

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124963.1 + phase: 1 /pseudo
(826 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
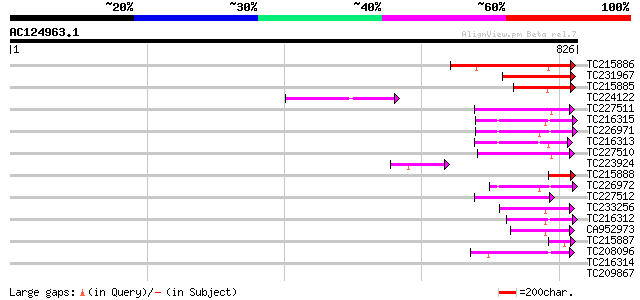
Score E
Sequences producing significant alignments: (bits) Value
TC215886 similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein li... 206 4e-53
TC231967 similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein li... 180 2e-45
TC215885 homologue to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ... 122 5e-28
TC224122 weakly similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-pro... 96 5e-20
TC227511 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {A... 94 2e-19
TC216315 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-pro... 87 4e-17
TC226971 homologue to GB|AAF36454.1|7108521|AF127564 ubiquitin-p... 85 1e-16
TC216313 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-pro... 85 1e-16
TC227510 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {A... 84 3e-16
TC223924 weakly similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-pro... 72 8e-13
TC215888 homologue to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ... 70 3e-12
TC226972 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-pro... 65 2e-10
TC227512 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {A... 60 3e-09
TC233256 weakly similar to UP|HUL5_YEAST (P53119) Probable ubiqu... 59 1e-08
TC216312 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-pro... 58 1e-08
CA952973 54 3e-07
TC215887 similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein li... 53 6e-07
TC208096 weakly similar to UP|Q9SU29 (Q9SU29) Polyubiquitin-like... 45 2e-04
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 38 0.021
TC209867 similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein li... 37 0.046
>TC215886 similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ligase 3
(KAKTUS protein), partial (13%)
Length = 1112
Score = 206 bits (523), Expect = 4e-53
Identities = 113/216 (52%), Positives = 137/216 (63%), Gaps = 33/216 (15%)
Frame = +2
Query: 642 FRNSKIEDLCLDFSLPGYDQTFLLPFS-IY*ITNLYSF-------------------FLV 681
FR + IEDLCLDF+LPGY + L P I I NL + F
Sbjct: 116 FRGAPIEDLCLDFTLPGYPEYILKPGDEIVDINNLEEYISMVVEATVKTGIMRQMEAFRA 295
Query: 682 GFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQ 741
GFNQVF I L++FS +EL+ +LCG W L H KFDHGYTA+SP I+NLL I+
Sbjct: 296 GFNQVFDISSLQIFSPQELDYLLCGRRELWKTETLADHIKFDHGYTAKSPAIVNLLGIMG 475
Query: 742 EFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNH-------------TDTDL 788
EF E++RAF QFVT +PRLPPGGLA L+PKLT+V+K+S + D DL
Sbjct: 476 EFTPEQQRAFCQFVTGAPRLPPGGLAVLNPKLTIVRKLSSSAANASSNGNGPSELADDDL 655
Query: 789 PSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
PSVMTCANYLKLPPYS+KE M +KLLYAI+EG+G F
Sbjct: 656 PSVMTCANYLKLPPYSTKEIMYKKLLYAISEGQGSF 763
>TC231967 similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ligase 3
(KAKTUS protein), partial (6%)
Length = 592
Score = 180 bits (457), Expect = 2e-45
Identities = 85/106 (80%), Positives = 97/106 (91%)
Frame = +1
Query: 719 HFKFDHGYTARSPPIMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQK 778
H KFDHGYTA SPPI+NLLEI++EF+NE+RRAF+QFVT +PRLPPGGLASL+PKLT+V+K
Sbjct: 10 HIKFDHGYTASSPPIINLLEIVREFDNEQRRAFLQFVTGAPRLPPGGLASLNPKLTIVRK 189
Query: 779 ISYNHTDTDLPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
N DTDLPSVMTCANYLKLPPYSSKERMK+KLLYAITEG+G F
Sbjct: 190 HCSNRADTDLPSVMTCANYLKLPPYSSKERMKEKLLYAITEGQGSF 327
>TC215885 homologue to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ligase 3
(KAKTUS protein), partial (5%)
Length = 853
Score = 122 bits (307), Expect = 5e-28
Identities = 64/103 (62%), Positives = 76/103 (73%), Gaps = 13/103 (12%)
Frame = +2
Query: 735 NLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYN------------ 782
NLLEI+ EF E++RAF QFVT +PRLPPGGLA L+PKLT+V+K+S
Sbjct: 2 NLLEIMGEFTPEQQRAFCQFVTGAPRLPPGGLAVLNPKLTIVRKLSSTAVNNSSNGNGPS 181
Query: 783 -HTDTDLPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
D DLPSVMTCANYLKLPPYS+KE M +KLLYAI+EG+G F
Sbjct: 182 ESADDDLPSVMTCANYLKLPPYSTKEIMYKKLLYAISEGQGSF 310
>TC224122 weakly similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ligase
3 (KAKTUS protein), partial (3%)
Length = 672
Score = 96.3 bits (238), Expect = 5e-20
Identities = 74/167 (44%), Positives = 93/167 (55%), Gaps = 2/167 (1%)
Frame = +3
Query: 403 S*KSSCLFCGGS*NPE*PKITTTFF*TG*VFEQEVDREVGTTNGGSFGIVHWSYALLVLS 462
+*K+ C S* P * K ++F T *V +Q VDRE T+ G FG +H YA+ S
Sbjct: 153 A*KNLCFCPRES**PG*SKNNSSFCPTD*VCKQ*VDREARATDEGLFGCLHLWYAIAA*S 332
Query: 463 IDDLVSLPIQLCDKMQVL*IGDIWPATEP--SSRV**QP*NNERQKIEPQWIAAQEMSSF 520
I+ +S I KMQ+L IG IW AT P S R+ N+ Q EP I ++E S
Sbjct: 333 INGFMSFLI*F*GKMQILQIGTIWSATSPTFSQRL----KNSN*QATEPCRIVSKENFSS 500
Query: 521 SRPNFGVCCTNDEPECLSKSRS*SGI*W*SWLRFRTDFGILYLGL*G 567
SRP+F +CCTND P C S S* GI* S T+FGILYL + G
Sbjct: 501 SRPDFRICCTNDGPACQ**SGS*GGI**RSGHWSWTNFGILYLSVPG 641
>TC227511 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {Arabidopsis
thaliana;} , partial (44%)
Length = 1712
Score = 94.0 bits (232), Expect = 2e-19
Identities = 56/149 (37%), Positives = 84/149 (55%), Gaps = 3/149 (2%)
Frame = +2
Query: 677 SFFLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNL 736
S FL GF Q+ + + +F+E EL+L++ G +S +DL H + GY +
Sbjct: 1010 SHFLRGFQQLMQKDWIDMFNEHELQLLISGSLDSLDIDDLRLHTNYAGGYHGEHYVMEMF 1189
Query: 737 LEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTD---LPSVMT 793
E+L+ F+ E R+ F++FVT R P G L+P + +Q+ S N + LP+ T
Sbjct: 1190 WEVLKGFSLENRKKFLKFVTGCSRGPLLGFRYLEP-MFCIQRASGNAAEESLDRLPTSAT 1366
Query: 794 CANYLKLPPYSSKERMKQKLLYAITEGRG 822
C N LKLPPY+SKE+++ KLLYAI G
Sbjct: 1367 CMNLLKLPPYTSKEQLETKLLYAINADAG 1453
>TC216315 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein ligase 1
{Arabidopsis thaliana;} , partial (12%)
Length = 1938
Score = 86.7 bits (213), Expect = 4e-17
Identities = 54/152 (35%), Positives = 89/152 (58%), Gaps = 4/152 (2%)
Frame = +1
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL GFN++ P E + +F+++ELEL++ G P +DL + ++ GY+ SP I E
Sbjct: 1312 FLEGFNELIPRELISIFNDKELELLISGLPEI-DLDDLRANTEYS-GYSGASPVIQWFWE 1485
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI----SYNHTDTDLPSVMTC 794
++Q F+ E++ +QFVT + ++P G ++L ++ Q+ +Y +D LPS TC
Sbjct: 1486 VVQGFSKEDKARLLQFVTGTSKVPLEGFSALQG-ISGAQRFQIHKAYGSSD-HLPSAHTC 1659
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
N L LP Y SK+ ++++LL AI E F F
Sbjct: 1660 FNQLDLPEYPSKQHLEERLLLAIHEANEGFGF 1755
>TC226971 homologue to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein
ligase 1 {Arabidopsis thaliana;} , partial (7%)
Length = 1359
Score = 85.1 bits (209), Expect = 1e-16
Identities = 55/152 (36%), Positives = 87/152 (57%), Gaps = 4/152 (2%)
Frame = +1
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL GFN++ P E + +F+++ELEL++ G P +DL + ++ GYT S + E
Sbjct: 505 FLEGFNELVPRELISIFNDKELELLISGLPEI-DLDDLKANTEYT-GYTVASNVVQWFWE 678
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLD----PKLTVVQKISYNHTDTDLPSVMTC 794
+++ FN E+ +QFVT + ++P G +L P+ + K +Y D LPS TC
Sbjct: 679 VVKAFNKEDMARLLQFVTGTSKVPLEGFKALQGISGPQRFQIHK-AYGAPDR-LPSAHTC 852
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
N L LP Y+SKE+++++LL AI E F F
Sbjct: 853 FNQLDLPEYTSKEQLQERLLLAIHEASEGFGF 948
>TC216313 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein ligase
1 {Arabidopsis thaliana;} , partial (7%)
Length = 785
Score = 85.1 bits (209), Expect = 1e-16
Identities = 51/146 (34%), Positives = 87/146 (58%), Gaps = 3/146 (2%)
Frame = +1
Query: 677 SFFLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNL 736
++FL GFN++ P E + +F+++ELEL++ G P+ +DL + ++ GY+A SP I
Sbjct: 349 NYFLEGFNEMIPRELISIFNDKELELLISGLPDI-DLDDLRANTEYS-GYSAASPVIQWF 522
Query: 737 LEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNH---TDTDLPSVMT 793
E++Q + E++ +QFVT + ++P G ++L ++ QK + + LPS T
Sbjct: 523 WEVVQGLSKEDKARLLQFVTGTSKVPLEGFSALQ-GISGSQKFQIHKA*GSPEHLPSAHT 699
Query: 794 CANYLKLPPYSSKERMKQKLLYAITE 819
C N L LP Y SK ++++LL AI E
Sbjct: 700 CFNQLDLPEYPSKHHLEERLLLAIHE 777
>TC227510 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {Arabidopsis
thaliana;} , partial (14%)
Length = 730
Score = 83.6 bits (205), Expect = 3e-16
Identities = 51/144 (35%), Positives = 78/144 (53%), Gaps = 3/144 (2%)
Frame = +2
Query: 682 GFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQ 741
GF + + +F+E EL+L++ G +S +DL H + GY + E+L+
Sbjct: 5 GFQXXMQKDWIDMFNEHELQLLISGSLDSLDIDDLRLHTNYAGGYHNEHFVMEMFWEVLK 184
Query: 742 EFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDL---PSVMTCANYL 798
F+ E R+ F++FVT R P G L+P + + + S N + L P+ TC N L
Sbjct: 185 GFSLENRKKFLKFVTGCSRGPLLGFRYLEP-MFCIPRASGNAVEESLDRLPTSATCMNLL 361
Query: 799 KLPPYSSKERMKQKLLYAITEGRG 822
KLPPY+SKE+++ KLLYAI G
Sbjct: 362 KLPPYTSKEQLETKLLYAINADAG 433
>TC223924 weakly similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ligase
3 (KAKTUS protein), partial (7%)
Length = 755
Score = 72.4 bits (176), Expect = 8e-13
Identities = 47/103 (45%), Positives = 55/103 (52%), Gaps = 18/103 (17%)
Frame = +1
Query: 556 TDFGILYLGL*GVSKSRFRPVERG------------------YSVWSLSSPMVENAR*I* 597
T+FGILYLG+ GV + VER + +W++SS MV NAR I
Sbjct: 10 TNFGILYLGVSGVPEIWLGHVERRCFFLYPQDKHGG*GHRNPFLLWTISSSMVINARYIW 189
Query: 598 WPKDF*STKEICPFRSSCC*SYSRW*APRSPYLQSLS*TYLWK 640
W *S KE P R SCC*S SRW RSP L+S *TY WK
Sbjct: 190 WHTVL*SNKEFFPSRPSCC*STSRWKDLRSPLLKSFL*TYSWK 318
Score = 35.4 bits (80), Expect = 0.10
Identities = 13/19 (68%), Positives = 18/19 (94%)
Frame = +2
Query: 641 AFRNSKIEDLCLDFSLPGY 659
+FR+ +IEDLCLDF+LPG+
Sbjct: 455 SFRDKRIEDLCLDFTLPGF 511
Score = 29.6 bits (65), Expect = 5.7
Identities = 12/21 (57%), Positives = 16/21 (76%)
Frame = +1
Query: 691 HLRVFSEEELELILCGEPNSW 711
HLR+F+EEELE +L +SW
Sbjct: 670 HLRIFNEEELERMLSESGDSW 732
>TC215888 homologue to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ligase 3
(KAKTUS protein), partial (3%)
Length = 454
Score = 70.5 bits (171), Expect = 3e-12
Identities = 33/40 (82%), Positives = 35/40 (87%)
Frame = +3
Query: 785 DTDLPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
D DLPSVMTCANYLKLPPYS+KE M +KLLYAI EGRG F
Sbjct: 48 DDDLPSVMTCANYLKLPPYSTKEIMYKKLLYAINEGRGSF 167
>TC226972 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein ligase
1 {Arabidopsis thaliana;} , partial (3%)
Length = 957
Score = 64.7 bits (156), Expect = 2e-10
Identities = 46/132 (34%), Positives = 72/132 (53%), Gaps = 4/132 (3%)
Frame = +1
Query: 699 ELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEFNNEERRAFVQFVTRS 758
++EL++ G P +DL + ++ GYT S + E+++ FN E+ +QFVT +
Sbjct: 4 QVELLISGLPEI-DLDDLKANTEYT-GYTVASNVVQWFWEVVKAFNKEDMARLLQFVTGT 177
Query: 759 PRLPPGGLASLD----PKLTVVQKISYNHTDTDLPSVMTCANYLKLPPYSSKERMKQKLL 814
++P G +L P+ V K +Y D LPS TC N L LP Y+SKE+++++LL
Sbjct: 178 SKVPLEGFKALQGISGPQRFQVHK-AYGAPDR-LPSAHTCFNQLDLPEYTSKEQLQERLL 351
Query: 815 YAITEGRGCFLF 826
AI E F F
Sbjct: 352 LAIHEASEGFGF 387
>TC227512 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {Arabidopsis
thaliana;} , partial (17%)
Length = 541
Score = 60.5 bits (145), Expect = 3e-09
Identities = 38/117 (32%), Positives = 62/117 (52%)
Frame = +2
Query: 677 SFFLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNL 736
S FL GF Q+ + + +F+E EL+L++ G +S +DL +H + GY + I
Sbjct: 191 SHFLRGFQQLIQKDWIDMFNEHELQLLISGSLDSLDVDDLRQHTNYAGGYHSDHHVIEMF 370
Query: 737 LEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMT 793
E+L+ F+ E ++ F++FVT R P G L+P L +Q+ N D L + T
Sbjct: 371 WEVLKGFSLENKKKFLKFVTGCSRGPLLGFQYLEP-LFCIQRAGSNDPDEALDRLPT 538
>TC233256 weakly similar to UP|HUL5_YEAST (P53119) Probable
ubiquitin--protein ligase HUL5 , partial (3%)
Length = 732
Score = 58.5 bits (140), Expect = 1e-08
Identities = 39/117 (33%), Positives = 55/117 (46%), Gaps = 8/117 (6%)
Frame = +2
Query: 714 NDLLKHFKFDHGYTARSPPIMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKL 773
+DL + ++ GY S I E+++ F +ER ++FVT R P G L P
Sbjct: 38 DDLKNNTRYTGGYNEGSRTIKIFWEVIKGFEPKERCMLLKFVTSCSRAPLLGFKYLQPPF 217
Query: 774 TVVQKI-------SYNHTDTD-LPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
T+ + + D D LPS TC N LKLP Y ++ KLLYAI+ G
Sbjct: 218 TIHKVACDVPLWATIGGQDVDRLPSASTCYNTLKLPTYKRPGTLRAKLLYAISSNAG 388
>TC216312 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein ligase
1 {Arabidopsis thaliana;} , partial (4%)
Length = 621
Score = 58.2 bits (139), Expect = 1e-08
Identities = 38/106 (35%), Positives = 57/106 (52%), Gaps = 4/106 (3%)
Frame = +2
Query: 725 GYTARSPPIMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI----S 780
GY+ SP I E++Q + E++ +QFVT + ++P G ++L ++ QK +
Sbjct: 131 GYSXASPVIQWFWEVVQGLSKEDKARLLQFVTGTSKVPLEGFSALQG-ISGSQKFQIHKA 307
Query: 781 YNHTDTDLPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
Y D LPS TC N L LP Y SK ++++LL AI E F F
Sbjct: 308 YGSPD-HLPSAHTCFNQLDLPEYPSKHHLEERLLLAIHEASEGFGF 442
>CA952973
Length = 425
Score = 53.9 bits (128), Expect = 3e-07
Identities = 36/101 (35%), Positives = 48/101 (46%), Gaps = 8/101 (7%)
Frame = +3
Query: 730 SPPIMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI-------SYN 782
S PI E+++ F +ER ++FVT R P G L P T+ + +
Sbjct: 3 SRPIKIFWEVIKGFEPKERCMLLKFVTSCSRAPLLGFKYLQPPFTIHKVACDVPLWATIG 182
Query: 783 HTDTD-LPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
D D LPS TC N LKLP Y ++ KLLYAI+ G
Sbjct: 183 GQDVDRLPSASTCYNTLKLPTYKRPGTLRAKLLYAISSNAG 305
>TC215887 similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ligase 3
(KAKTUS protein), partial (3%)
Length = 937
Score = 52.8 bits (125), Expect = 6e-07
Identities = 32/72 (44%), Positives = 36/72 (49%), Gaps = 32/72 (44%)
Frame = +2
Query: 785 DTDLPSVMTCANYLKLPPYSSK--------------------------------ERMKQK 812
D DLPSVMTCANYLKLPPYS+K E M +K
Sbjct: 62 DDDLPSVMTCANYLKLPPYSTKVEHCKALLVI*SCMSKCLLVLNC*YLISILKQEIMYKK 241
Query: 813 LLYAITEGRGCF 824
LLYAI+EG+G F
Sbjct: 242 LLYAISEGQGSF 277
>TC208096 weakly similar to UP|Q9SU29 (Q9SU29) Polyubiquitin-like protein,
partial (19%)
Length = 789
Score = 44.7 bits (104), Expect = 2e-04
Identities = 41/157 (26%), Positives = 65/157 (41%), Gaps = 5/157 (3%)
Frame = +2
Query: 672 ITNLYSFFLVGFNQVFPIEHLRVF-----SEEELELILCGEPNSWTFNDLLKHFKFDHGY 726
I+ S F+ GF + L+ + E+L+ +L G ++ + D H +++ GY
Sbjct: 92 ISEQVSHFVKGFADILSNSKLQQYFFQSLDLEDLDWMLHGSEDTISVEDWKAHTEYN-GY 268
Query: 727 TARSPPIMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDT 786
I EI+ ++R+ + F T LP G L +L + + +
Sbjct: 269 KETDIQISWFWEIVGRMTADQRKVLLFFWTSVKYLPVEGFRGLASRLYIYRSLEPGDR-- 442
Query: 787 DLPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGC 823
LPS TC L P YSS MK +L E GC
Sbjct: 443 -LPSSHTCFFRLCFPAYSSMAVMKDRLEVITQEHIGC 550
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 37.7 bits (86), Expect = 0.021
Identities = 19/40 (47%), Positives = 26/40 (64%)
Frame = +3
Query: 780 SYNHTDTDLPSVMTCANYLKLPPYSSKERMKQKLLYAITE 819
+Y +D LPS TC N L LP Y SK+ ++++LL AI E
Sbjct: 444 AYGSSD-HLPSAHTCFNQLDLPEYPSKQHLEERLLLAIHE 560
>TC209867 similar to UP|Q6WWW4 (Q6WWW4) HECT ubiquitin-protein ligase 3
(KAKTUS protein), partial (10%)
Length = 691
Score = 36.6 bits (83), Expect = 0.046
Identities = 19/45 (42%), Positives = 26/45 (57%), Gaps = 4/45 (8%)
Frame = +2
Query: 626 RSPYLQSLS*TYLWKA----FRNSKIEDLCLDFSLPGYDQTFLLP 666
R Y++S+ +Y F + IEDLCLDF+LPGY + L P
Sbjct: 503 RKHYIESIGGSYTDTIVNLYFHGAPIEDLCLDFTLPGYPEYTLKP 637
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.358 0.160 0.582
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,665,960
Number of Sequences: 63676
Number of extensions: 649587
Number of successful extensions: 6968
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 3801
Number of HSP's successfully gapped in prelim test: 252
Number of HSP's that attempted gapping in prelim test: 3045
Number of HSP's gapped (non-prelim): 4279
length of query: 826
length of database: 12,639,632
effective HSP length: 105
effective length of query: 721
effective length of database: 5,953,652
effective search space: 4292583092
effective search space used: 4292583092
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC124963.1


