

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.11 + phase: 0 /pseudo
(240 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
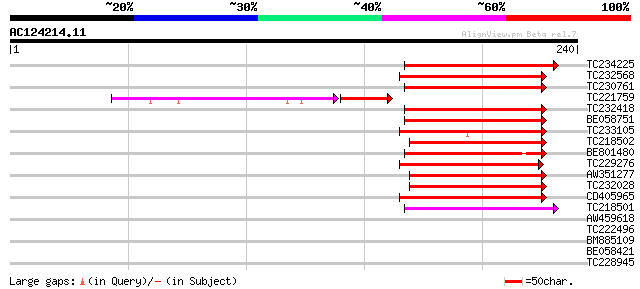
Score E
Sequences producing significant alignments: (bits) Value
TC234225 homologue to UP|O81348 (O81348) Symbiotic ammonium tran... 96 2e-20
TC232568 weakly similar to UP|Q9ZSQ9 (Q9ZSQ9) Transporter homolo... 67 9e-12
TC230761 weakly similar to UP|O81348 (O81348) Symbiotic ammonium... 66 2e-11
TC221759 homologue to UP|O81348 (O81348) Symbiotic ammonium tran... 57 7e-11
TC232418 weakly similar to UP|O81348 (O81348) Symbiotic ammonium... 60 7e-10
BE058751 59 2e-09
TC233105 58 3e-09
TC218502 58 4e-09
BE801480 56 1e-08
TC229276 weakly similar to UP|SRE2_HUMAN (Q12772) Sterol regulat... 56 2e-08
AW351277 54 5e-08
TC232028 53 1e-07
CD405965 47 8e-06
TC218501 UP|Q6Y562 (Q6Y562) ATPase 6 (Fragment), partial (5%) 43 1e-04
AW459618 31 0.58
TC222496 29 1.7
BM885109 28 4.9
BE058421 28 4.9
TC228945 27 8.3
>TC234225 homologue to UP|O81348 (O81348) Symbiotic ammonium transporter,
partial (27%)
Length = 721
Score = 95.5 bits (236), Expect = 2e-20
Identities = 48/65 (73%), Positives = 55/65 (83%)
Frame = +1
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIEARF +RNVLIR+HCEK KGV+EKTI +IEKL+LKV NSS +TFGS LDITIIAQ
Sbjct: 31 LPEIEARFYERNVLIRIHCEKNKGVIEKTISEIEKLHLKVINSSALTFGSFILDITIIAQ 210
Query: 228 PTLSF 232
+ F
Sbjct: 211 MDMEF 225
>TC232568 weakly similar to UP|Q9ZSQ9 (Q9ZSQ9) Transporter homolog
(Fragment), partial (17%)
Length = 955
Score = 66.6 bits (161), Expect = 9e-12
Identities = 30/62 (48%), Positives = 46/62 (73%)
Frame = +1
Query: 166 KRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITII 225
+RLP +EAR +++VL+R+HC+K KG++ + +I+ L+L V NSS + FG LDITI+
Sbjct: 172 ERLPRVEARVLEKDVLLRIHCQKQKGLLLNILVEIQNLHLFVVNSSVLPFGDSVLDITIV 351
Query: 226 AQ 227
AQ
Sbjct: 352 AQ 357
>TC230761 weakly similar to UP|O81348 (O81348) Symbiotic ammonium
transporter, partial (13%)
Length = 697
Score = 65.9 bits (159), Expect = 2e-11
Identities = 30/60 (50%), Positives = 43/60 (71%)
Frame = +1
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPE++ R ++VLI +HCEK KG++ K + ++E +NL V NSS + FG LDITIIA+
Sbjct: 352 LPEVKVRVLQKDVLIIIHCEKQKGIMLKILSQLENVNLSVVNSSVLRFGKITLDITIIAK 531
>TC221759 homologue to UP|O81348 (O81348) Symbiotic ammonium transporter,
partial (70%)
Length = 818
Score = 57.0 bits (136), Expect(2) = 7e-11
Identities = 53/132 (40%), Positives = 63/132 (47%), Gaps = 36/132 (27%)
Frame = +1
Query: 44 VLLLVTLHNTIHTFI-QTSTLELQWKLL-----------------------RH*RLNLFR 79
VLLL + + H+ I QTST + WK RH + NL
Sbjct: 208 VLLLERFYKSTHSLITQTSTPKHPWKHHKLELKDMQNSSEITAGITTKVNNRHQKHNLLH 387
Query: 80 TRIFFRLSI*IS*ISWD**SLRMR*LVLKTTMQLLLT-*FLKEP-----------LRPKR 127
+IFF L I I+ +WD**+LR R VLK T LL T * LKEP RPKR
Sbjct: 388 AQIFFHLLIRITQANWD**NLRWRWRVLKLTTMLLQTC*SLKEP*EIKTIFSRLAKRPKR 567
Query: 128 *QHVLNSLFLKT 139
+HVLN L LKT
Sbjct: 568 LKHVLNCLNLKT 603
Score = 26.6 bits (57), Expect(2) = 7e-11
Identities = 16/22 (72%), Positives = 17/22 (76%)
Frame = +2
Query: 141 **LKESGVRSSASASLLYLPLF 162
**LK S RSSA+ SLLYL LF
Sbjct: 608 **LKGSEERSSANGSLLYLLLF 673
>TC232418 weakly similar to UP|O81348 (O81348) Symbiotic ammonium
transporter, partial (17%)
Length = 890
Score = 60.5 bits (145), Expect = 7e-10
Identities = 31/60 (51%), Positives = 39/60 (64%)
Frame = +2
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPE+EAR + VLI +HCEK GV K + +E L+L VT SS + FG+ AL ITI Q
Sbjct: 518 LPEMEARVMGKEVLIEIHCEKENGVELKILDHLENLHLSVTGSSVLPFGNSALCITITTQ 697
>BE058751
Length = 436
Score = 59.3 bits (142), Expect = 2e-09
Identities = 30/60 (50%), Positives = 39/60 (65%)
Frame = +1
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPE+E R + VLI +HCEK GV K + +E L+L VT SS + FG+ +L ITI AQ
Sbjct: 97 LPEMEVRVLGKEVLIEIHCEKENGVELKILDHLENLHLSVTGSSVLPFGNSSLCITITAQ 276
>TC233105
Length = 586
Score = 58.2 bits (139), Expect = 3e-09
Identities = 31/63 (49%), Positives = 42/63 (66%), Gaps = 1/63 (1%)
Frame = +1
Query: 166 KRLPEIEARFCDRNVLIRVHCEKIKGV-VEKTIHKIEKLNLKVTNSSFMTFGSCALDITI 224
K LP++EAR + VLI +HCEK G+ + K + +E L+L VT SS + FG+ L ITI
Sbjct: 286 KMLPDVEARVTENEVLIEIHCEKEDGLELIKILDHLENLHLCVTASSVLPFGNSTLSITI 465
Query: 225 IAQ 227
IAQ
Sbjct: 466 IAQ 474
>TC218502
Length = 873
Score = 57.8 bits (138), Expect = 4e-09
Identities = 26/58 (44%), Positives = 41/58 (69%)
Frame = +3
Query: 170 EIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
E+E+R + +L+R+HC+K KG++ K + +I+ +L V NSS + FG LDITI+AQ
Sbjct: 357 EVESRVSGKEMLLRIHCQKQKGLLVKLLAEIQSHHLFVANSSVLPFGDSILDITIVAQ 530
>BE801480
Length = 435
Score = 56.2 bits (134), Expect = 1e-08
Identities = 27/60 (45%), Positives = 44/60 (73%)
Frame = +2
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPE+EAR + VLI+++C K KG++ K + ++E+L+L ++ S+ + FG+ LDITI AQ
Sbjct: 98 LPEVEARVLGKEVLIKIYCGKQKGILLKIMSQLERLHLYISTSNVLPFGN-TLDITITAQ 274
>TC229276 weakly similar to UP|SRE2_HUMAN (Q12772) Sterol regulatory element
binding protein-2 (SREBP-2) (Sterol regulatory
element-binding transcription factor 2), partial (3%)
Length = 1069
Score = 55.8 bits (133), Expect = 2e-08
Identities = 29/61 (47%), Positives = 41/61 (66%)
Frame = +2
Query: 166 KRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITII 225
K+ P++EAR ++VLIRV CEK K +V K + K+E NL + S+ + FG+ AL IT I
Sbjct: 650 KKCPKVEARVSGKDVLIRVTCEKQKDIVLKLLAKLEAHNLCIVCSNVLPFGNSALSITSI 829
Query: 226 A 226
A
Sbjct: 830 A 832
>AW351277
Length = 442
Score = 54.3 bits (129), Expect = 5e-08
Identities = 28/58 (48%), Positives = 35/58 (60%)
Frame = -1
Query: 170 EIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
++EAR + VLI +HCEK GV K + E L VT SS + FG+ L ITIIAQ
Sbjct: 442 DVEARVMENEVLIEIHCEKEDGVELKILDHFENLQXXVTASSVLPFGNSTLGITIIAQ 269
>TC232028
Length = 805
Score = 53.1 bits (126), Expect = 1e-07
Identities = 27/58 (46%), Positives = 37/58 (63%)
Frame = +2
Query: 170 EIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
E+EAR D+ VLI +HCEK K +V K + L+L T+S+ + FG+ L I IIAQ
Sbjct: 443 EVEARVLDKEVLIGIHCEKQKDIVFKIHALLRNLHLSTTSSTVLPFGTSTLIINIIAQ 616
>CD405965
Length = 243
Score = 47.0 bits (110), Expect = 8e-06
Identities = 23/62 (37%), Positives = 40/62 (64%)
Frame = +3
Query: 166 KRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITII 225
K P +R ++ VLIR++CEK K ++ K + ++ ++L + +SS + FG+ L+I II
Sbjct: 3 KSPPSS*SRGLEKEVLIRIYCEKRKDIMLKLLALLKDVHLSIASSSILQFGNSILNIIII 182
Query: 226 AQ 227
AQ
Sbjct: 183 AQ 188
>TC218501 UP|Q6Y562 (Q6Y562) ATPase 6 (Fragment), partial (5%)
Length = 604
Score = 42.7 bits (99), Expect = 1e-04
Identities = 22/65 (33%), Positives = 39/65 (59%)
Frame = +1
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
L +E+R + +L+++ K +G++ K + I+ +L V NSS + FG+ L ITI+AQ
Sbjct: 34 LXXVESRVSGKEMLLKIXXXKQRGLLVKLLGXIQSNHLFVANSSVLPFGNSILGITIVAQ 213
Query: 228 PTLSF 232
S+
Sbjct: 214 MXESY 228
>AW459618
Length = 372
Score = 30.8 bits (68), Expect = 0.58
Identities = 12/19 (63%), Positives = 15/19 (78%)
Frame = +2
Query: 170 EIEARFCDRNVLIRVHCEK 188
E+EAR D+ VLI +HCEK
Sbjct: 311 EVEARVLDKEVLIGIHCEK 367
>TC222496
Length = 766
Score = 29.3 bits (64), Expect = 1.7
Identities = 12/28 (42%), Positives = 20/28 (70%), Gaps = 1/28 (3%)
Frame = +3
Query: 162 FLDYKRLPEIEARFC-DRNVLIRVHCEK 188
FLDY ++P + +C D+ VL++V+ EK
Sbjct: 12 FLDYPKMPTKQVTYCNDKAVLVKVYVEK 95
>BM885109
Length = 421
Score = 27.7 bits (60), Expect = 4.9
Identities = 16/47 (34%), Positives = 26/47 (55%), Gaps = 1/47 (2%)
Frame = -1
Query: 195 KTIHKIEKLNLKVT-NSSFMTFGSCALDITIIAQPTLSFYVFHLRTF 240
KT+ K+ K+ + VT + +F+T + A + I SFY +H TF
Sbjct: 256 KTMKKVTKMTMLVTQDENFLTLHALAKCLKICCWLWSSFYSWHCGTF 116
>BE058421
Length = 345
Score = 27.7 bits (60), Expect = 4.9
Identities = 9/26 (34%), Positives = 18/26 (68%)
Frame = +1
Query: 185 HCEKIKGVVEKTIHKIEKLNLKVTNS 210
HC+K++G+ + +HK++ + TNS
Sbjct: 172 HCQKVEGLGARVVHKLDHMLNNDTNS 249
>TC228945
Length = 449
Score = 26.9 bits (58), Expect = 8.3
Identities = 10/28 (35%), Positives = 17/28 (60%)
Frame = +1
Query: 2 YEISMNIVKTCI*SCRK*WRILYFFIIS 29
Y I + + TC+* CR W+ +F ++S
Sbjct: 289 YRICLIYIATCL**CRNWWQQHFFLVVS 372
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.364 0.163 0.551
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,756,073
Number of Sequences: 63676
Number of extensions: 147912
Number of successful extensions: 1983
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1974
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1979
length of query: 240
length of database: 12,639,632
effective HSP length: 94
effective length of query: 146
effective length of database: 6,654,088
effective search space: 971496848
effective search space used: 971496848
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124214.11


