

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124952.2 - phase: 0
(880 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
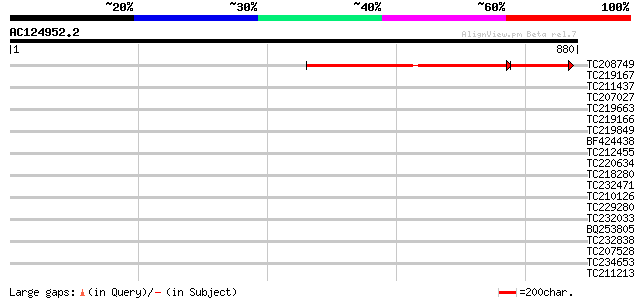
Score E
Sequences producing significant alignments: (bits) Value
TC208749 weakly similar to UP|Q6WNS7 (Q6WNS7) Transcription fact... 546 0.0
TC219167 similar to GB|AAA36308.1|386949|HUMMHRD5 MHC HLA-RD pro... 42 0.001
TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%) 39 0.008
TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%) 38 0.017
TC219663 similar to GB|AAA36308.1|386949|HUMMHRD5 MHC HLA-RD pro... 37 0.029
TC219166 similar to GB|AAA36308.1|386949|HUMMHRD5 MHC HLA-RD pro... 37 0.029
TC219849 weakly similar to UP|Q7RP82 (Q7RP82) Casein kinase ii b... 36 0.064
BF424438 36 0.084
TC212455 weakly similar to UP|Q7QIA8 (Q7QIA8) AgCP3984 (Fragment... 35 0.11
TC220634 35 0.11
TC218280 similar to UP|Q9LMF3 (Q9LMF3) F16A14.27, partial (12%) 35 0.14
TC232471 similar to UP|NLFE_HUMAN (P18615) Negative elongation f... 35 0.19
TC210126 similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C2... 35 0.19
TC229280 similar to UP|NK61_HUMAN (P78426) Homeobox protein Nkx-... 34 0.32
TC232033 34 0.32
BQ253805 34 0.32
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 33 0.42
TC207528 similar to UP|Q76LJ5 (Q76LJ5) Serine threonine Kinase, ... 33 0.55
TC234653 weakly similar to UP|Q6VSS7 (Q6VSS7) TOX, partial (10%) 33 0.55
TC211213 similar to PIR|T48518|T48518 transcription factor like ... 33 0.71
>TC208749 weakly similar to UP|Q6WNS7 (Q6WNS7) Transcription factor Sox10a,
partial (4%)
Length = 1980
Score = 546 bits (1408), Expect(2) = 0.0
Identities = 265/319 (83%), Positives = 293/319 (91%)
Frame = +1
Query: 461 DPQQSVPSRDGGLTASTSSTDDACLLVKRPRRIDRIEVVGARQKRGDVSFSERLVGVKEY 520
+PQ S+PSRDGGLTASTSS DDA LLV+ PR+IDRIEVVGARQK+GDVSFSERLVGVKEY
Sbjct: 4 EPQFSMPSRDGGLTASTSSKDDAYLLVQCPRKIDRIEVVGARQKKGDVSFSERLVGVKEY 183
Query: 521 TVYKIKVWSGKDQWEVEKRYRDFLTLHRCMKSLFNEQGWTLPLPWSSVEKESKIFRSASL 580
TVYKIKVWSGKDQWEVE+RYRDFLTL+R MK+LFNEQGW LPLPWSSVEKE++IFRSAS
Sbjct: 184 TVYKIKVWSGKDQWEVERRYRDFLTLYRYMKTLFNEQGWKLPLPWSSVEKETQIFRSASP 363
Query: 581 DIIAKRSVLIQECLQSILCSRFFSSPPRALVWFLSPEDSHPSSPVSNSPVSLSSFTRGEN 640
DII KRSVLIQ+CLQSI+ SRF SSPPRAL+WF+S +DS+P SPVS+ SFTRGEN
Sbjct: 364 DIIVKRSVLIQDCLQSIIRSRFSSSPPRALIWFISHQDSYPISPVSH------SFTRGEN 525
Query: 641 IRNSSTWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKP 700
IR+ S GKTISLIVEIP NKS +QLLE+QHHTCAGCH+HFDDG + IWDFVQTFGWGKP
Sbjct: 526 IRSISNLGKTISLIVEIPPNKSVKQLLESQHHTCAGCHKHFDDGKTLIWDFVQTFGWGKP 705
Query: 701 RLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVN 760
RLCEYTGQLFCSSCHTN+TAVLPARVLH+WDFT+YPVSQLAKSYLDSI+E PMLCVTAVN
Sbjct: 706 RLCEYTGQLFCSSCHTNQTAVLPARVLHNWDFTYYPVSQLAKSYLDSIYEQPMLCVTAVN 885
Query: 761 PFLVSKVPALLHVMSVRKK 779
PFL+SKVPALLH+MSVRKK
Sbjct: 886 PFLLSKVPALLHIMSVRKK 942
Score = 191 bits (484), Expect(2) = 0.0
Identities = 89/97 (91%), Positives = 96/97 (98%)
Frame = +2
Query: 778 KKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKGVFAALPVMVETVSR 837
KKIGTMLPYVRCPFRRSIN+G+G+RRYLLESNDFFALRDLIDLS+GVFAALPVMV+TVSR
Sbjct: 938 KKIGTMLPYVRCPFRRSINRGLGSRRYLLESNDFFALRDLIDLSRGVFAALPVMVDTVSR 1117
Query: 838 KILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
KILEHITDQCL+CCDVG PC+ARQDCSDPSSLIFPFQ
Sbjct: 1118 KILEHITDQCLICCDVGDPCNARQDCSDPSSLIFPFQ 1228
>TC219167 similar to GB|AAA36308.1|386949|HUMMHRD5 MHC HLA-RD protein {Homo
sapiens;} , partial (11%)
Length = 600
Score = 42.0 bits (97), Expect = 0.001
Identities = 32/119 (26%), Positives = 48/119 (39%), Gaps = 10/119 (8%)
Frame = +3
Query: 233 RESNNRVEEHGVSSVRKEEKDSLIVN---DEVEETKDIGDRE-------ALEEVRDRDRD 282
++S +E S E +D I+ D + ET+ DR+ E RDRDRD
Sbjct: 66 KDSKRSDDESKGMSTEGEARDEPILKSKADHIGETRRNRDRDRDRERDRGRERDRDRDRD 245
Query: 283 RDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGAS 341
R + RDRD + + + D L + + D G K R+S +D S
Sbjct: 246 RTKSRDRDRGRDSDKEREREEAERDKLKDRGHRSRERAKDSGHSEKSKRHSSHDRVSYS 422
>TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%)
Length = 857
Score = 39.3 bits (90), Expect = 0.008
Identities = 21/55 (38%), Positives = 33/55 (59%)
Frame = +1
Query: 93 ELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSE 147
E G+EGDE +++ ++G E+ DG++NG E + + E EEE+EEE E
Sbjct: 439 EFSGEEGDEGGDPEDEANGEGSDDGDEDDDGDDNGDEEDDDDGEDEEEDEEEDEE 603
>TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%)
Length = 1005
Score = 38.1 bits (87), Expect = 0.017
Identities = 42/203 (20%), Positives = 79/203 (38%), Gaps = 1/203 (0%)
Frame = +3
Query: 240 EEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRD-RDRDAPVSCEVQ 298
E+ G KE+KD + K GD E +E +D+++++ ++ +D+D
Sbjct: 291 EDDGKDEGNKEKKD---------KEKGDGDGEEKKEKKDKEKEKKKEKKDKD-------- 419
Query: 299 CADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKS 358
E D K D G + +GN+ K D+ D ++E D + E K
Sbjct: 420 -----------EETDTLKEKGKNDEGEDDEGNKKKKKDKKEKEKDHKKEKKDKEEGE-KE 563
Query: 359 DQFCDNRDDVSTSNVSVHVENVDAKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHV 418
D S V V V ++D + K G +++ E+ +K
Sbjct: 564 D-----------SKVEVSVRDIDIEEIKKEGEKEDKGKDGGKEVKEKKKKEDK--DKKEK 704
Query: 419 ISKIEDSELSEFYDEVVQEMEEI 441
K+ + ++ + Q++E+I
Sbjct: 705 KKKVTGKDKTKDLSTLKQKLEKI 773
>TC219663 similar to GB|AAA36308.1|386949|HUMMHRD5 MHC HLA-RD protein {Homo
sapiens;} , partial (14%)
Length = 1176
Score = 37.4 bits (85), Expect = 0.029
Identities = 28/103 (27%), Positives = 44/103 (42%)
Frame = +3
Query: 244 VSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNS 303
+S R +KD + + DR+ E+ RDRDR+RDRDRD + D
Sbjct: 282 ISRERGRDKDRERERSREHSHERVRDRDHREDRHHRDRDRNRDRDRD-----RERDRDRG 446
Query: 304 PDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQR 346
D D + D ++ D + + +R+ + D GD R
Sbjct: 447 RDRDRTRDRDRERG---RDRDRDREYDRHRERDRDYEVGDPDR 566
Score = 37.0 bits (84), Expect = 0.038
Identities = 19/50 (38%), Positives = 27/50 (54%)
Frame = +3
Query: 241 EHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRD 290
E G R+ E+ ++ V + DR + R+RDRDRDR+RDRD
Sbjct: 291 ERGRDKDRERERSREHSHERVRDRDHREDRHHRDRDRNRDRDRDRERDRD 440
Score = 34.3 bits (77), Expect = 0.24
Identities = 29/112 (25%), Positives = 49/112 (42%), Gaps = 12/112 (10%)
Frame = +3
Query: 233 RESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIG-DREAL-----EEVRDRDRDRDRD 286
R+ ++R + H R ++D + E +D G DR+ E RDRDRDR+ D
Sbjct: 354 RDRDHREDRHHRDRDRNRDRDR-----DRERDRDRGRDRDRTRDRDRERGRDRDRDREYD 518
Query: 287 RDRDAPVSCEV------QCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRY 332
R R+ EV + D D D + + +++ + G+ N+Y
Sbjct: 519 RHRERDRDYEVGDPDRGRSRDRESDYDRVESKHGERNHDYEPEDDRGRHNQY 674
>TC219166 similar to GB|AAA36308.1|386949|HUMMHRD5 MHC HLA-RD protein {Homo
sapiens;} , partial (12%)
Length = 774
Score = 37.4 bits (85), Expect = 0.029
Identities = 23/75 (30%), Positives = 34/75 (44%), Gaps = 1/75 (1%)
Frame = +3
Query: 274 EEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYS 333
E RDRDRDR + RDRD + + + L + D + D G K R+S
Sbjct: 288 ERDRDRDRDRTKSRDRDRGRDSDKEREREEAERGKLKDRDHRSRERAKDSGHSEKSKRHS 467
Query: 334 KND-EAGASGDAQRE 347
+D ++ S + RE
Sbjct: 468 SHDRDSHGSSYSSRE 512
>TC219849 weakly similar to UP|Q7RP82 (Q7RP82) Casein kinase ii beta chain,
partial (14%)
Length = 587
Score = 36.2 bits (82), Expect = 0.064
Identities = 22/59 (37%), Positives = 31/59 (52%), Gaps = 9/59 (15%)
Frame = +2
Query: 115 NNGIEESDGNEN-GGEVGEREIETEEEEEEEFSEGD--------DSMFNYGSGCDGDNE 164
N G EE+D NE GE G+ + +EE+EE+ S+ D D +Y DGD+E
Sbjct: 107 NQGKEENDENEKKDGEDGDEDENAQEEDEEDISDDDYNQNIDFDDDEDDYNDVDDGDDE 283
>BF424438
Length = 393
Score = 35.8 bits (81), Expect = 0.084
Identities = 18/30 (60%), Positives = 21/30 (70%), Gaps = 2/30 (6%)
Frame = +2
Query: 263 ETKDIGDREALEEVRDRDRD--RDRDRDRD 290
+ +D GDR+ RDRDRD RDRDRDRD
Sbjct: 239 DRRDHGDRDRDRRDRDRDRDDRRDRDRDRD 328
>TC212455 weakly similar to UP|Q7QIA8 (Q7QIA8) AgCP3984 (Fragment), partial
(11%)
Length = 944
Score = 35.4 bits (80), Expect = 0.11
Identities = 23/65 (35%), Positives = 29/65 (44%)
Frame = +1
Query: 226 SDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDR 285
SD ++ R EHG SV V++ ++ R E RDRDR RDR
Sbjct: 136 SDYDHDEERVRVRGREHGHGSV-------------VDDRREYSTRYHNREARDRDRSRDR 276
Query: 286 DRDRD 290
D DRD
Sbjct: 277 DVDRD 291
Score = 32.3 bits (72), Expect = 0.93
Identities = 20/58 (34%), Positives = 29/58 (49%)
Frame = +1
Query: 233 RESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRD 290
R S+ +E V +E +V+D E + +REA + R RDRD DRD R+
Sbjct: 130 RRSDYDHDEERVRVRGREHGHGSVVDDRREYSTRYHNREARDRDRSRDRDVDRDLHRE 303
Score = 30.4 bits (67), Expect = 3.5
Identities = 47/201 (23%), Positives = 79/201 (38%), Gaps = 17/201 (8%)
Frame = +1
Query: 259 DEVEETKDIGDREALEE---------VRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLL 309
D E ++ GDR +E RDR R+++RDR RD + +
Sbjct: 409 DRSREREEDGDRRQEKERDRSWDTVFERDRRREKERDRSRDRTIGGK------------- 549
Query: 310 PEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVS 369
+ DP+ + D E + +RY D A+G + + + DN D
Sbjct: 550 RDRDPENERD--DKNRERRDDRYGHKDRDTANGKDR---------HLRHEDGNDNGDRYR 696
Query: 370 TSNVSVHVENV---DAKSFKNLKPIVLPSNGGTRKILERSSTSTNV----LEKSHV-ISK 421
S H EN + K + V S G T ++ E TS+ V LE+ + + +
Sbjct: 697 KH--SRHEENEYHWERKRNSDNPVKVYNSMGSTAEVDESKLTSSEVEPDDLEEDTLQLPE 870
Query: 422 IEDSELSEFYDEVVQEMEEIL 442
E+ +L+ +E + E I+
Sbjct: 871 QEEEDLNRIKEESRRRREAIM 933
>TC220634
Length = 583
Score = 35.4 bits (80), Expect = 0.11
Identities = 43/191 (22%), Positives = 78/191 (40%), Gaps = 28/191 (14%)
Frame = +2
Query: 162 DNENEFYSMKGENVSSEFYSSRSVSLYEE-----GEVRNENPLLMNSSVAFGSHDLD--- 213
D ENE SM EN + S+ EE E N N + D D
Sbjct: 14 DKENELQSMFQENDELRSREAESIKKLEELSKLLEEATTRNHTEENGDLTDSEKDYDLLP 193
Query: 214 ---DFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEE-------KDSLIVNDEVEE 263
+F +NG V E ++VE SV +EE +DS++ ND+ E+
Sbjct: 194 KVVEFSEENGLVG----------EDISKVE----LSVNQEELKQNNMQEDSILSNDKAEK 331
Query: 264 TK-----DIGDREALEEVRDRDRDRDRDRDRDAPV----SCEVQCADNSPDLDLLPEE-D 313
+ ++ + +E ++++ +++D + SC+++ + SP+ + PE +
Sbjct: 332 IESPKHEEVSGKRKEDETKEKEESKEKDDSVEVEYKMWESCKIEKKEFSPEREAEPESFE 511
Query: 314 PQKSLNITDGG 324
+ + I GG
Sbjct: 512 EEVNSKIEKGG 544
>TC218280 similar to UP|Q9LMF3 (Q9LMF3) F16A14.27, partial (12%)
Length = 624
Score = 35.0 bits (79), Expect = 0.14
Identities = 29/95 (30%), Positives = 40/95 (41%)
Frame = +3
Query: 119 EESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSE 178
E+ N+ G E E + E EEE+EEE E +F G GC +EF +
Sbjct: 132 EDLHPNQPGSESPEAKAEGEEEDEEEEGECGFCLFMKGGGC----RDEFIGWE------- 278
Query: 179 FYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLD 213
+ +E E NE+ L + V GS LD
Sbjct: 279 -------NCVQEAENNNEDLLEKCAQVTAGSQALD 362
>TC232471 similar to UP|NLFE_HUMAN (P18615) Negative elongation factor E
(NELF-E) (RD protein), partial (11%)
Length = 480
Score = 34.7 bits (78), Expect = 0.19
Identities = 18/48 (37%), Positives = 27/48 (55%), Gaps = 5/48 (10%)
Frame = +3
Query: 248 RKEEKDSLIVNDEVEETKDIGDREALEEV-----RDRDRDRDRDRDRD 290
+K ++ L+ + + K DR+ E RDRDRDRDRDR+R+
Sbjct: 231 QKRRQEQLLSRNHSSDPKPSSDRDRDRERDRDHHRDRDRDRDRDRERE 374
>TC210126 similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23), partial
(7%)
Length = 735
Score = 34.7 bits (78), Expect = 0.19
Identities = 21/55 (38%), Positives = 28/55 (50%)
Frame = +3
Query: 93 ELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSE 147
E G+EGDE D E G ++ D N+N + + E E EEE+EEE E
Sbjct: 411 EFSGEEGDEEGDPE-DDPEANGGGSDDDDDDNDNEEDDDDGEDEEEEEDEEEDEE 572
>TC229280 similar to UP|NK61_HUMAN (P78426) Homeobox protein Nkx-6.1, partial
(5%)
Length = 463
Score = 33.9 bits (76), Expect = 0.32
Identities = 18/44 (40%), Positives = 24/44 (53%)
Frame = +2
Query: 115 NNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSG 158
N+ +EE N NG E ++E EEEEEEE +D + G G
Sbjct: 188 NHNLEEEHSNLNGEEQMAEQVEAEEEEEEE----EDRQADEGEG 307
>TC232033
Length = 865
Score = 33.9 bits (76), Expect = 0.32
Identities = 33/111 (29%), Positives = 50/111 (44%), Gaps = 8/111 (7%)
Frame = +2
Query: 277 RDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKND 336
RDRDR RDRD+DRD + ++ D D ++++ D E +
Sbjct: 17 RDRDRSRDRDQDRD-------RRFEHMHDYD-----QAREAMLENDWIREDDRVENEQEH 160
Query: 337 EAGASGDAQRE-NLDLDNFEFKSDQ-------FCDNRDDVSTSNVSVHVEN 379
G GD R+ N+DL N + K D+ F ++R D S N ++V N
Sbjct: 161 SRGHGGDVDRDHNIDL-NVDRKMDRTSDPDRSFGEDRKDQSRRNNDLNVTN 310
>BQ253805
Length = 425
Score = 33.9 bits (76), Expect = 0.32
Identities = 18/44 (40%), Positives = 24/44 (53%)
Frame = +3
Query: 115 NNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSG 158
N+ +EE N NG E ++E EEEEEEE +D + G G
Sbjct: 210 NHNLEEEHSNLNGEEQMAEQVEAEEEEEEE----EDRQADEGEG 329
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 33.5 bits (75), Expect = 0.42
Identities = 31/120 (25%), Positives = 55/120 (45%), Gaps = 1/120 (0%)
Frame = +2
Query: 233 RESNNRVEEHGVS-SVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDA 291
+E + +VEE+ V V +++++ V++ V++ +D D E E+ D + D + D + DA
Sbjct: 233 KELSEQVEENEVKVLVEEDDEEYESVDEVVDDAEDDEDEE--EDDDDDEGDDNDDEEDDA 406
Query: 292 PVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDL 351
P D+ D D E D Q+ D ++ + DE + Q E DL
Sbjct: 407 P-----DGGDDDDDEDDEEEGDVQRGGEPDDDDNDSDDDSDDDEDEDEEDEEEQGEEEDL 571
>TC207528 similar to UP|Q76LJ5 (Q76LJ5) Serine threonine Kinase, partial (4%)
Length = 757
Score = 33.1 bits (74), Expect = 0.55
Identities = 23/94 (24%), Positives = 43/94 (45%)
Frame = +3
Query: 296 EVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFE 355
+V+ D S + + PEEDP++ + +G + + + KN E A D + EN+ +N
Sbjct: 18 DVKMEDGSDEEEEDPEEDPEEYEEMENGSPQHEASN-DKNAEQEAKADTKSENITTNN-- 188
Query: 356 FKSDQFCDNRDDVSTSNVSVHVENVDAKSFKNLK 389
D+ S + V E ++K+ +K
Sbjct: 189 -------KTTDETSKEEIKVKDEVQESKADPQVK 269
>TC234653 weakly similar to UP|Q6VSS7 (Q6VSS7) TOX, partial (10%)
Length = 871
Score = 33.1 bits (74), Expect = 0.55
Identities = 28/120 (23%), Positives = 45/120 (37%), Gaps = 20/120 (16%)
Frame = +2
Query: 69 DFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNG------IEESD 122
D S + D+ + G + + LS T D L NN + ++
Sbjct: 155 DADSGNGKSSDNKNQEVTNGQNPTPAANTQNGVLSETKEDVKNLPPNNEESSKLEVTDAA 334
Query: 123 GNENGGEVGEREIETEEEEEE--------------EFSEGDDSMFNYGSGCDGDNENEFY 168
G +N GE E + E E++ E EGD+ + Y G +G+N+ E Y
Sbjct: 335 GGDNDGEYEEGMVYDEAGEDQYYEGDDQRDADYEYEQGEGDEGYYEYEQGEEGENQEEDY 514
>TC211213 similar to PIR|T48518|T48518 transcription factor like protein -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(45%)
Length = 789
Score = 32.7 bits (73), Expect = 0.71
Identities = 21/68 (30%), Positives = 33/68 (47%)
Frame = -1
Query: 585 KRSVLIQECLQSILCSRFFSSPPRALVWFLSPEDSHPSSPVSNSPVSLSSFTRGENIRNS 644
+R++L C++++L PP ++W+ E HPSS N+ L S GE+
Sbjct: 783 ERTMLSDSCIETMLFMSLMKYPPS*MLWYHG*E*GHPSSMSRNTNSELRS---GESKLVP 613
Query: 645 STWGKTIS 652
S TIS
Sbjct: 612 SGGSSTIS 589
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.133 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,464,650
Number of Sequences: 63676
Number of extensions: 607695
Number of successful extensions: 3659
Number of sequences better than 10.0: 95
Number of HSP's better than 10.0 without gapping: 2997
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3312
length of query: 880
length of database: 12,639,632
effective HSP length: 106
effective length of query: 774
effective length of database: 5,889,976
effective search space: 4558841424
effective search space used: 4558841424
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC124952.2


