

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124217.1 - phase: 0 /pseudo
(1312 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
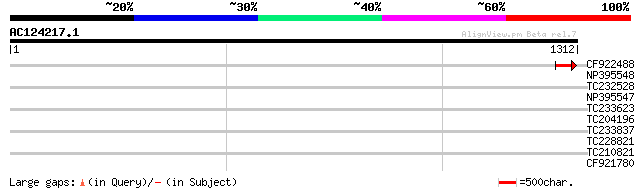
Score E
Sequences producing significant alignments: (bits) Value
CF922488 47 5e-05
NP395548 reverse transcriptase [Glycine max] 40 0.007
TC232528 weakly similar to UP|Q6WAY5 (Q6WAY5) Gag/pol polyprotei... 39 0.020
NP395547 reverse transcriptase [Glycine max] 37 0.043
TC233623 homologue to GB|AAL25580.1|16648779|AY058166 At1g69160/... 32 2.4
TC204196 similar to UP|Q9XHD5 (Q9XHD5) B12D protein, partial (94%) 32 2.4
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 31 4.1
TC228821 weakly similar to UP|Q8LRM1 (Q8LRM1) Nam-like protein 4... 31 4.1
TC210821 similar to UP|O22265 (O22265) Expressed protein (At2g47... 30 6.9
CF921780 30 6.9
>CF922488
Length = 741
Score = 47.0 bits (110), Expect = 5e-05
Identities = 21/48 (43%), Positives = 34/48 (70%)
Frame = +3
Query: 1264 LLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFIS 1311
LL F+ + +GIE++ NK K I T+K++Q FLG++N+++RFIS
Sbjct: 468 LLDFIDS*RGIEVDSNKVKVILEMAKPHTEKQVQGFLGRLNYIVRFIS 611
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 40.0 bits (92), Expect = 0.007
Identities = 20/50 (40%), Positives = 30/50 (60%)
Frame = +1
Query: 1263 ILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
I+LG I+ +GIE++Q K I P S K +++FLG+ F RFI +
Sbjct: 601 IVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQARFYRRFIKD 750
>TC232528 weakly similar to UP|Q6WAY5 (Q6WAY5) Gag/pol polyprotein
(Fragment), partial (3%)
Length = 449
Score = 38.5 bits (88), Expect = 0.020
Identities = 20/73 (27%), Positives = 39/73 (53%)
Frame = +1
Query: 700 HLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSG 759
H K L + K + KVL+D G +N++P+S L K+ + S+LKP ++ + +
Sbjct: 220 HNKALHVSVKCMDHIVAKVLIDNGYSLNVMPKSTLDKLPFNASHLKPSSMVVRAFDGTRR 399
Query: 760 SSFGVVEVDLVVG 772
G +++ + +G
Sbjct: 400 EVRGEIDLPVQIG 438
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 37.4 bits (85), Expect = 0.043
Identities = 19/50 (38%), Positives = 28/50 (56%)
Frame = +1
Query: 1263 ILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
I+LG I+K+GIE+ + K I P K + +FLG + F RFI +
Sbjct: 601 IVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGHVGFYRRFIKD 750
>TC233623 homologue to GB|AAL25580.1|16648779|AY058166 At1g69160/F4N2_9
{Arabidopsis thaliana;} , partial (10%)
Length = 421
Score = 31.6 bits (70), Expect = 2.4
Identities = 21/71 (29%), Positives = 33/71 (45%), Gaps = 3/71 (4%)
Frame = +3
Query: 489 DTPPTRHQNRGQVRTFTPSSKSPTEKWV--FSSGKKSS-YTSPPAKWVKRFTGTDNQNKA 545
++P R + R + F SS + ++ SSG ++ Y P K K F ++NK
Sbjct: 60 ESPGGRRRRRSSISHFRSSSTADSKSLYSSLSSGFRTPPYVQTPTKSCKEFRTFSSENKH 239
Query: 546 PVSKKFAYNNN 556
+S YNNN
Sbjct: 240 ALSFSAKYNNN 272
>TC204196 similar to UP|Q9XHD5 (Q9XHD5) B12D protein, partial (94%)
Length = 741
Score = 31.6 bits (70), Expect = 2.4
Identities = 23/60 (38%), Positives = 34/60 (56%), Gaps = 7/60 (11%)
Frame = -3
Query: 696 HMKIHL--KPLFIKAKVNQIGLNKVLVDGG-----VVVNLLPQSFLKKIGLSESNLKPHN 748
H +IH+ KPLFI++ + ++K VDGG V ++LPQ L I LS S + H+
Sbjct: 472 HNRIHISHKPLFIQSNLVATWVSKEAVDGGHNLRSSVAHILPQEML-SILLSLSKVLQHS 296
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 30.8 bits (68), Expect = 4.1
Identities = 12/21 (57%), Positives = 18/21 (85%)
Frame = +2
Query: 1264 LLGFVINKKGIEINQNKTKAI 1284
LLGF++++KGIEI+ K KA+
Sbjct: 338 LLGFIVSQKGIEIDPEKVKAL 400
>TC228821 weakly similar to UP|Q8LRM1 (Q8LRM1) Nam-like protein 4, partial
(10%)
Length = 913
Score = 30.8 bits (68), Expect = 4.1
Identities = 16/41 (39%), Positives = 25/41 (60%), Gaps = 3/41 (7%)
Frame = -1
Query: 830 TLLLRLTTSPKRTSTNNWRKL---CHVLLLNLVIKTMTISF 867
T+L + + S RTST +WR L C +++ NL++ T SF
Sbjct: 754 TILFKSSFSLLRTSTRDWRMLIIACVLIISNLIVSFSTESF 632
>TC210821 similar to UP|O22265 (O22265) Expressed protein
(At2g47450/T30B22.25), partial (29%)
Length = 639
Score = 30.0 bits (66), Expect = 6.9
Identities = 25/96 (26%), Positives = 36/96 (37%), Gaps = 9/96 (9%)
Frame = -3
Query: 463 PQAKRKRKWTDNK----PKFIFNKRGVPYKDTPPTRHQNRGQVRTFTP-----SSKSPTE 513
P + R W PK + +P+ PP R + RGQ R+ +P +S SPT
Sbjct: 391 PHSPRAPMWAPPNRPI*PKTPASLHPLPFSRAPPQRRRTRGQPRSLSPNPPPQTSPSPTR 212
Query: 514 KWVFSSGKKSSYTSPPAKWVKRFTGTDNQNKAPVSK 549
S + + S P V G P+ K
Sbjct: 211 WHHRPSTAQDTPNSHPFLSVPESPGPPRTRTLPLPK 104
>CF921780
Length = 238
Score = 30.0 bits (66), Expect = 6.9
Identities = 28/81 (34%), Positives = 33/81 (40%), Gaps = 8/81 (9%)
Frame = +3
Query: 463 PQAKRKRKWTDNKPKFIFNKRGVPYK--------DTPPTRHQNRGQVRTFTPSSKSPTEK 514
P RKR T K K K+GVPY+ +TPP +TF PSS
Sbjct: 39 PPRARKRAKTPKKKKL--EKKGVPYRAGIFPTNSETPP---------KTFFPSS------ 167
Query: 515 WVFSSGKKSSYTSPPAKWVKR 535
F K +PPAK V R
Sbjct: 168 --FPPFKFPPQKNPPAKGVPR 224
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.358 0.160 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,743,306
Number of Sequences: 63676
Number of extensions: 749747
Number of successful extensions: 8170
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 4511
Number of HSP's successfully gapped in prelim test: 358
Number of HSP's that attempted gapping in prelim test: 3546
Number of HSP's gapped (non-prelim): 5112
length of query: 1312
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1203
effective length of database: 5,698,948
effective search space: 6855834444
effective search space used: 6855834444
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC124217.1


