

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124951.10 - phase: 0
(446 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
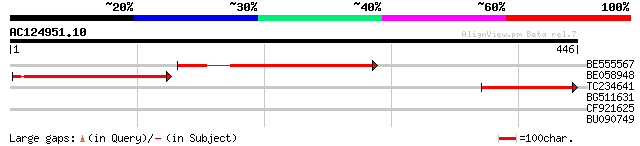
Sequences producing significant alignments: (bits) Value
BE555567 similar to GP|18087557|gb AT3g27930/K24A2_2 {Arabidopsi... 238 4e-63
BE058948 204 7e-53
TC234641 weakly similar to UP|Q8W106 (Q8W106) AT3g27930/K24A2_2,... 140 2e-33
BG511631 similar to GP|18087557|gb| AT3g27930/K24A2_2 {Arabidops... 34 0.12
CF921625 28 6.4
BU090749 28 8.3
>BE555567 similar to GP|18087557|gb AT3g27930/K24A2_2 {Arabidopsis thaliana},
partial (28%)
Length = 422
Score = 238 bits (607), Expect = 4e-63
Identities = 119/157 (75%), Positives = 128/157 (80%)
Frame = +2
Query: 133 SCAYYPRYGFGAFGVFPLLLKNRTSSQIATFTADKVRQLYRETSQDSGVMGLRYGSGNLS 192
+CAYYP+YGFGAFGVFPLLLK R E+SQD G+MGLRYGSGNLS
Sbjct: 5 ACAYYPKYGFGAFGVFPLLLKKR------------------ESSQDYGLMGLRYGSGNLS 130
Query: 193 CGVTLMPFAKKDELPKSVWLVSKIGRVTAGVQYEPHHENAKLSNLMNWSCAMAYGVGSQS 252
GVTL+PFA KDELPKS WLVSK+GR+TAGVQYEP NAKLSNLMNWSCAM YGVGS S
Sbjct: 131 FGVTLLPFAMKDELPKSAWLVSKMGRLTAGVQYEPQQGNAKLSNLMNWSCAMGYGVGSGS 310
Query: 253 PLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLE 289
PL PSFNF+LELVKSSQF+ASFYQHMVVQRRVKNPLE
Sbjct: 311 PLCPSFNFNLELVKSSQFIASFYQHMVVQRRVKNPLE 421
>BE058948
Length = 411
Score = 204 bits (519), Expect = 7e-53
Identities = 97/125 (77%), Positives = 107/125 (85%)
Frame = +3
Query: 3 IWKWIVSWYRNEVESPPPVVLVPPLFDFPPIAARNRMLESSYDVVFGKLALRCLFNDYFQ 62
+WKW+ S R PPPVVLVPPLFDFPP+AARNRML SSYDVVFGKLAL LF DYF
Sbjct: 42 MWKWLFS--REPAPPPPPVVLVPPLFDFPPLAARNRMLHSSYDVVFGKLALTSLFEDYFH 215
Query: 63 QPKHFVTRIMLKPIDDPHVDLIATVSGPLDQKPDENINGNALFRWQSDVNDPHTFMDLYV 122
Q ++F TRIMLKPI+DPHVDLIATVSGPLD KP ++I G+ALFRWQSDVNDPHTFMDLYV
Sbjct: 216 QARNFTTRIMLKPIEDPHVDLIATVSGPLDHKPKDSIAGDALFRWQSDVNDPHTFMDLYV 395
Query: 123 STSDP 127
STS+P
Sbjct: 396 STSNP 410
>TC234641 weakly similar to UP|Q8W106 (Q8W106) AT3g27930/K24A2_2, partial
(16%)
Length = 384
Score = 140 bits (352), Expect = 2e-33
Identities = 65/75 (86%), Positives = 73/75 (96%)
Frame = +3
Query: 372 ADGQVQYGFGLQSESLREASYQRADPNFVMLTPSKEHLAEGIVWETGKRPMLQSDIDAGH 431
ADG++QYGFG+QSESLREASYQRADPN+VMLT SKEHLA+GIVW+ GKRPML+SDIDAGH
Sbjct: 12 ADGKMQYGFGIQSESLREASYQRADPNYVMLTQSKEHLAKGIVWKAGKRPMLKSDIDAGH 191
Query: 432 FDGLPRELRPLDKIL 446
FDGLPREL+PLDKIL
Sbjct: 192 FDGLPRELQPLDKIL 236
>BG511631 similar to GP|18087557|gb| AT3g27930/K24A2_2 {Arabidopsis
thaliana}, partial (3%)
Length = 297
Score = 34.3 bits (77), Expect = 0.12
Identities = 15/28 (53%), Positives = 19/28 (67%)
Frame = -2
Query: 257 SFNFSLELVKSSQFVASFYQHMVVQRRV 284
S + + + QF+ASFYQHM VQRRV
Sbjct: 152 SILLKFQSLNNLQFIASFYQHMAVQRRV 69
>CF921625
Length = 523
Score = 28.5 bits (62), Expect = 6.4
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = +1
Query: 10 WYRNEVESPPPVVLVPPLFDFPP 32
W + +V +PPP + P +F FPP
Sbjct: 241 WEKKKVFNPPPPLFNPKIFFFPP 309
>BU090749
Length = 405
Score = 28.1 bits (61), Expect = 8.3
Identities = 13/41 (31%), Positives = 21/41 (50%)
Frame = -2
Query: 355 SWWKPSFTFSISATRDRADGQVQYGFGLQSESLREASYQRA 395
+WW + + + A+G+V + GLQS SL E +A
Sbjct: 404 AWWLEFSEVASALGEEEAEGEVDHKTGLQSSSLMEERIWKA 282
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.135 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,512,259
Number of Sequences: 63676
Number of extensions: 279922
Number of successful extensions: 1490
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1461
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1488
length of query: 446
length of database: 12,639,632
effective HSP length: 100
effective length of query: 346
effective length of database: 6,272,032
effective search space: 2170123072
effective search space used: 2170123072
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC124951.10


