

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124958.16 + phase: 0
(248 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
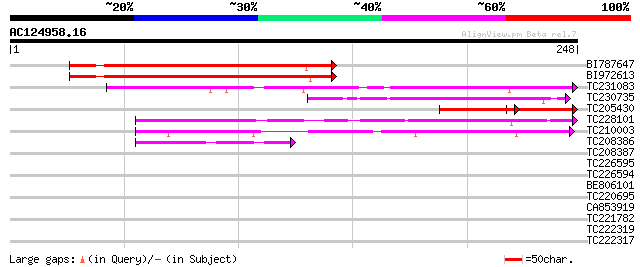
Score E
Sequences producing significant alignments: (bits) Value
BI787647 172 1e-43
BI972613 171 3e-43
TC231083 96 2e-20
TC230735 65 4e-11
TC205430 similar to UP|O22600 (O22600) Glycine-rich protein, par... 53 1e-07
TC228101 49 2e-06
TC210003 47 1e-05
TC208386 46 1e-05
TC208387 39 0.003
TC226595 similar to UP|Q9SHG8 (Q9SHG8) SOUL-like protein, partia... 35 0.024
TC226594 weakly similar to UP|Q9SHG8 (Q9SHG8) SOUL-like protein,... 34 0.071
BE806101 32 0.21
TC220695 weakly similar to UP|Q8W1C6 (Q8W1C6) Ring-H2 zinc finge... 28 3.9
CA853919 homologue to PIR|T11622|T11 extensin class 1 precursor ... 28 3.9
TC221782 28 5.1
TC222319 UP|Q39833 (Q39833) Alfa-carboxyltransferase precursor (... 27 6.6
TC222317 UP|Q9LLQ8 (Q9LLQ8) Carboxyl transferase alpha subunit ... 27 6.6
>BI787647
Length = 423
Score = 172 bits (436), Expect = 1e-43
Identities = 90/121 (74%), Positives = 99/121 (81%), Gaps = 4/121 (3%)
Frame = +2
Query: 27 NLLLPTSFNSHFTLSHSRSIMGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYD 86
+LL P S ++HF S S+MG+VFGKI VET KYEV K+T +YEIR YAPSV AEVTYD
Sbjct: 68 SLLSPNSQHTHF---RSLSLMGLVFGKISVETAKYEVIKSTSEYEIRKYAPSVVAEVTYD 238
Query: 87 PSQFKGNKDGGFMVLANYIGALGNPQNTKPEKIAMTAPVITK----GSAEKIAMTAPVVT 142
PSQFKGNKDGGFM+LANYIGA+G PQNTKPEKIAMTAPVITK G E IAMTAPVVT
Sbjct: 239 PSQFKGNKDGGFMILANYIGAVGKPQNTKPEKIAMTAPVITKDSVGGDGETIAMTAPVVT 418
Query: 143 K 143
K
Sbjct: 419 K 421
>BI972613
Length = 423
Score = 171 bits (433), Expect = 3e-43
Identities = 91/124 (73%), Positives = 100/124 (80%), Gaps = 7/124 (5%)
Frame = +2
Query: 27 NLLLPTSFNSHFTLSHSRSIMGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYD 86
+LL P S ++HF S S+MG+VFGKI VET KYEV K+T +YEIR YAPSV AEVTYD
Sbjct: 53 SLLSPKSQHTHF---RSLSLMGLVFGKITVETAKYEVIKSTSEYEIRKYAPSVVAEVTYD 223
Query: 87 PSQFKGNKDGGFMVLANYIGALGNPQNTKPEKIAMTAPVITKGS-------AEKIAMTAP 139
PSQFKGNKDGGFM+LANYIGA+G PQNTKPEKIAMTAPVITK S EKIAMTAP
Sbjct: 224 PSQFKGNKDGGFMILANYIGAVGKPQNTKPEKIAMTAPVITKDSVGGGGSDGEKIAMTAP 403
Query: 140 VVTK 143
VVTK
Sbjct: 404 VVTK 415
>TC231083
Length = 1086
Score = 95.5 bits (236), Expect = 2e-20
Identities = 73/214 (34%), Positives = 103/214 (48%), Gaps = 8/214 (3%)
Frame = +1
Query: 43 SRSIMGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYD-PSQFKGN-KDGGFMV 100
SR++ + +ET ++V YEIR P AE T S F N F
Sbjct: 361 SRNLEEALMSVPDLETVDFKVLSRMDQYEIREVEPYFVAETTMPGKSGFDFNGASRSFNA 540
Query: 101 LANYIGALGNPQNTKPEKIAMTAPVIT---KGSAEKIAMTAPVVTKSSEEGERNKMVTMQ 157
LA Y+ +NT EK+ MT PV T + K+ MT PV+T E+ + KM
Sbjct: 541 LAEYLFG----KNTTKEKMEMTTPVFTSKNQSDGVKMDMTTPVLTTKMEDQDNWKM---S 699
Query: 158 FILPSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDG 217
F++PS Y P P D V I+E + VV FSG +DE +K++ KLR +L+ D
Sbjct: 700 FVMPSKY--GANLPLPKDSSVRIKEVPRKIVAVVSFSGFVNDEEIKQRELKLRDALKSDS 873
Query: 218 ---FKVIGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
K + +YNPP+TLP R NE+ + +E
Sbjct: 874 QFEIKEGTSVEVAQYNPPFTLPFQRRNEIALEVE 975
>TC230735
Length = 612
Score = 64.7 bits (156), Expect = 4e-11
Identities = 45/118 (38%), Positives = 67/118 (56%), Gaps = 3/118 (2%)
Frame = +3
Query: 131 AEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGV 190
A +I MT PV T++++ + +K V++Q +LP E E P P E V +R+ V
Sbjct: 6 ASEIPMTTPVFTETND-ADLSK-VSIQIVLPLDKE-TESLPNPNQETVRLRKVEGGIAAV 176
Query: 191 VKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPP---WTLPMFRTNEVMI 245
+KFSG +++ V+EK + LR ++ +DG K LL RYN P WT M NEV+I
Sbjct: 177 MKFSGKPTEDTVREKEKTLRANIIKDGLKPQSGCLLARYNDPGRTWTFIM--RNEVLI 344
>TC205430 similar to UP|O22600 (O22600) Glycine-rich protein, partial (19%)
Length = 421
Score = 53.1 bits (126), Expect = 1e-07
Identities = 22/31 (70%), Positives = 27/31 (86%)
Frame = +3
Query: 218 FKVIGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
++ +G LLGRYNPPWT+P FRTNEVMIP+E
Sbjct: 156 WRKMGLRLLGRYNPPWTIPAFRTNEVMIPVE 248
Score = 52.8 bits (125), Expect = 1e-07
Identities = 25/35 (71%), Positives = 31/35 (88%)
Frame = +2
Query: 189 GVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGD 223
GV SGVAS++VV+EKVEKLR SLE+DGFKV+G+
Sbjct: 83 GVGNVSGVASEQVVREKVEKLRESLEKDGFKVVGE 187
>TC228101
Length = 952
Score = 48.9 bits (115), Expect = 2e-06
Identities = 51/196 (26%), Positives = 79/196 (40%), Gaps = 3/196 (1%)
Frame = +3
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
+E+P++ V + D+EIR+Y SV F+ GF L + N
Sbjct: 81 IESPQHTVVHSESDFEIRLYRTSVWMSAPALDISFEKATWNGFHRLFQFTEGA----NLN 248
Query: 116 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
+I MT PV+T P +G + LP ++ P P +
Sbjct: 249 FSRIPMTIPVLTTA--------VPGAGPLQSQG-----YYVSLYLPVKFQGDPPVPLP-E 386
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGF---KVIGDFLLGRYNPP 232
+ E V KFSG A DE + ++ EKL SL R + K + + +YN P
Sbjct: 387 LNIKPYEFSSHCVAVRKFSGFAKDERIVKEAEKLATSLSRSPWAESKTGRGYSIAQYNTP 566
Query: 233 WTLPMFRTNEVMIPIE 248
+ + R NEV + I+
Sbjct: 567 IRI-VKRKNEVWVDID 611
>TC210003
Length = 896
Score = 46.6 bits (109), Expect = 1e-05
Identities = 48/197 (24%), Positives = 82/197 (41%), Gaps = 5/197 (2%)
Frame = -2
Query: 56 VETPKYEVTKTTQ-DYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYI-GALGNPQN 113
+E P Y V + D+++R+Y S + F+ + GF L YI GA N
Sbjct: 748 IELPNYTVILPEESDFQLRLYNESSWISARVSGTSFEQSYKLGFSRLYQYIHGANSN--- 578
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+ KIA TAPV+T R+ + F+ S + P+P
Sbjct: 577 -----------------SSKIAFTAPVLTSVPSSPPRDGYIVRMFV---STHFQGKPPQP 458
Query: 174 TDE-RVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKV--IGDFLLGRYN 230
E ++ I + + V KF+G A D+ + +++E L +L ++ + + + +YN
Sbjct: 457 NPELKLRIEKWKTQCIAVRKFTGYAKDDNINKEIEALVTTLNKNSATIQDTSFYTIAKYN 278
Query: 231 PPWTLPMFRTNEVMIPI 247
R NEV I +
Sbjct: 277 ASSHNTADRLNEVWIKV 227
>TC208386
Length = 757
Score = 46.2 bits (108), Expect = 1e-05
Identities = 27/70 (38%), Positives = 39/70 (55%)
Frame = +1
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
+E+PKY++ K T++YE+R Y P + E D K + GF +A YI +N+
Sbjct: 571 LESPKYQILKRTENYEVRQYNPFIVVETNGD----KLSGSTGFNDVAGYIFG----KNST 726
Query: 116 PEKIAMTAPV 125
EKI MT PV
Sbjct: 727 TEKIPMTTPV 756
>TC208387
Length = 487
Score = 38.5 bits (88), Expect = 0.003
Identities = 23/64 (35%), Positives = 35/64 (53%)
Frame = +2
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
+E+PKY++ K T++YE+R Y P + E D K + GF +A YI +N+
Sbjct: 317 LESPKYQILKRTENYEVRQYNPFIVVETNGD----KLSGSTGFNDVAGYIFG----KNST 472
Query: 116 PEKI 119
EKI
Sbjct: 473 TEKI 484
>TC226595 similar to UP|Q9SHG8 (Q9SHG8) SOUL-like protein, partial (59%)
Length = 950
Score = 35.4 bits (80), Expect = 0.024
Identities = 41/160 (25%), Positives = 62/160 (38%), Gaps = 2/160 (1%)
Frame = +1
Query: 56 VETPKYEVTKTTQDYEIRIYAPSV--AAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
+E P Y+V YEIR Y V + D S + + GF L +YI +N
Sbjct: 193 IECPSYDVIHVGNGYEIRRYNSPVWISNSPIQDISLVEATRT-GFRRLFDYI----QGKN 357
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+KI MTAPVI +E + P S F++ K +A P
Sbjct: 358 NYKQKIEMTAPVI----SEVLPSDGPFCESS-------------FVVSFYVPKENQANPP 486
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSL 213
+ + ++ V +F G D V E+ L+ S+
Sbjct: 487 PAKGLQVQRWKTVFVAVRQFGGFVKDSSVGEEAAALKASI 606
>TC226594 weakly similar to UP|Q9SHG8 (Q9SHG8) SOUL-like protein, partial
(77%)
Length = 901
Score = 33.9 bits (76), Expect = 0.071
Identities = 41/160 (25%), Positives = 62/160 (38%), Gaps = 2/160 (1%)
Frame = +3
Query: 56 VETPKYEVTKTTQDYEIRIYAPSV--AAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
+E P Y+V YEIR Y V + D S + + G F L +YI +N
Sbjct: 84 IECPSYDVIHFGNGYEIRRYNSPVWISNSPILDISLVEATRTG-FRRLFDYIQG----KN 248
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+KI MTAPVI +E + P S F++ K +A P
Sbjct: 249 NYKQKIEMTAPVI----SEVLPRDGPFCESS-------------FVVSFYVPKENQANPP 377
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSL 213
+ + ++ V +F G D V E+ L+ S+
Sbjct: 378 PAKGLHVQRWKTVFAAVRQFGGFVKDSSVGEEAAALKASI 497
>BE806101
Length = 421
Score = 32.3 bits (72), Expect = 0.21
Identities = 17/61 (27%), Positives = 31/61 (49%), Gaps = 5/61 (8%)
Frame = +3
Query: 56 VETPKYEVTKTTQDYEIRIYAPSV-----AAEVTYDPSQFKGNKDGGFMVLANYIGALGN 110
+E+P++ V + D+EIR+Y SV A +++++ + + G NY A N
Sbjct: 30 IESPQHTVVHSESDFEIRLYRTSVWMSAPALDISFEKATWNGFHRYSIHFFFNYSTAHSN 209
Query: 111 P 111
P
Sbjct: 210 P 212
>TC220695 weakly similar to UP|Q8W1C6 (Q8W1C6) Ring-H2 zinc finger protein,
partial (15%)
Length = 556
Score = 28.1 bits (61), Expect = 3.9
Identities = 15/40 (37%), Positives = 21/40 (52%)
Frame = +2
Query: 24 THKNLLLPTSFNSHFTLSHSRSIMGMVFGKIGVETPKYEV 63
TH +L PT FNSH L H S + +F + T K ++
Sbjct: 80 THHSLSSPTFFNSHLELYHLTSDVLKIFIICSLSTLKLDM 199
>CA853919 homologue to PIR|T11622|T11 extensin class 1 precursor - cowpea,
partial (30%)
Length = 454
Score = 28.1 bits (61), Expect = 3.9
Identities = 29/110 (26%), Positives = 48/110 (43%), Gaps = 1/110 (0%)
Frame = -2
Query: 117 EKIAMTAPVITKGSA-EKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
E + M V+ GS+ E+ VV S GE + + ++ I Y E + +
Sbjct: 348 EVVVMEMVVVENGSSREEEEREMVVVGSGSSRGEEERGMVVEEI----YSNREVVGEIYN 181
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFL 225
+ EEG K GVV ++ + +E V +VE + KV+G+ L
Sbjct: 180 SK---EEEGMVKVGVVIYNSMGEEEKVMVEVESDNSMGV*ELVKVVGEHL 40
>TC221782
Length = 750
Score = 27.7 bits (60), Expect = 5.1
Identities = 12/22 (54%), Positives = 16/22 (72%)
Frame = -1
Query: 22 STTHKNLLLPTSFNSHFTLSHS 43
S T +N+LLP+ +HFTL HS
Sbjct: 408 SATTENVLLPSLSLTHFTLEHS 343
>TC222319 UP|Q39833 (Q39833) Alfa-carboxyltransferase precursor (Carboxyl
transferase alpha subunit) , complete
Length = 2588
Score = 27.3 bits (59), Expect = 6.6
Identities = 16/52 (30%), Positives = 27/52 (51%)
Frame = +2
Query: 164 YEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLER 215
Y +A +A TD + +REE + D ++K+K+EKLR+ E+
Sbjct: 1766 YSEAVKATGLTDSLLKLREEVSK----ANADNQIVDPLLKDKIEKLRVEFEQ 1909
>TC222317 UP|Q9LLQ8 (Q9LLQ8) Carboxyl transferase alpha subunit , complete
Length = 2521
Score = 27.3 bits (59), Expect = 6.6
Identities = 16/52 (30%), Positives = 27/52 (51%)
Frame = +3
Query: 164 YEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLER 215
Y +A +A TD + +REE + D ++K+K+EKLR+ E+
Sbjct: 1785 YSEAVKATGLTDSLLKLREEVSK----ANADSQIVDPLLKDKIEKLRVEFEQ 1928
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.133 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,806,923
Number of Sequences: 63676
Number of extensions: 103076
Number of successful extensions: 538
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 521
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 529
length of query: 248
length of database: 12,639,632
effective HSP length: 95
effective length of query: 153
effective length of database: 6,590,412
effective search space: 1008333036
effective search space used: 1008333036
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC124958.16


