

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124217.4 + phase: 0
(583 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
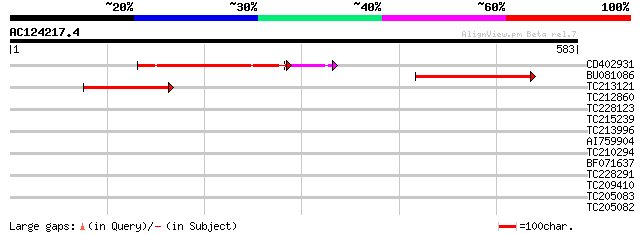
Score E
Sequences producing significant alignments: (bits) Value
CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (ja... 151 4e-41
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 147 2e-35
TC213121 89 7e-18
TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and... 35 0.12
TC228123 similar to UP|Q9LW21 (Q9LW21) Arabidopsis thaliana geno... 32 1.0
TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%) 31 1.3
TC213996 weakly similar to GB|AAD30225.1|4835758|T8K14 ESTs gb|T... 30 2.3
AI759904 similar to GP|14164420|db P0710A02.18 {Oryza sativa (ja... 30 3.0
TC210294 30 3.9
BF071637 30 3.9
TC228291 homologue to UP|Q9FXT8 (Q9FXT8) 26S proteasome regulato... 29 5.1
TC209410 weakly similar to UP|O24097 (O24097) MtN19 protein prec... 29 5.1
TC205083 weakly similar to UP|Q8GT67 (Q8GT67) Xyloglucan-specifi... 29 6.7
TC205082 weakly similar to UP|Q8GT67 (Q8GT67) Xyloglucan-specifi... 28 8.7
>CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 617
Score = 151 bits (381), Expect(2) = 4e-41
Identities = 78/158 (49%), Positives = 110/158 (69%)
Frame = +2
Query: 132 GTGKTFIWKAMSAAIRSRGEIVLTVASSGIAALLIPGGRTAHSRFGIPFTVDECSTCGVK 191
GT KTF+WK +SA +RS+G ++ + +A+LL+P G TAHS F IP + + STC +
Sbjct: 11 GTSKTFLWKTLSAGMRSKG-LMSSSCLKRLASLLLPRG*TAHSTFSIPLVIKDDSTCNIN 187
Query: 192 PNTPLAQLVVQARLIIWDEAPMMHKFCFEALDRSCRDVMKEVDKRNKNIPFGGKVVVFGG 251
+ ++L++ +LIIWDE P+M+KFCFEALDR+ +D+M +K N PFGGKVV+ GG
Sbjct: 188 QGSARSKLLLHTKLIIWDETPVMNKFCFEALDRTLQDIMASQNKDNATKPFGGKVVL-GG 364
Query: 252 DFRQILPVIPRGIRPEIVHATINSSHLWRHCEVLRLTK 289
DFRQILPVI +G R +IV + IN+S +VL+L K
Sbjct: 365 DFRQILPVIRKGSRQDIVGSAINASK-----KVLKLKK 463
Score = 35.8 bits (81), Expect(2) = 4e-41
Identities = 22/55 (40%), Positives = 31/55 (56%)
Frame = +1
Query: 283 EVLRLTKNMRLLNGASEADIEDRRKFSEWVLSIGDGTIGEDNLVDKTITIPTDLL 337
E L KNMRL + D +D R+F +W+L I DG E++ + I IP +LL
Sbjct: 442 EGLEA*KNMRLGTTNNNEDRDDIREFVDWILMIRDGNRDEED--EGEIDIPKNLL 600
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 147 bits (370), Expect = 2e-35
Identities = 70/123 (56%), Positives = 91/123 (73%)
Frame = +2
Query: 418 TPKFLNTIVASGLPNHKLRLKEGVPVMLLRNLDTKNGLCNGTRLIITRMGRYVLEGKVIS 477
T +F+N++ S LPNHK++LK +MLLRNLD GLCN TRLIIT +++E K++S
Sbjct: 56 TTEFINSLSTSRLPNHKIKLKVDSRIMLLRNLDQNEGLCNSTRLIITMFADHIIEAKIMS 235
Query: 478 GSNVGDRVFIPRLSLSPSDVRIPFKFQRKQFPLAVSFAMTINKSQGQSLQNVGVYLPAPV 537
G G+ V+IPRL+ SPS PFK R++FP+ VS+AMTINKSQGQ L +VG+YLP PV
Sbjct: 236 GKGQGNTVYIPRLATSPSQSPWPFKLIRRKFPIIVSYAMTINKSQGQLLASVGLYLPTPV 415
Query: 538 FSH 540
FSH
Sbjct: 416 FSH 424
>TC213121
Length = 713
Score = 88.6 bits (218), Expect = 7e-18
Identities = 40/92 (43%), Positives = 61/92 (65%)
Frame = +2
Query: 77 NHFIHEEINYNRRELQEEHNQLMSKMTDEQHKVYDTIMARVDEKKPGIFFLYGYGGTGKT 136
N I+ E+ Y++ L ++ +TDE +++ IM + + G++FLYGYGGTGKT
Sbjct: 335 NKLIYNELAYDKDILAA*FSRCYHSLTDEHASIFNKIMHMIATQSGGVYFLYGYGGTGKT 514
Query: 137 FIWKAMSAAIRSRGEIVLTVASSGIAALLIPG 168
F+WK +S+ I S IVLT+ASSGI ++L+PG
Sbjct: 515 FVWKTLSSTIHSNSGIVLTMASSGILSMLLPG 610
>TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and
recombination protein PIF1, mitochondrial precursor,
partial (8%)
Length = 596
Score = 34.7 bits (78), Expect = 0.12
Identities = 20/46 (43%), Positives = 30/46 (64%)
Frame = +2
Query: 513 SFAMTINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTSRGGLKI 558
++AM+I+K QG +L+ V L + F G +YVA+SRV S GL +
Sbjct: 2 AWAMSIHKCQGMTLERVHTDL-SRAFGCGMVYVALSRVRSLEGLHL 136
>TC228123 similar to UP|Q9LW21 (Q9LW21) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MDJ14, partial (9%)
Length = 831
Score = 31.6 bits (70), Expect = 1.0
Identities = 19/67 (28%), Positives = 36/67 (53%), Gaps = 3/67 (4%)
Frame = -1
Query: 393 EEKVYLSYDSPDERNVCGDAMDDV---HTPKFLNTIVASGLPNHKLRLKEGVPVMLLRNL 449
E+ Y + SP RN+ + D+ +TP F ++ + +++ L++G+PV L +
Sbjct: 267 EDPHY*NCPSPLNRNLVVSLIHDMFYPNTPFFHSSHIYQ*YTHNQASLRKGLPVFLFSKI 88
Query: 450 DTKNGLC 456
+KN LC
Sbjct: 87 ASKNPLC 67
>TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%)
Length = 340
Score = 31.2 bits (69), Expect = 1.3
Identities = 15/22 (68%), Positives = 17/22 (77%)
Frame = -3
Query: 504 QRKQFPLAVSFAMTINKSQGQS 525
QR FPL V FAMT NKS+GQ+
Sbjct: 338 QR**FPLIVCFAMTTNKSEGQT 273
>TC213996 weakly similar to GB|AAD30225.1|4835758|T8K14 ESTs gb|T22000 and
gb|AA585765 come from this gene. {Arabidopsis thaliana;}
, partial (40%)
Length = 595
Score = 30.4 bits (67), Expect = 2.3
Identities = 15/45 (33%), Positives = 25/45 (55%)
Frame = -1
Query: 7 CVSFLHQCYTLPLTYVNKCYVFDADLTLTDEQLQIYALADVEKLL 51
CV +H+CY +P+ + + + D LTD L A+ +VE +L
Sbjct: 451 CVQGIHECYVIPINVIIEVFDEDQRKVLTD-GLDSKAMVEVEFIL 320
>AI759904 similar to GP|14164420|db P0710A02.18 {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 300
Score = 30.0 bits (66), Expect = 3.0
Identities = 16/54 (29%), Positives = 28/54 (51%), Gaps = 2/54 (3%)
Frame = -1
Query: 115 ARVDEKKPGIFFLYGY--GGTGKTFIWKAMSAAIRSRGEIVLTVASSGIAALLI 166
A V + + G F+ G+ G G + + + AIRSRG +V+ V G+ ++
Sbjct: 207 AEVKKLRCGAGFIVGFAVGAVGGVGVRREQAEAIRSRGFVVVVVIGGGVCTAVV 46
>TC210294
Length = 769
Score = 29.6 bits (65), Expect = 3.9
Identities = 16/38 (42%), Positives = 22/38 (57%)
Frame = -1
Query: 458 GTRLIITRMGRYVLEGKVISGSNVGDRVFIPRLSLSPS 495
G R++ +R G G+ +S S GD + RLSLSPS
Sbjct: 385 GDRVLASRDGGITTGGERVSTSREGDSIESGRLSLSPS 272
>BF071637
Length = 364
Score = 29.6 bits (65), Expect = 3.9
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = -3
Query: 254 RQILPVIPRGIRPEIVHATINSSHLWRH 281
+Q+ V+ G+ P VHAT SH+++H
Sbjct: 272 QQLQVVLQNGVMPSHVHATQKESHMYKH 189
>TC228291 homologue to UP|Q9FXT8 (Q9FXT8) 26S proteasome regulatory particle
triple-A ATPase subunit4, partial (62%)
Length = 1079
Score = 29.3 bits (64), Expect = 5.1
Identities = 19/49 (38%), Positives = 26/49 (52%)
Frame = +3
Query: 113 IMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSRGEIVLTVASSGI 161
+ RV K P LYG GTGKT + +A+++ I + L V SS I
Sbjct: 51 LFIRVGIKPPKGVLLYGPPGTGKTLLARAIASNIEAN---FLKVVSSAI 188
>TC209410 weakly similar to UP|O24097 (O24097) MtN19 protein precursor,
partial (16%)
Length = 670
Score = 29.3 bits (64), Expect = 5.1
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = +2
Query: 5 FICVSFLHQCYTLPLTYVNKCYVFD 29
F C+S+L CY + L Y+ Y+F+
Sbjct: 593 FKCISYLAMCYVICLLYIK*FYIFE 667
>TC205083 weakly similar to UP|Q8GT67 (Q8GT67) Xyloglucan-specific fungal
endoglucanase inhibitor protein, partial (25%)
Length = 1345
Score = 28.9 bits (63), Expect = 6.7
Identities = 12/35 (34%), Positives = 20/35 (56%)
Frame = +1
Query: 174 SRFGIPFTVDECSTCGVKPNTPLAQLVVQARLIIW 208
S FG+ F + + V PN P+ LV+Q+ ++ W
Sbjct: 769 SPFGLCFESNGVGSSQVGPNVPIIDLVLQSEMVKW 873
>TC205082 weakly similar to UP|Q8GT67 (Q8GT67) Xyloglucan-specific fungal
endoglucanase inhibitor protein, partial (25%)
Length = 1773
Score = 28.5 bits (62), Expect = 8.7
Identities = 12/35 (34%), Positives = 20/35 (56%)
Frame = +1
Query: 174 SRFGIPFTVDECSTCGVKPNTPLAQLVVQARLIIW 208
S FG+ F + + V PN P+ LV+Q+ ++ W
Sbjct: 1174 SPFGLCFESNGVGSSQVGPNVPVIDLVLQSEMVKW 1278
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,898,795
Number of Sequences: 63676
Number of extensions: 334002
Number of successful extensions: 1673
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 1664
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1671
length of query: 583
length of database: 12,639,632
effective HSP length: 102
effective length of query: 481
effective length of database: 6,144,680
effective search space: 2955591080
effective search space used: 2955591080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC124217.4


