

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.5 - phase: 0
(266 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
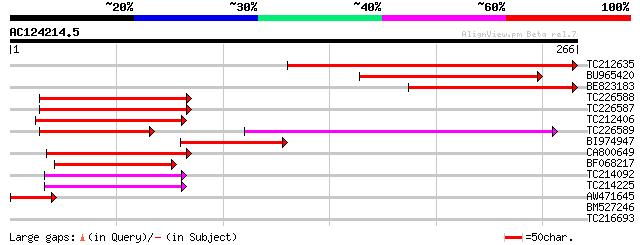
Score E
Sequences producing significant alignments: (bits) Value
TC212635 167 6e-42
BU965420 100 9e-22
BE823183 96 2e-20
TC226588 96 2e-20
TC226587 89 2e-18
TC212406 87 6e-18
TC226589 75 2e-14
BI974947 71 6e-13
CA800649 weakly similar to GP|16612258|gb| AT3g01590/F4P13_13 {A... 67 6e-12
BF068217 weakly similar to GP|8777366|dbj| apospory-associated p... 63 2e-10
TC214092 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated prote... 55 4e-08
TC214225 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated prote... 55 4e-08
AW471645 50 1e-06
BM527246 33 0.13
TC216693 27 7.3
>TC212635
Length = 837
Score = 167 bits (422), Expect = 6e-42
Identities = 97/136 (71%), Positives = 107/136 (78%)
Frame = +2
Query: 131 VR*GLKVLRHLITWTTFPKKHELQNKEML*HLNPRWIVFILAPQT*SQF*IMRGNEHIL* 190
VR*GLKV RH IT T F K++ LQNKEM *HLN RWI FILA QT* Q *IMRGNEH+L*
Sbjct: 134 VR*GLKVWRHWIT*TIFSKRNGLQNKEMQ*HLNLRWIEFILALQT*LQC*IMRGNEHLL* 313
Query: 191 ERKVSLMLPCGIHGRRNRSRWWTSVMKSTNKCFV*MVQL*RNL*T*SLERSGQGGFNSQL 250
R VSLML CGIHGRRN+++W T VM+S N CFV M Q *RNL*T*S R+G GG +S+L
Sbjct: 314 GRMVSLMLLCGIHGRRNQNQWQTLVMRSINICFVLMGQ**RNL*T*SQARNGLGGCSSRL 493
Query: 251 CDQVFVVTV*VSTEVV 266
C QVF VTV VSTEVV
Sbjct: 494 CRQVFSVTVWVSTEVV 541
>BU965420
Length = 453
Score = 100 bits (248), Expect = 9e-22
Identities = 58/86 (67%), Positives = 67/86 (77%)
Frame = +1
Query: 165 RWIVFILAPQT*SQF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWTSVMKSTNKCFV 224
RWI ILA QT* + *IMRGNEH+L* R VSLML CGIHGRRN+++ T VM+S NKCFV
Sbjct: 196 RWIECILALQT*LRC*IMRGNEHLL*GRMVSLMLLCGIHGRRNQNQCQTLVMRSINKCFV 375
Query: 225 *MVQL*RNL*T*SLERSGQGGFNSQL 250
M Q *RNL*T*S R+G GG +S+L
Sbjct: 376 LMGQ**RNL*T*SRARNGLGGCSSRL 453
>BE823183
Length = 589
Score = 95.9 bits (237), Expect = 2e-20
Identities = 54/79 (68%), Positives = 62/79 (78%)
Frame = -2
Query: 188 IL*ERKVSLMLPCGIHGRRNRSRWWTSVMKSTNKCFV*MVQL*RNL*T*SLERSGQGGFN 247
+L* R VSLML CGIHGRRN+++W T VM+S N CFV M Q *RNL*T*S R+G GG +
Sbjct: 528 LL*GRMVSLMLLCGIHGRRNQNQWQTLVMRSINICFVLMGQ**RNL*T*SQARNGLGGCS 349
Query: 248 SQLCDQVFVVTV*VSTEVV 266
S+LC QVF VTV VSTEVV
Sbjct: 348 SRLCRQVFAVTVWVSTEVV 292
>TC226588
Length = 501
Score = 95.5 bits (236), Expect = 2e-20
Identities = 44/71 (61%), Positives = 55/71 (76%)
Frame = +2
Query: 15 EFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGG 74
E +K NG+D+V+LR P+G+SA V L+GA VTSW+NEQ EELLF S K + K PK IRGG
Sbjct: 212 ELSKGINGLDKVILRDPRGSSAEVYLYGAHVTSWKNEQAEELLFLSSKAIFKPPKPIRGG 391
Query: 75 IPICFPQAGHI 85
IPICFPQ ++
Sbjct: 392 IPICFPQFSNL 424
>TC226587
Length = 566
Score = 89.4 bits (220), Expect = 2e-18
Identities = 41/71 (57%), Positives = 53/71 (73%)
Frame = +3
Query: 15 EFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGG 74
E +K NG+D+V+LR +G+SA V L+GA VTSW+N+ EELLF S K + K PK IRGG
Sbjct: 183 ELSKGINGLDKVILRDARGSSAEVYLYGAHVTSWKNDHAEELLFLSSKAIFKPPKPIRGG 362
Query: 75 IPICFPQAGHI 85
IPICFPQ ++
Sbjct: 363 IPICFPQFSNL 395
>TC212406
Length = 585
Score = 87.4 bits (215), Expect = 6e-18
Identities = 39/71 (54%), Positives = 52/71 (72%)
Frame = +3
Query: 13 ATEFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIR 72
+ E K NG+++++LR +G+S V L+G VTSW+N+ GEELLF S K + K PKAIR
Sbjct: 258 SVELCKGVNGLEKILLRESRGSSTEVYLYGGHVTSWKNDHGEELLFLSNKAIFKTPKAIR 437
Query: 73 GGIPICFPQAG 83
GGIP+CFPQ G
Sbjct: 438 GGIPLCFPQFG 470
>TC226589
Length = 1426
Score = 75.5 bits (184), Expect = 2e-14
Identities = 62/147 (42%), Positives = 76/147 (51%)
Frame = +3
Query: 111 MPSHLVSHLHTTHTCWFLT*VR*GLKVLRHLITWTTFPKKHELQNKEML*HLNPRWIVFI 170
M SH SHL H *V+ GLK RH I TT K + L N+ ML*HLN +I
Sbjct: 444 MGSHFHSHLLIIHISLCQI*VKCGLKD*RHWIILTTCRKGNALLNRGML*HLNRNLTGYI 623
Query: 171 LAPQT*SQF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWTSVMKSTNKCFV*MVQL* 230
A QF IMR E + + V LML CGI G R + VM S + CFV* + L
Sbjct: 624 *ALLQRLQFLIMRRREQLYYVKMVFLMLWCGILGTRKQRLCLILVMMSISTCFV*RLLLL 803
Query: 231 RNL*T*SLERSGQGGFNSQLCDQVFVV 257
++L *+L R+G+ N QL V VV
Sbjct: 804 KSLSL*NLARNGRED*NFQLFLLVIVV 884
Score = 59.7 bits (143), Expect = 1e-09
Identities = 28/54 (51%), Positives = 38/54 (69%)
Frame = +2
Query: 15 EFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAP 68
E +K NG+D+V+LR +G+SA V L+GA VTSW+N+ EELLF S K + P
Sbjct: 59 ELSKGINGLDKVILRDARGSSAEVYLYGAHVTSWKNDHAEELLFLSSKFSNLGP 220
>BI974947
Length = 402
Score = 70.9 bits (172), Expect = 6e-13
Identities = 38/50 (76%), Positives = 40/50 (80%)
Frame = +2
Query: 81 QAGHIVLSSAFECLLQQMET*P*YHE*GISMPSHLVSHLHTTHTCWFLT* 130
+AGHIVL+S F CLLQQMET* *Y E GI M SHLVSHLH THT W LT*
Sbjct: 86 RAGHIVLNSVFGCLLQQMET*L*YLESGI*MASHLVSHLHITHT*WSLT* 235
>CA800649 weakly similar to GP|16612258|gb| AT3g01590/F4P13_13 {Arabidopsis
thaliana}, partial (30%)
Length = 436
Score = 67.4 bits (163), Expect = 6e-12
Identities = 31/68 (45%), Positives = 45/68 (65%)
Frame = +2
Query: 18 KDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPI 77
+D +G+ +++L P+G+SA V L+G Q+ SW+N + EELLF S K K KAIRGGI
Sbjct: 173 QDKDGLPRIILTEPKGSSAEVLLYGGQIISWKNHRKEELLFMSSKANKKQHKAIRGGISA 352
Query: 78 CFPQAGHI 85
C + G +
Sbjct: 353 CLARFGDL 376
>BF068217 weakly similar to GP|8777366|dbj| apospory-associated protein C
{Arabidopsis thaliana}, partial (25%)
Length = 295
Score = 62.8 bits (151), Expect = 2e-10
Identities = 31/57 (54%), Positives = 39/57 (68%)
Frame = +2
Query: 22 GIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPIC 78
G+ + +L P+G+ A V L+GA VTSW NEQ EELLF S + + P IRGGIPIC
Sbjct: 2 GLYKDILLDPRGSYAEVFLYGAHVTSWMNEQAEELLFLSGEAMFMPP*PIRGGIPIC 172
>TC214092 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated protein C-like
protein, partial (89%)
Length = 1287
Score = 54.7 bits (130), Expect = 4e-08
Identities = 25/67 (37%), Positives = 38/67 (56%)
Frame = +1
Query: 17 TKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIP 76
T+ + ++VL +P G+ A + L G +TSW+ G++LLF + K I GG+P
Sbjct: 286 TEGEGNLPKLVLTSPAGSEAEIYLFGGCITSWKVPSGKDLLFVRPDAVFNGNKPISGGVP 465
Query: 77 ICFPQAG 83
CFPQ G
Sbjct: 466 HCFPQFG 486
>TC214225 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated protein C-like
protein, partial (86%)
Length = 1334
Score = 54.7 bits (130), Expect = 4e-08
Identities = 25/67 (37%), Positives = 38/67 (56%)
Frame = +3
Query: 17 TKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIP 76
T+ + ++VL +P G+ A + L G +TSW+ G++LLF + K I GG+P
Sbjct: 84 TEGEGNLPKLVLTSPAGSEAEIYLFGGFITSWKVPSGKDLLFVRPDAVFNGNKPISGGVP 263
Query: 77 ICFPQAG 83
CFPQ G
Sbjct: 264 HCFPQFG 284
>AW471645
Length = 103
Score = 49.7 bits (117), Expect = 1e-06
Identities = 20/22 (90%), Positives = 21/22 (94%)
Frame = +3
Query: 1 MGHSAAVWDFRAATEFTKDWNG 22
MGHSAAVWD+RAATE TKDWNG
Sbjct: 36 MGHSAAVWDYRAATEITKDWNG 101
>BM527246
Length = 422
Score = 33.1 bits (74), Expect = 0.13
Identities = 34/106 (32%), Positives = 46/106 (43%)
Frame = +2
Query: 83 GHIVLSSAFECLLQQMET*P*YHE*GISMPSHLVSHLHTTHTCWFLT*VR*GLKVLRHLI 142
GH+VLS FE L + + * E + H + H + V+ L+ RHLI
Sbjct: 98 GHVVLSFGFESL-SVLASSF*SPESETRIIKHFLLHCQ*ATIYRYQISVKYVLRDWRHLI 274
Query: 143 TWTTFPKKHELQNKEML*HLNPRWIVFILAPQT*SQF*IMRGNEHI 188
T T Q+++M L RWI AP Q * MR EH+
Sbjct: 275 TLTIC*TDQGSQSRQMQLLLMARWIGCTCAPPQKLQ**TMRRKEHL 412
>TC216693
Length = 805
Score = 27.3 bits (59), Expect = 7.3
Identities = 16/54 (29%), Positives = 25/54 (45%)
Frame = +2
Query: 39 SLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPICFPQAGHIVLSSAFE 92
S+H + +W + L +TSCK + I P+C P I+ SS+ E
Sbjct: 416 SMHCCALGTWLADLINMLSWTSCKLFNAVLLLIAPLQPLCVP*GLQILFSSSME 577
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.351 0.152 0.549
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,188,967
Number of Sequences: 63676
Number of extensions: 192972
Number of successful extensions: 1838
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 1823
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1835
length of query: 266
length of database: 12,639,632
effective HSP length: 96
effective length of query: 170
effective length of database: 6,526,736
effective search space: 1109545120
effective search space used: 1109545120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC124214.5


