

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
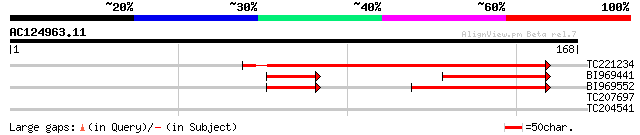
Query= AC124963.11 - phase: 0 /pseudo
(168 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC221234 similar to UP|Q6V9S8 (Q6V9S8) Glycuronosyltransferase-l... 91 3e-19
BI969441 42 1e-06
BI969552 43 6e-06
TC207697 similar to UP|O81698 (O81698) Proline-rich protein, par... 28 2.7
TC204541 homologue to UP|Q93YH7 (Q93YH7) Selenium binding protei... 27 6.0
>TC221234 similar to UP|Q6V9S8 (Q6V9S8) Glycuronosyltransferase-like protein,
partial (45%)
Length = 738
Score = 90.9 bits (224), Expect = 3e-19
Identities = 56/91 (61%), Positives = 61/91 (66%)
Frame = +1
Query: 70 HCFGLLSKRKPNQLNCQRY*GKLVSCIDMLFSVKSSWT*KLN*IIKEI*LLDTLNIIDLV 129
H FG K KP NCQRY*GKL SCI M FS K S *KLN I + I L TL+ +
Sbjct: 49 HAFG---KLKPIPKNCQRY*GKLASCIGMWFSRKISRN*KLNSITRGILRLSTLSTTGSM 219
Query: 130 GLFTLLAFLMFMIFNSSNSLEILRSLGHGQR 160
L+TLLAF MFMI NSS SLEILRSL HGQ+
Sbjct: 220 ALYTLLAFPMFMISNSSISLEILRSLEHGQQ 312
>BI969441
Length = 504
Score = 41.6 bits (96), Expect(2) = 1e-06
Identities = 21/32 (65%), Positives = 26/32 (80%)
Frame = +3
Query: 129 VGLFTLLAFLMFMIFNSSNSLEILRSLGHGQR 160
+ L+T+LAF MFMI NSS SL+ILRSL H Q+
Sbjct: 165 MALYTVLAFPMFMISNSSISLQILRSLEHXQQ 260
Score = 26.6 bits (57), Expect(2) = 1e-06
Identities = 12/16 (75%), Positives = 12/16 (75%)
Frame = +1
Query: 77 KRKPNQLNCQRY*GKL 92
K KP NCQRY*GKL
Sbjct: 10 KLKPLPKNCQRY*GKL 57
>BI969552
Length = 522
Score = 42.7 bits (99), Expect(2) = 6e-06
Identities = 23/41 (56%), Positives = 29/41 (70%)
Frame = +1
Query: 120 LDTLNIIDLVGLFTLLAFLMFMIFNSSNSLEILRSLGHGQR 160
L TL+ + L+T+LAF M MI NSS SL+ILRSL H Q+
Sbjct: 175 LSTLSTTGSMALYTVLAFPMIMISNSSISLQILRSLEHXQQ 297
Score = 23.1 bits (48), Expect(2) = 6e-06
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = +3
Query: 77 KRKPNQLNCQRY*GKL 92
K P NC+RY*GKL
Sbjct: 42 KLTPIPKNCERY*GKL 89
>TC207697 similar to UP|O81698 (O81698) Proline-rich protein, partial (91%)
Length = 1487
Score = 27.7 bits (60), Expect = 2.7
Identities = 18/50 (36%), Positives = 23/50 (46%), Gaps = 7/50 (14%)
Frame = +2
Query: 44 HQQAQNYLITMFF*EGWQIL*SLLINHC----FGL---LSKRKPNQLNCQ 86
H Q YL +F WQ+L NHC F L L + +QL+CQ
Sbjct: 611 HHNFQTYLCLLFIKLKWQLLPVSYSNHCRHRGFHLCNHLCHHRSDQLHCQ 760
>TC204541 homologue to UP|Q93YH7 (Q93YH7) Selenium binding protein (Fragment),
complete
Length = 1732
Score = 26.6 bits (57), Expect = 6.0
Identities = 17/45 (37%), Positives = 21/45 (45%), Gaps = 10/45 (22%)
Frame = +1
Query: 72 FGLLSKRKPNQLN----------CQRY*GKLVSCIDMLFSVKSSW 106
+GLL +RKP N C R+ GK V C SV+S W
Sbjct: 1144 WGLLPERKPYSSNNRRW*YLAI*CSRHPGK*VECWPSDDSVESGW 1278
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.362 0.162 0.552
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,283,664
Number of Sequences: 63676
Number of extensions: 98402
Number of successful extensions: 1333
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1043
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1214
length of query: 168
length of database: 12,639,632
effective HSP length: 90
effective length of query: 78
effective length of database: 6,908,792
effective search space: 538885776
effective search space used: 538885776
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC124963.11


