

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124216.10 - phase: 0
(543 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
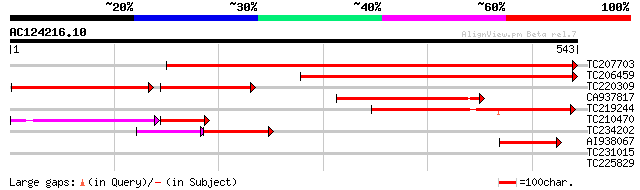
Score E
Sequences producing significant alignments: (bits) Value
TC207703 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (57%) 574 e-164
TC206459 weakly similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial ... 369 e-102
TC220309 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (31%) 190 5e-76
CA937817 172 4e-43
TC219244 similar to GB|AAO11608.1|27363378|BT002692 At3g53950/F5... 164 9e-41
TC210470 similar to GB|AAO11608.1|27363378|BT002692 At3g53950/F5... 113 3e-36
TC234202 weakly similar to UP|Q93Z02 (Q93Z02) At1g19900/F6F9_4, ... 91 5e-30
AI938067 similar to GP|9369366|gb|A F10A5.18 {Arabidopsis thalia... 69 5e-12
TC231015 weakly similar to UP|Q8S9J7 (Q8S9J7) AT5g65760/MPA24_11... 29 4.7
TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like ara... 28 8.0
>TC207703 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (57%)
Length = 1396
Score = 574 bits (1479), Expect = e-164
Identities = 283/396 (71%), Positives = 329/396 (82%), Gaps = 3/396 (0%)
Frame = +3
Query: 151 DGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPKNNI-GVYRLPFLEQTNDAGAENN 209
D GLAARRWYATNHILPDGRQIIIGGR+QFNYEFYPK Y LPFL QTNDA AENN
Sbjct: 6 DDGLAARRWYATNHILPDGRQIIIGGRRQFNYEFYPKTQAKNTYSLPFLVQTNDANAENN 185
Query: 210 LYPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNL 269
LYPFV LNVDGNLFIF+NNRAILFDY KN VVRT+PQIPGGDPR YPS+GS VLLPLKNL
Sbjct: 186 LYPFVFLNVDGNLFIFSNNRAILFDYNKNSVVRTYPQIPGGDPRCYPSTGSAVLLPLKNL 365
Query: 270 QSKFIEAEVLICGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVETMPRARVMGD 329
++ +EAEVLICGGAP+G+YQ A +F+ AL TCARIKITDPNP W +ETMP ARVM D
Sbjct: 366 RAPKVEAEVLICGGAPRGAYQNALSGKFVPALETCARIKITDPNPKWDMETMPGARVMSD 545
Query: 330 MVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHTPRMYHST 389
MV+LPNGDVLI+NGA GTAGWE GR+P+L+P LYK NN +G+RFE+Q S PRMYHS+
Sbjct: 546 MVLLPNGDVLIVNGAAVGTAGWELGRNPLLSPFLYKPNNRVGSRFEVQTSSDIPRMYHSS 725
Query: 390 AILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIVASTSELQ 449
A+L+RDGRVLV GSNPHI Y F NVLFPTEL +EAFSP YLEP F++VRP IV S+ +
Sbjct: 726 AVLLRDGRVLVAGSNPHIYYKFTNVLFPTELRLEAFSPWYLEPGFSSVRPTIVFPASQTK 905
Query: 450 -KHGQKLGLRFQV-KAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTTYD 507
K+GQ L LRF+V A L + V VTML+PPFNTHSFSMNQR+LVLE + ++ V +TY+
Sbjct: 906 LKYGQTLRLRFEVMSATLVGDSVSVTMLSPPFNTHSFSMNQRMLVLEPHDLSKVGESTYE 1085
Query: 508 VQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQIL 543
V+VT P S +LAPPGFYLLF+VH+EIPS+GIW+Q++
Sbjct: 1086VEVTAPVSAVLAPPGFYLLFLVHQEIPSQGIWVQMV 1193
>TC206459 weakly similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (37%)
Length = 970
Score = 369 bits (946), Expect = e-102
Identities = 178/266 (66%), Positives = 215/266 (79%), Gaps = 1/266 (0%)
Frame = +3
Query: 279 LICGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVETMPRARVMGDMVMLPNGDV 338
LICGGAP+G+++ +F+GAL TCARIKITDP WV+ETMP ARVM DMV+LPNGDV
Sbjct: 3 LICGGAPRGAFRNTLSGKFVGALRTCARIKITDPKANWVMETMPGARVMSDMVLLPNGDV 182
Query: 339 LIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHTPRMYHSTAILVRDGRV 398
LI+NGA GTAGWE GR+PVLNP LYK N +G RFE+QNPSH PRMYHS A+L+RDGRV
Sbjct: 183 LIVNGAAVGTAGWELGRNPVLNPFLYKPNKRVGMRFEVQNPSHIPRMYHSGAVLLRDGRV 362
Query: 399 LVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIVASTSELQ-KHGQKLGL 457
L+ GSNPH YNF VLFPTEL +EAFSP YLEP F+NVRP IV+ S+ + K+GQ L L
Sbjct: 363 LLAGSNPHTYYNFTKVLFPTELRLEAFSPWYLEPGFSNVRPAIVSPASQTKLKYGQTLRL 542
Query: 458 RFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPI 517
RF+V A L + V VTMLAPPFNTHSFSMNQRLLVL+ + ++ V +T++V+VT P S +
Sbjct: 543 RFKVSATLVGDSVSVTMLAPPFNTHSFSMNQRLLVLKPHHLSGVGESTHEVEVTAPASAV 722
Query: 518 LAPPGFYLLFVVHKEIPSEGIWIQIL 543
LAPPGFYLLFVVH+E+PS GIW+Q++
Sbjct: 723 LAPPGFYLLFVVHQEVPSHGIWVQMV 800
>TC220309 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (31%)
Length = 822
Score = 190 bits (483), Expect(2) = 5e-76
Identities = 89/136 (65%), Positives = 108/136 (78%)
Frame = +3
Query: 2 TTATYLLFLLPILITTITTTPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTD 61
T++ L L+ I+I I AA GQWQ+L K+IGIVAMHMQLLHNDR++IFDRTD
Sbjct: 75 TSSPLFLLLIIIIIIPILHFHAAEAAAKGQWQLLHKNIGIVAMHMQLLHNDRVIIFDRTD 254
Query: 62 FGLSKLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKG 121
FGLS L LP+G+CR++P E VKTDCTAHSVEY++ +NTFR LFVQT+VWCSS S +P G
Sbjct: 255 FGLSNLTLPDGRCRNNPNEMVVKTDCTAHSVEYDVVANTFRALFVQTNVWCSSASASPDG 434
Query: 122 TLVQTGGYNDGDRTIR 137
TLVQTGG+NDGDR +R
Sbjct: 435 TLVQTGGFNDGDRAVR 482
Score = 113 bits (282), Expect(2) = 5e-76
Identities = 58/92 (63%), Positives = 62/92 (67%), Gaps = 1/92 (1%)
Frame = +1
Query: 145 CDWQEFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPKNNIGVYRLPFLEQTNDA 204
CDW+E D GLAARRWYATNHILPDGRQIIIGGR+QFNYEFYPK + FL QTNDA
Sbjct: 502 CDWKEIDDGLAARRWYATNHILPDGRQIIIGGRRQFNYEFYPKTQA*TLQXTFLGQTNDA 681
Query: 205 GAENNLYPFVILNVD-GNLFIFANNRAILFDY 235
A N L P G L NNRAIL +
Sbjct: 682 YAXNKLLPVGFSKRGLGXLLSX*NNRAILIGF 777
Score = 33.1 bits (74), Expect = 0.32
Identities = 21/67 (31%), Positives = 28/67 (41%), Gaps = 1/67 (1%)
Frame = +2
Query: 184 FYPKNNIGVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAIL-FDYTKNVVVR 242
F P++ + Y L F + N YP LNVD F + D +N VVR
Sbjct: 620 FTPRHKLRHYXLLFWVKPTTLTLXTNFYPLGFLNVDWEXFYLXKTTGLS*LDSIRNNVVR 799
Query: 243 TFPQIPG 249
T+ PG
Sbjct: 800 TYL*XPG 820
>CA937817
Length = 427
Score = 172 bits (435), Expect = 4e-43
Identities = 84/141 (59%), Positives = 101/141 (71%)
Frame = +1
Query: 314 PTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGAR 373
P WV+E MP R+MGDMVMLPNGDVLIINGA SGT G+E DP LNPVLY+ + P+G R
Sbjct: 7 PRWVMEDMPFGRIMGDMVMLPNGDVLIINGAMSGTQGFEMASDPCLNPVLYRPDQPVGLR 186
Query: 374 FELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPR 433
F + NP PRMYH+TA L+ D RVL+ GSNPH+ Y FN+V FPTEL +EAFSP YL
Sbjct: 187 FLVLNPGTVPRMYHATANLLPDARVLLAGSNPHVLYRFNDVEFPTELRVEAFSPEYLSAD 366
Query: 434 FANVRPRIVASTSELQKHGQK 454
AN+RP ++ E + G K
Sbjct: 367 RANLRP-VIEEVPETVRFGGK 426
>TC219244 similar to GB|AAO11608.1|27363378|BT002692 At3g53950/F5K20_250
{Arabidopsis thaliana;} , partial (34%)
Length = 858
Score = 164 bits (415), Expect = 9e-41
Identities = 91/200 (45%), Positives = 121/200 (60%), Gaps = 4/200 (2%)
Frame = +1
Query: 347 GTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPH 406
GT G+E DP LNPVLY+ + P+G RF + NP PRMYH+TA L+ D RVL+ GSNPH
Sbjct: 1 GTQGFEMASDPCLNPVLYRPDQPVGLRFMVLNPGTVPRMYHATANLLPDARVLLAGSNPH 180
Query: 407 IGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIVASTSELQKHGQKLGLRFQVKAALD 466
+ Y F++V FPTEL +EAFSP YL AN+RP I E + G +F V ++D
Sbjct: 181 VLYRFDDVEFPTELRLEAFSPEYLSADRANLRPVI-----EEVPQTVRFGGKFDVVVSVD 345
Query: 467 ---KNLVYVTMLAPPFNTHSFSMNQRLLVLE-SNKVNIVEGTTYDVQVTMPGSPILAPPG 522
+V V + + PF THSFS QRL+ L S+ V Y + VT P S +APPG
Sbjct: 346 LPVVGIVEVNLASAPFATHSFSQGQRLVKLTVSSAVPDGSDGRYRIGVTAPPSGAVAPPG 525
Query: 523 FYLLFVVHKEIPSEGIWIQI 542
+Y+ F V++ +PS WI +
Sbjct: 526 YYMAFAVNQGVPSIAKWIHV 585
>TC210470 similar to GB|AAO11608.1|27363378|BT002692 At3g53950/F5K20_250
{Arabidopsis thaliana;} , partial (31%)
Length = 609
Score = 113 bits (282), Expect(2) = 3e-36
Identities = 59/143 (41%), Positives = 80/143 (55%)
Frame = +3
Query: 1 MTTATYLLFLLPILITTITTTPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRT 60
M +Y LFLL + T + A G W++L GI +MH + + +V+ DRT
Sbjct: 21 MENLSYFLFLLFLFAT------HARADLPGTWELLVPDAGIASMHTAVTRFNTVVLLDRT 182
Query: 61 DFGLSKLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPK 120
+ G S+ LP G CR D + +K DC AHSV ++ +N RPL + TD WCSSG P
Sbjct: 183 NIGPSRKLLPKGHCRSDKNDAVLKLDCYAHSVHLDLATNQIRPLKILTDTWCSSGQFLPD 362
Query: 121 GTLVQTGGYNDGDRTIRMFDTCN 143
GTL+QTGG DG + IR F C+
Sbjct: 363 GTLLQTGGDLDGLKKIRKFSPCD 431
Score = 57.4 bits (137), Expect(2) = 3e-36
Identities = 28/48 (58%), Positives = 30/48 (62%), Gaps = 1/48 (2%)
Frame = +1
Query: 145 CDWQEFDG-GLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPKNNIG 191
CDW+E D LA RWYATN ILPDG IIIGGR EF+P G
Sbjct: 463 CDWEELDDIELAEGRWYATNQILPDGSVIIIGGRGSNTVEFFPPKRNG 606
>TC234202 weakly similar to UP|Q93Z02 (Q93Z02) At1g19900/F6F9_4, partial
(16%)
Length = 631
Score = 90.9 bits (224), Expect(2) = 5e-30
Identities = 40/67 (59%), Positives = 51/67 (75%)
Frame = +1
Query: 186 PKNNIGVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFP 245
P + ++ LPF+ QTND ENNLYPFV LN++ LFIFAN + ILFDYTKNV+++TFP
Sbjct: 217 PHHAKNIFSLPFVVQTNDPHEENNLYPFVFLNMESTLFIFANIKVILFDYTKNVIIQTFP 396
Query: 246 QIPGGDP 252
+P GDP
Sbjct: 397 IVPHGDP 417
Score = 58.9 bits (141), Expect(2) = 5e-30
Identities = 28/67 (41%), Positives = 39/67 (57%), Gaps = 1/67 (1%)
Frame = +3
Query: 122 TLVQTGGYNDGDRTIRMFDTCNNCDWQEFD-GGLAARRWYATNHILPDGRQIIIGGRKQF 180
+LVQ G+N+G++ + F C DW E L+AR WY+TNH+LP+ QIIIG
Sbjct: 15 SLVQIDGFNNGEKKVCTFSPCPTYDWLELSFATLSARHWYSTNHLLPNDCQIIIGS*DNS 194
Query: 181 NYEFYPK 187
+PK
Sbjct: 195 TTNSFPK 215
>AI938067 similar to GP|9369366|gb|A F10A5.18 {Arabidopsis thaliana}, partial
(4%)
Length = 196
Score = 68.9 bits (167), Expect = 5e-12
Identities = 36/59 (61%), Positives = 42/59 (71%)
Frame = -1
Query: 470 VYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLFV 528
VYVTMLAPPFN HSFSM QRLLVLE + ++ V T V+VT P S +L P G Y+ FV
Sbjct: 178 VYVTMLAPPFNQHSFSMKQRLLVLEPHHLSGVGETRPKVKVTAPASNVLGPRGNYMRFV 2
>TC231015 weakly similar to UP|Q8S9J7 (Q8S9J7) AT5g65760/MPA24_11, partial
(29%)
Length = 720
Score = 29.3 bits (64), Expect = 4.7
Identities = 15/54 (27%), Positives = 26/54 (47%)
Frame = +3
Query: 181 NYEFYPKNNIGVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAILFD 234
++ F P++N + + T GA+NN FV +GN+ F N +F+
Sbjct: 183 HFNFNPQSNHTFQQRYLINDTFWGGAKNNAPIFVYTGNEGNIEWFTQNTGFMFE 344
>TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (57%)
Length = 1652
Score = 28.5 bits (62), Expect = 8.0
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = -3
Query: 329 DMVMLPNGDVLIINGAGSGTAGWEYGRD 356
D+ L GDV + GAG G G E+G D
Sbjct: 417 DLAGLEVGDVDVAGGAGGGAGGLEHGGD 334
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,118,185
Number of Sequences: 63676
Number of extensions: 367941
Number of successful extensions: 1632
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1613
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1623
length of query: 543
length of database: 12,639,632
effective HSP length: 102
effective length of query: 441
effective length of database: 6,144,680
effective search space: 2709803880
effective search space used: 2709803880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC124216.10


