

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124966.11 - phase: 0
(447 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
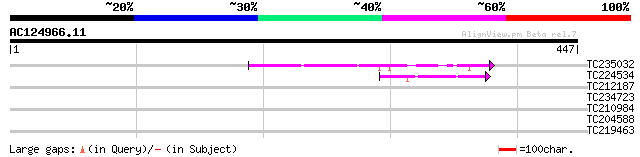
Score E
Sequences producing significant alignments: (bits) Value
TC235032 weakly similar to UP|Q7XDX0 (Q7XDX0) Contains similarit... 82 5e-16
TC224534 similar to UP|Q6USA0 (Q6USA0) Transposase (Fragment), p... 42 7e-04
TC212187 weakly similar to UP|Q6USA0 (Q6USA0) Transposase (Fragm... 40 0.003
TC234723 similar to UP|Q9FHY5 (Q9FHY5) Emb|CAB72466.1 (At5g41980... 38 0.008
TC210984 33 0.26
TC204588 similar to UP|Q7X9G7 (Q7X9G7) Alpha-L-arabinofuranosida... 29 3.8
TC219463 homologue to UP|Q94KL5 (Q94KL5) BEL1-like homeobox 4, p... 28 8.4
>TC235032 weakly similar to UP|Q7XDX0 (Q7XDX0) Contains similarity to
ribosomal protein, partial (24%)
Length = 689
Score = 82.0 bits (201), Expect = 5e-16
Identities = 67/208 (32%), Positives = 99/208 (47%), Gaps = 14/208 (6%)
Frame = +3
Query: 189 LQRILHVSEMRGFPGMIGSIDCMHWEWKNCPKAWEGQFTRGDKGTTTVILEAVASHDLWI 248
+Q ++ + E+RGF M+GSID +HW+ K C AW+GQ+ GD G T++L+ V S L I
Sbjct: 12 IQYLMQICEIRGFQSMLGSIDYLHWK*KTCLVAWKGQYCHGD-GPPTIMLKVVVSQYLRI 188
Query: 249 WHAFFGCPGTLNDINVLDRSPVFDDVEQGKAPRVNFFVNQRP-----YNMAYYLA--DGI 301
HA+ G G+ N+INVL+ S V G++P P + +YL+ D
Sbjct: 189 *HAYLGVEGSNNNINVLNESCV*RRY-GGRSPYCTICS**NPI*HGVLSDIWYLSRLDDA 365
Query: 302 YPSYPTFVKSIRLPQSEPDKLFAKHQESCRKDIERAFGVLQARFKIIREPARLWDIADLG 361
+P ++S + K+ S +K + F ++R W L
Sbjct: 366 CEDHPKAIRS*K-------KIVYSISRSNKKGSHLKY------FNLVRN--YTWSCVWLE 500
Query: 362 -------IIMRSCIILHNMIVEDERDTY 382
M CIILHNMIV+DE TY
Sbjct: 501 HEHISSIYYMYVCIILHNMIVKDEPHTY 584
>TC224534 similar to UP|Q6USA0 (Q6USA0) Transposase (Fragment), partial (50%)
Length = 968
Score = 41.6 bits (96), Expect = 7e-04
Identities = 30/100 (30%), Positives = 49/100 (49%), Gaps = 12/100 (12%)
Frame = +1
Query: 292 NMAYYLADGIYPSYPTFVKSI------------RLPQSEPDKLFAKHQESCRKDIERAFG 339
N Y+L D Y + P F+ PQ+ + LF S + IER+FG
Sbjct: 472 NGKYFLVDAGYTNGPGFLTPY*RTRYHRNEWIGNTPQNYKE-LFNLRHASAKNVIERSFG 648
Query: 340 VLQARFKIIREPARLWDIADLGIIMRSCIILHNMIVEDER 379
VL+ R+ I+R P+ +DI I+ +C +L N I ++++
Sbjct: 649 VLKKRWSILRTPS-FFDIKTQIRIINTCFMLRNFIRDEQQ 765
>TC212187 weakly similar to UP|Q6USA0 (Q6USA0) Transposase (Fragment),
partial (45%)
Length = 973
Score = 39.7 bits (91), Expect = 0.003
Identities = 36/128 (28%), Positives = 59/128 (45%), Gaps = 23/128 (17%)
Frame = +1
Query: 295 YYLADGIYPSYPTFVKSIR-----LPQSEPD-------KLFAKHQESCRKDIERAFGVLQ 342
+YL D YP++ F+ S LPQ ++F + S + IERAFG+ +
Sbjct: 514 FYLVDSCYPTFMGFLGSYNKTRYHLPQFRIGPRIRGRVEVFNYYHSSLQSTIERAFGLCK 693
Query: 343 ARFKIIREPARLWDIADLGIIMRSCIILHNMIVE-----------DERDTYAQRWTDFEQ 391
AR+KI+ + + + I+ +C+ +HN I DE + Y TD ++
Sbjct: 694 ARWKILGNMSP-FALKTQNQIIVACMTIHNFIQRNDKSDGEFDSLDEDNEYID--TDKDE 864
Query: 392 SEGSGSST 399
SE S+T
Sbjct: 865 SEVGPSTT 888
>TC234723 similar to UP|Q9FHY5 (Q9FHY5) Emb|CAB72466.1 (At5g41980), partial
(24%)
Length = 837
Score = 38.1 bits (87), Expect = 0.008
Identities = 39/171 (22%), Positives = 67/171 (38%), Gaps = 23/171 (13%)
Frame = +2
Query: 295 YYLADGIYPSYPTFVK-------------SIRLPQSEPDKLFAKHQESCRKDIERAFGVL 341
YY+ D Y + P FV S PQ + +LF + R I+R FG+L
Sbjct: 143 YYIVDSKYQNVPGFVAPYSSTPYYSKEFLSDYHPQ-DASELFNQRHSLLRHVIDRTFGIL 319
Query: 342 QARFKIIREPARLWDIADLGIIMRSCIILHNMIVEDERDTYAQRWTDFEQSEGSGSSTAQ 401
+ARF I+ + +++ +C LHN I ++ D + + +E+ +
Sbjct: 320 KARFPILMSAPSYPLQTQVKLVVAAC-ALHNYIRREKPDDWLFKM--YEEGSFPMEESQP 490
Query: 402 PYSTEVLPTFANHARARSELRDPNVH----------HELQADLVKHIWTKF 442
P EV P + + P+VH +L+ + +W F
Sbjct: 491 PLEMEVPPKMDVETQTQ-----PSVHTFDSEEIALASQLRDSIATEMWNDF 628
>TC210984
Length = 1085
Score = 33.1 bits (74), Expect = 0.26
Identities = 25/83 (30%), Positives = 39/83 (46%), Gaps = 1/83 (1%)
Frame = +3
Query: 321 KLFAKHQESCRKDIERAFGVLQARFKIIREPARLWDIADLGIIMRSCIILHNMIVEDER- 379
+LF S R IER FG+ ++RF I + +++ +C LHN + ++ R
Sbjct: 426 ELFNLRHASLRNVIERIFGIFKSRFTIFKSAPPFLFKTQAELVL-ACAALHNFLRKECRS 602
Query: 380 DTYAQRWTDFEQSEGSGSSTAQP 402
D + TD E S SS+ P
Sbjct: 603 DEFPVEPTD----ESSSSSSVLP 659
>TC204588 similar to UP|Q7X9G7 (Q7X9G7) Alpha-L-arabinofuranosidase, partial
(84%)
Length = 2003
Score = 29.3 bits (64), Expect = 3.8
Identities = 22/79 (27%), Positives = 32/79 (39%)
Frame = -3
Query: 232 GTTTVILEAVASHDLWIWHAFFGCPGTLNDINVLDRSPVFDDVEQGKAPRVNFFVNQRPY 291
GT T + LWI FFGC + I D S VF D + N F +
Sbjct: 1752 GTVTFMSFPTF*SRLWIGTTFFGCEKEFSSITFADVSTVFVDPD-----NCNEFGSSPSR 1588
Query: 292 NMAYYLADGIYPSYPTFVK 310
+ + A + P + TF++
Sbjct: 1587 EILRFTA--VVPKFTTFIR 1537
>TC219463 homologue to UP|Q94KL5 (Q94KL5) BEL1-like homeobox 4, partial (21%)
Length = 1111
Score = 28.1 bits (61), Expect = 8.4
Identities = 17/54 (31%), Positives = 26/54 (47%)
Frame = +3
Query: 329 SCRKDIERAFGVLQARFKIIREPARLWDIADLGIIMRSCIILHNMIVEDERDTY 382
+CR+ + G L+A FK+ + P ILH+MI++D DTY
Sbjct: 699 TCRRKLRSPLGTLEA-FKLNQFP-----------------ILHHMIIKDNNDTY 806
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.138 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,983,622
Number of Sequences: 63676
Number of extensions: 269072
Number of successful extensions: 1283
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1276
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1281
length of query: 447
length of database: 12,639,632
effective HSP length: 100
effective length of query: 347
effective length of database: 6,272,032
effective search space: 2176395104
effective search space used: 2176395104
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC124966.11


