDEDDEDGHDQDDDGEDEEFSGEEGDEEGDPEDDPEA 467
Query: 221 VESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEE 258
+D + +ND E ++ +EE + E EED EE
Sbjct: 468 NGGGSDDDDDDNDNEEDDDDGEDEEEE--EDEEEDEEE 575
Score = 36.2 bits (82), Expect = 0.080
Identities = 17/78 (21%), Positives = 42/78 (53%), Gaps = 2/78 (2%)
Frame = +3
Query: 215 EQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEE--DPEEENDMVESAEEDPEE 272
E +N E +++D +++++ ++D E+E E +E DPE++ + +D ++
Sbjct: 318 ENKNSETEDDDKDGDEDDEDGHDQDDDGEDEEFSGEEGDEEGDPEDDPEANGGGSDDDDD 497
Query: 273 ENDKSDGEGVLNLDEEQD 290
+ND + + +EE++
Sbjct: 498 DNDNEEDDDDGEDEEEEE 551
>BE475116
Length = 367
Score = 44.7 bits (104), Expect = 2e-04
Identities = 33/89 (37%), Positives = 48/89 (53%), Gaps = 4/89 (4%)
Frame = +1
Query: 189 QENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDM 248
+E I+ESE + P EE IVE P +E IVESE +E + E A+E E++
Sbjct: 7 EETLIIESETKKPSEETLIVE----PAKETVIVESEARKPSEETLINEPAKETVIVESEA 174
Query: 249 VESAEE----DPEEENDMVESAEEDPEEE 273
+ +EE +P +E +VES + P EE
Sbjct: 175 RKPSEETLINEPAKETVIVESEAQKPLEE 261
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,056,539
Number of Sequences: 63676
Number of extensions: 881896
Number of successful extensions: 6454
Number of sequences better than 10.0: 240
Number of HSP's better than 10.0 without gapping: 4611
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5560
length of query: 1084
length of database: 12,639,632
effective HSP length: 107
effective length of query: 977
effective length of database: 5,826,300
effective search space: 5692295100
effective search space used: 5692295100
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC124966.7


