

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.3 - phase: 0
(529 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
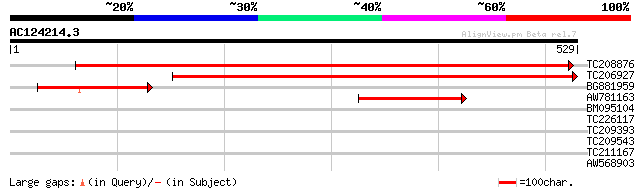
TC208876 UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase, complete 667 0.0
TC206927 homologue to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase... 521 e-148
BG881959 homologue to GP|4324495|gb|A glutamyl-tRNA reductase pr... 124 1e-28
AW781163 similar to GP|4324495|gb| glutamyl-tRNA reductase precu... 103 2e-22
BM095104 similar to GP|15215794|gb AT5g19180/T24G5_80 {Arabidops... 34 0.18
TC226117 similar to UP|Q9LL85 (Q9LL85) DNA-binding protein p24, ... 31 1.6
TC209393 similar to UP|Q9FYW4 (Q9FYW4) BAC19.12, partial (26%) 30 2.0
TC209543 similar to UP|CH10_ARATH (P34893) 10 kDa chaperonin (Pr... 30 3.5
TC211167 29 5.9
AW568903 similar to PIR|T05187|T05 chitinase (EC 3.2.1.14) class... 28 7.7
>TC208876 UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase, complete
Length = 1896
Score = 667 bits (1720), Expect = 0.0
Identities = 340/466 (72%), Positives = 404/466 (85%), Gaps = 1/466 (0%)
Frame = +1
Query: 62 SPLEILKTSSADRYTKEKSSIIVIGLNVHTAPVEMREKLAIPEAQWPQVIQELCALNHIE 121
S LE LKTS+ADRYTKE+SS++VIGL+VH+ PVEMREKLAIPEA+WP+ I ELC+LNHIE
Sbjct: 223 SALEQLKTSAADRYTKERSSVMVIGLSVHSTPVEMREKLAIPEAEWPRAIAELCSLNHIE 402
Query: 122 EAAVLSTCNRIEIYLVALSQHRGVREVTDWISKKSGVSVPEISKHQILLYNKDATQHLFE 181
EAAVLSTCNR+EIY+VALS+HRGV+EVT+W+SK SG+ V ++ +HQ LLYNKDATQHLFE
Sbjct: 403 EAAVLSTCNRMEIYVVALSKHRGVKEVTEWMSKTSGIPVADLCQHQFLLYNKDATQHLFE 582
Query: 182 VAAGLDSLVLGEGQILSQVKQVVKSGQGVPGFDRKISGLFKQAISVGKRVRTETNISSGS 241
V+AGLDSLVLGEGQIL+QVKQVVK GQGV GF R ISGLFK AI+VGKRVRTETNI++G+
Sbjct: 583 VSAGLDSLVLGEGQILAQVKQVVKVGQGVNGFGRNISGLFKHAITVGKRVRTETNIAAGA 762
Query: 242 VSVSSAAVELALMKRLESSFGDAKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSEEKVN 301
VSVSSAAVELALMK E+S +A++LVIGAGKMGKLVIKHLVAKGC KMVVVNRSEE+V
Sbjct: 763 VSVSSAAVELALMKLPEASHANARMLVIGAGKMGKLVIKHLVAKGCTKMVVVNRSEERVA 942
Query: 302 AIRKELKDVDIVYRPLSEMMECAAEADVIFTGTASESLLFSKENVEILPPVGQGV-RRRL 360
AIR+E+KDV+I+Y+PLSEM+ C EADV+FT TASE+ LF K++V+ LPP V RRL
Sbjct: 943 AIREEIKDVEIIYKPLSEMLTCIGEADVVFTSTASENPLFLKDDVKELPPATDEVGGRRL 1122
Query: 361 FVDISIPRNVDPGVSELENALVYNVDDLREVVDANKEDRQRKAMEARGIIKEELNTFEAW 420
FVDIS+PRNV +S+LE+ VYNVDDL+EVV ANKEDR RKAMEA+ II EE FEAW
Sbjct: 1123FVDISVPRNVGSCLSDLESVRVYNVDDLKEVVAANKEDRLRKAMEAQAIIGEESKQFEAW 1302
Query: 421 KDSLETVPTIKKFRAYVERIRASELEKCLSKMRGDVSKEQKEAMYALSMGIVNKLLHGPM 480
+DSLETVPTIKK RAY ERIR +ELEKCL KM D++K+ + A+ LS GIVNKLLHGPM
Sbjct: 1303RDSLETVPTIKKLRAYAERIRLAELEKCLGKMGDDINKKTQRAVDDLSRGIVNKLLHGPM 1482
Query: 481 QHLRCDGNDTKCLDEVLENMRALNRMYDLETEISLMEEKIRVKMEK 526
QHLRCDG+D++ L E LENM ALNRM++LETEIS++E+KIR K+E+
Sbjct: 1483QHLRCDGSDSRTLSETLENMHALNRMFNLETEISVLEQKIRAKVEQ 1620
>TC206927 homologue to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase, partial
(69%)
Length = 1499
Score = 521 bits (1343), Expect = e-148
Identities = 270/378 (71%), Positives = 318/378 (83%), Gaps = 1/378 (0%)
Frame = +3
Query: 153 SKKSGVSVPEISKHQILLYNKDATQHLFEVAAGLDSLVLGEGQILSQVKQVVKSGQGVPG 212
+K S V V E+S+H+ LLYN DATQHLFEV+AGLDSLVLGEGQILSQVKQVVK GQGV G
Sbjct: 27 AKTSSVPVSELSQHRFLLYNNDATQHLFEVSAGLDSLVLGEGQILSQVKQVVKVGQGVNG 206
Query: 213 FDRKISGLFKQAISVGKRVRTETNISSGSVSVSSAAVELALMKRLESSFGDAKVLVIGAG 272
F R ISGLFK AI+VGKRVRTETNI+SG+VSVSSAAVELA MK E+S +A++LVIGAG
Sbjct: 207 FGRNISGLFKHAITVGKRVRTETNIASGAVSVSSAAVELAYMKLPEASHDNARMLVIGAG 386
Query: 273 KMGKLVIKHLVAKGCRKMVVVNRSEEKVNAIRKELKDVDIVYRPLSEMMECAAEADVIFT 332
KMGKLVIKHLVAK C+KMVVVNR+EE+V AIR+ELKD++I+Y+PLSEM+ CA EAD++FT
Sbjct: 387 KMGKLVIKHLVAKSCKKMVVVNRTEERVAAIREELKDIEIIYKPLSEMLTCAGEADLVFT 566
Query: 333 GTASESLLFSKENVEILPPVGQGV-RRRLFVDISIPRNVDPGVSELENALVYNVDDLREV 391
TASE+ LF K++V+ LPP Q V RR F+DIS+PRNV VS+LE+ VYNVDDL+EV
Sbjct: 567 STASENPLFLKQHVKDLPPANQDVGGRRFFIDISVPRNVGSCVSDLESVRVYNVDDLKEV 746
Query: 392 VDANKEDRQRKAMEARGIIKEELNTFEAWKDSLETVPTIKKFRAYVERIRASELEKCLSK 451
V ANKEDR RKAMEA+ II EE FEAW+DSLETVPTIKK RAY ERIR +ELEKCL K
Sbjct: 747 VAANKEDRLRKAMEAQAIIAEESKQFEAWRDSLETVPTIKKLRAYAERIRLAELEKCLGK 926
Query: 452 MRGDVSKEQKEAMYALSMGIVNKLLHGPMQHLRCDGNDTKCLDEVLENMRALNRMYDLET 511
M D+ K+ + A+ LS GIVNKLLHGPMQHLRCDGND++ L E LENM ALNRM++LET
Sbjct: 927 MGDDIPKKTRRAVDDLSRGIVNKLLHGPMQHLRCDGNDSRTLSETLENMNALNRMFNLET 1106
Query: 512 EISLMEEKIRVKMEKAKK 529
EIS++EEKIR K+E+ +K
Sbjct: 1107EISVLEEKIRAKVEQNQK 1160
>BG881959 homologue to GP|4324495|gb|A glutamyl-tRNA reductase precursor
{Glycine max}, partial (15%)
Length = 403
Score = 124 bits (311), Expect = 1e-28
Identities = 70/116 (60%), Positives = 85/116 (72%), Gaps = 9/116 (7%)
Frame = +2
Query: 27 STFAPQRLSFTT-PKIPKCTLRSDNPLPQNFIVSKPSP--------LEILKTSSADRYTK 77
S+ +P R SFTT P+ + TL + + + S S LE LK S+ADRYTK
Sbjct: 56 SSPSPSRASFTTFPRQNRRTLIQRGFIRCDALASDASSVAPNNATALEQLKISAADRYTK 235
Query: 78 EKSSIIVIGLNVHTAPVEMREKLAIPEAQWPQVIQELCALNHIEEAAVLSTCNRIE 133
E+SSIIVIGL+VHTAPVEMREKLAIPEA+WP+ I ELC+LNHIEEAA+LSTCNR+E
Sbjct: 236 ERSSIIVIGLSVHTAPVEMREKLAIPEAEWPRAIAELCSLNHIEEAAILSTCNRME 403
>AW781163 similar to GP|4324495|gb| glutamyl-tRNA reductase precursor
{Glycine max}, partial (18%)
Length = 314
Score = 103 bits (257), Expect = 2e-22
Identities = 55/102 (53%), Positives = 71/102 (68%), Gaps = 1/102 (0%)
Frame = +3
Query: 326 EADVIFTGTASESLLFSKENVEILPPVGQGVR-RRLFVDISIPRNVDPGVSELENALVYN 384
+AD++FT ASE+ LF KE+V+ PP Q V RR F+DIS+PRNV VS+LE+ VY
Sbjct: 9 QADLVFTSAASENPLFLKEHVKDFPPASQEVGGRRFFIDISVPRNVGSCVSDLESVRVYE 188
Query: 385 VDDLREVVDANKEDRQRKAMEARGIIKEELNTFEAWKDSLET 426
VDDLREVV AN+ED + +A+ I EE F+ W+DSLET
Sbjct: 189 VDDLREVVAANEEDXRVEAVXGXAISGEEGEQFQGWRDSLET 314
>BM095104 similar to GP|15215794|gb AT5g19180/T24G5_80 {Arabidopsis
thaliana}, partial (24%)
Length = 429
Score = 33.9 bits (76), Expect = 0.18
Identities = 21/58 (36%), Positives = 34/58 (58%), Gaps = 1/58 (1%)
Frame = +3
Query: 264 AKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSE-EKVNAIRKELKDVDIVYRPLSEM 320
AKVLV+GAG +G ++K L G R + V++ E N R+ L ++ V +P +E+
Sbjct: 183 AKVLVVGAGGLGCELLKDLALSGFRNLEVIDMDRIEVTNLNRQFLFRLEDVGKPKAEV 356
>TC226117 similar to UP|Q9LL85 (Q9LL85) DNA-binding protein p24, partial
(62%)
Length = 1015
Score = 30.8 bits (68), Expect = 1.6
Identities = 20/62 (32%), Positives = 29/62 (46%)
Frame = +1
Query: 2 SMAASSSTDHTLRSSVKFSPAQFPKSTFAPQRLSFTTPKIPKCTLRSDNPLPQNFIVSKP 61
S+ + SS+ + SS+K FP F+ + L F TPK R + QN + S P
Sbjct: 88 SLLSYSSSSLSSSSSLKL----FPNHPFSSKSLPFNTPKPFSLRCRHSDLFDQNTLASTP 255
Query: 62 SP 63
P
Sbjct: 256 RP 261
>TC209393 similar to UP|Q9FYW4 (Q9FYW4) BAC19.12, partial (26%)
Length = 1422
Score = 30.4 bits (67), Expect = 2.0
Identities = 21/90 (23%), Positives = 41/90 (45%), Gaps = 15/90 (16%)
Frame = -1
Query: 20 SPAQFPKSTFAP--------QRLSFTTPKIPKCTLRSDNPLPQNFIVSKPSPLEILKTSS 71
+P + P TFA Q++S+ ++ +L +PLP+ + SKP P++++
Sbjct: 969 NPERPPSYTFALISAVLAYYQKISYQ--ELLPISLPEKSPLPKAAVTSKPPPIDVISADK 796
Query: 72 -------ADRYTKEKSSIIVIGLNVHTAPV 94
A KS ++ +G N+ P+
Sbjct: 795 PPPAVPPAAIPNPRKSRVVCLGPNIEEVPI 706
>TC209543 similar to UP|CH10_ARATH (P34893) 10 kDa chaperonin (Protein CPN10)
(Protein groES), partial (98%)
Length = 487
Score = 29.6 bits (65), Expect = 3.5
Identities = 17/55 (30%), Positives = 28/55 (50%)
Frame = -3
Query: 40 KIPKCTLRSDNPLPQNFIVSKPSPLEILKTSSADRYTKEKSSIIVIGLNVHTAPV 94
K P+ L S PQN IVSK L ++K + ++K+SI + + + P+
Sbjct: 389 KFPRIILTSIMQCPQNIIVSKQMILFVIKLHLSSPIFRQKNSITFLNSHRN*LPI 225
>TC211167
Length = 852
Score = 28.9 bits (63), Expect = 5.9
Identities = 19/52 (36%), Positives = 24/52 (45%), Gaps = 1/52 (1%)
Frame = -2
Query: 13 LRSSVKFSPAQF-PKSTFAPQRLSFTTPKIPKCTLRSDNPLPQNFIVSKPSP 63
L S+ KFSP++F P F+P + S P C S P SK SP
Sbjct: 761 LFSTSKFSPSKFSPSGLFSPSKFSPPELFSPSCKYSSSKFSPSKVSSSKFSP 606
>AW568903 similar to PIR|T05187|T05 chitinase (EC 3.2.1.14) class III acidic
- soybean, partial (41%)
Length = 414
Score = 28.5 bits (62), Expect = 7.7
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = +3
Query: 42 PKCTLRSDNPLPQNFIVSKPSPLEILKTSSAD 73
PK T + N L QNF+ +KP PL + S D
Sbjct: 312 PKDTKKVTNYLYQNFLSNKPHPLRSITLESID 407
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,181,629
Number of Sequences: 63676
Number of extensions: 220885
Number of successful extensions: 1092
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1080
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1089
length of query: 529
length of database: 12,639,632
effective HSP length: 102
effective length of query: 427
effective length of database: 6,144,680
effective search space: 2623778360
effective search space used: 2623778360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC124214.3


