

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123976.3 - phase: 0
(447 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
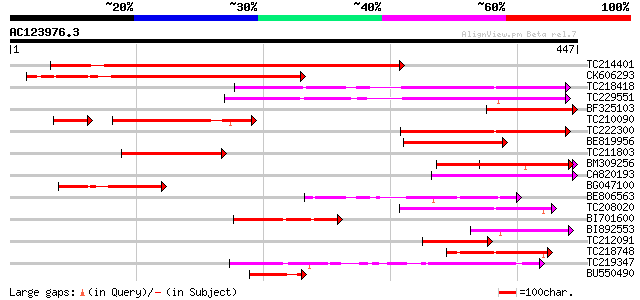
Score E
Sequences producing significant alignments: (bits) Value
TC214401 weakly similar to UP|Q949G3 (Q949G3) ABC1 protein, part... 300 1e-81
CK606293 207 1e-53
TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partia... 141 5e-34
TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, p... 137 1e-32
BF325103 117 1e-26
TC210090 similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter, par... 95 8e-24
TC222300 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {A... 106 2e-23
BE819956 similar to GP|20522008|db pleiotropic drug resistance l... 101 8e-22
TC211803 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter ... 93 2e-19
BM309256 similar to PIR|T02644|T0 ABC-type transport protein hom... 92 6e-19
CA820193 similar to PIR|T04229|T04 ABC-type transport protein F1... 77 1e-14
BG047100 similar to GP|20522008|db pleiotropic drug resistance l... 70 3e-12
BE806563 similar to GP|11994752|db ABC transporter-like protein ... 67 2e-11
TC208020 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter ... 64 1e-10
BI701600 63 3e-10
BI892553 60 2e-09
TC212091 similar to UP|Q9LFG8 (Q9LFG8) ABC transporter-like prot... 53 2e-07
TC218748 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter ... 51 9e-07
TC219347 similar to UP|Q9LF78 (Q9LF78) Mitochondrial half-ABC tr... 50 2e-06
BU550490 50 2e-06
>TC214401 weakly similar to UP|Q949G3 (Q949G3) ABC1 protein, partial (18%)
Length = 1177
Score = 300 bits (767), Expect = 1e-81
Identities = 149/279 (53%), Positives = 203/279 (72%)
Frame = +1
Query: 33 TSQVDEDEEALKWAAIEKLPTYDRLRTSIMQTFTEGDQPQPGNRQQHKEVDVTKLDMNER 92
TS+ ++DEEALKWAAIE+LPTY R+R SI+ + +EVD+ +L + ER
Sbjct: 358 TSEREDDEEALKWAAIERLPTYLRIRRSILNN----------EDGKGREVDIKQLGLTER 507
Query: 93 QQIIDKIFKVAEEDNEKYLRKFRNRIDKVGIRLPTVEVRFKNLTVEADSFVGSRALPTLP 152
+ I++++ K+AEEDNE++L K R R+D+VG+ +PT+EVRF+++ VEA +VG RALP++
Sbjct: 508 KIIVERLVKIAEEDNERFLLKLRERMDRVGLDIPTIEVRFEHINVEAQVYVGGRALPSML 687
Query: 153 NTALNILESLIGLFGFNTTKRTKLTILKNASGIVKPSRMALLLGPPSSGKTTLLLALAGK 212
N N++E + + + L IL+N SGI+KP RM LLLGPP SGKTTLLLALAGK
Sbjct: 688 NFFANVIEGFLNYLHIIPSPKKPLRILQNVSGIIKPRRMTLLLGPPGSGKTTLLLALAGK 867
Query: 213 LDSELRVQGDITYNGHRLNEFVPRKTSAYISQNDVHVGEMTVKETLDFSARCQGVGTRYD 272
LD +L G +TYNGH L EFVP++TSAYISQ D H+GEMTV+ETL FSARCQGVG Y+
Sbjct: 868 LDKDLNHSGRVTYNGHGLEEFVPQRTSAYISQYDNHIGEMTVRETLAFSARCQGVGQNYE 1047
Query: 273 LLSELARREKEAGIFPEAELDLFMKATAVKGTESSLITD 311
+L+EL RREK A I P+ ++D +MKA A+ +S++TD
Sbjct: 1048MLAELLRREKHAKIKPDPDIDAYMKAAALGRQRTSVVTD 1164
>CK606293
Length = 782
Score = 207 bits (526), Expect = 1e-53
Identities = 110/220 (50%), Positives = 153/220 (69%)
Frame = -3
Query: 14 SRSSWKMEEVFASGRYSRRTSQVDEDEEALKWAAIEKLPTYDRLRTSIMQTFTEGDQPQP 73
S S W+ + A+ +S Q + DEEALKWAAI+KLPT RLR +++ T +G+
Sbjct: 621 SSSIWRNSD--AAEIFSNSFHQ-ENDEEALKWAAIQKLPTVARLRKALI-TSPDGES--- 463
Query: 74 GNRQQHKEVDVTKLDMNERQQIIDKIFKVAEEDNEKYLRKFRNRIDKVGIRLPTVEVRFK 133
E+DV KL + E++ +++++ K A+EDNEK+L K ++RID+VGI LPT+EVRF+
Sbjct: 462 ------NEIDVKKLGLQEKKALLERLVKTAQEDNEKFLLKLKDRIDRVGIDLPTIEVRFE 301
Query: 134 NLTVEADSFVGSRALPTLPNTALNILESLIGLFGFNTTKRTKLTILKNASGIVKPSRMAL 193
NL++EA++ G+RALPT N +NILE L+ ++ L IL++ SGI+KP RM L
Sbjct: 300 NLSIEAEARAGTRALPTFTNFIVNILEGLLNSLHVLPNRKQHLNILEDVSGIIKPGRMTL 121
Query: 194 LLGPPSSGKTTLLLALAGKLDSELRVQGDITYNGHRLNEF 233
LLGPPSSGKTTLLLALAGKLD + + G +TYNGH +NEF
Sbjct: 120 LLGPPSSGKTTLLLALAGKLDPKKKFSGKVTYNGHGMNEF 1
>TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (49%)
Length = 1220
Score = 141 bits (356), Expect = 5e-34
Identities = 84/265 (31%), Positives = 140/265 (52%)
Frame = +1
Query: 178 ILKNASGIVKPSRMALLLGPPSSGKTTLLLALAGKLDSELRVQGDITYNGHRLNEFVPRK 237
+L+ +G +P L+GP SGK+TLL AL+ +L + + G I NG + +
Sbjct: 298 VLEGLTGYAEPGTFTALMGPSGSGKSTLLDALSSRLAANAFLSGTILLNGRKAK--LSFG 471
Query: 238 TSAYISQNDVHVGEMTVKETLDFSARCQGVGTRYDLLSELARREKEAGIFPEAELDLFMK 297
T+AY++Q+D +G +TV+ET+ +SAR + L + +K A
Sbjct: 472 TAAYVTQDDNLIGTLTVRETISYSARLR-------LPDNMPWADKRA------------- 591
Query: 298 ATAVKGTESSLITDYTLKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMIVGPTKTLF 357
+ + T+ +GL C DT++G+ RG+SGG+K+RV+ I+ + LF
Sbjct: 592 -----------LVESTIVAMGLQDCADTVIGNWHLRGISGGEKRRVSIALEILMRPRLLF 738
Query: 358 MDEISTGLDSSTTFQIVKCLQQIVHLTEGTILMSLLQPAPETFDLFDDIILISEGQVVYQ 417
+DE ++GLDS++ F + + L+ + T++ S+ QP+ E F+LFD + L+S G+ VY
Sbjct: 739 LDEPTSGLDSASAFFVTQTLRALAR-DGRTVIASIHQPSSEVFELFDQLYLLSSGKTVYF 915
Query: 418 GPREHIVEFFESCGFRCPERKGTAD 442
G EFF GF CP + +D
Sbjct: 916 GQASEAYEFFAQAGFPCPALRNPSD 990
>TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (34%)
Length = 1436
Score = 137 bits (345), Expect = 1e-32
Identities = 89/275 (32%), Positives = 143/275 (51%), Gaps = 2/275 (0%)
Frame = +3
Query: 170 TTKRTKLTILKNASGIVKPSRMALLLGPPSSGKTTLLLALAGKLDSELRVQGDITYNGHR 229
T + + IL +G +P R+ ++GP SGK+TLL ALAG+L S ++ G I NGH+
Sbjct: 51 TNGKNRKLILHGLTGYAQPGRLLAIIGPSGSGKSTLLDALAGRLTSNIKQTGKILINGHK 230
Query: 230 LNEFVPRKTSAYISQNDVHVGEMTVKETLDFSARCQGVGTRYDLLSELARREKEAGIFPE 289
+ + TS Y++Q+D + +T ETL +SA Q T +S ++E+
Sbjct: 231 --QELAYGTSGYVTQDDAMLSCLTAGETLYYSAMLQFPNT----MSVEEKKER------- 371
Query: 290 AELDLFMKATAVKGTESSLITDYTLKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMI 349
D TL+ +GL +T VG +G+SGGQ++R++ I
Sbjct: 372 --------------------ADMTLREMGLQDAINTRVGGWNCKGLSGGQRRRLSICIEI 491
Query: 350 VGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLT--EGTILMSLLQPAPETFDLFDDII 407
+ K LF+DE ++GLDS+ ++ ++ + ++ + TI+ S+ QP+ E F LF D+
Sbjct: 492 LTHPKLLFLDEPTSGLDSAASYYVMSGIANLIQRDGIQRTIVASVHQPSSEVFQLFHDLF 671
Query: 408 LISEGQVVYQGPREHIVEFFESCGFRCPERKGTAD 442
L+S G+ VY GP +FF S GF CP +D
Sbjct: 672 LLSSGETVYFGPASDANQFFASNGFPCPPLYNPSD 776
>BF325103
Length = 393
Score = 117 bits (292), Expect = 1e-26
Identities = 51/71 (71%), Positives = 66/71 (92%)
Frame = -2
Query: 377 LQQIVHLTEGTILMSLLQPAPETFDLFDDIILISEGQVVYQGPREHIVEFFESCGFRCPE 436
L+Q+VH+ + T+++SLLQPAPETFDLFDDIIL+SEG ++YQGPRE++++FFES GF+CPE
Sbjct: 389 LRQLVHVMDVTMIISLLQPAPETFDLFDDIILLSEGHIIYQGPRENVLDFFESVGFKCPE 210
Query: 437 RKGTADFLQEV 447
RKG ADFLQEV
Sbjct: 209 RKGIADFLQEV 177
>TC210090 similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter, partial (12%)
Length = 1189
Score = 95.1 bits (235), Expect(2) = 8e-24
Identities = 56/121 (46%), Positives = 76/121 (62%), Gaps = 8/121 (6%)
Frame = +3
Query: 82 VDVTKLDMNERQQIIDKIFKVAEEDNEKYLRKFRNRIDKVGIRLPTVEVRFKNLTVEAD- 140
VDV KL ER I+K+ K E DN + L+KFR RIDKVGI LPTVE+R++NL+VEA+
Sbjct: 282 VDVRKLGAQERHTFIEKLIKHIENDNLRLLQKFRKRIDKVGINLPTVELRYQNLSVEAEC 461
Query: 141 SFVGSRALPTLPNTALNILESLIGLFGFNTTK-------RTKLTILKNASGIVKPSRMAL 193
V + +PTL NT + F+TTK +K++I+KN +GI+KP R A+
Sbjct: 462 KIVQGKPIPTLWNTLKEWI--------FDTTKLSVLKSQNSKISIIKNDNGIIKPGRYAI 617
Query: 194 L 194
L
Sbjct: 618 L 620
Score = 33.5 bits (75), Expect(2) = 8e-24
Identities = 13/32 (40%), Positives = 24/32 (74%), Gaps = 1/32 (3%)
Frame = +2
Query: 35 QVDEDE-EALKWAAIEKLPTYDRLRTSIMQTF 65
+VD D EAL+WA I++LPT++R+ +++ +
Sbjct: 167 EVDNDAGEALQWAEIQRLPTFERITSALFDVY 262
>TC222300 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {Arabidopsis
thaliana;} , partial (37%)
Length = 801
Score = 106 bits (265), Expect = 2e-23
Identities = 53/134 (39%), Positives = 84/134 (62%)
Frame = +1
Query: 309 ITDYTLKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSS 368
ITD T+ +GL C D ++G RG+SGG+KKR++ I+ + LF+DE ++GLDS+
Sbjct: 4 ITDATIIEMGLQDCDDRLIGKWHLRGISGGEKKRLSIAL*ILTRPRLLFLDEPTSGLDSA 183
Query: 369 TTFQIVKCLQQIVHLTEGTILMSLLQPAPETFDLFDDIILISEGQVVYQGPREHIVEFFE 428
+ F +V+ L+ + E T++ S+ QP E F L+DD+ L+S G+ VY G + +EFF+
Sbjct: 184 SAFFVVQTLRNVAR-DERTVIYSIHQPDSEVFALYDDLFLLSGGETVYFGEAKSAIEFFD 360
Query: 429 SCGFRCPERKGTAD 442
GF CP ++ D
Sbjct: 361 EAGFPCPRKRNPYD 402
>BE819956 similar to GP|20522008|db pleiotropic drug resistance like protein
{Nicotiana tabacum}, partial (6%)
Length = 673
Score = 101 bits (251), Expect = 8e-22
Identities = 50/82 (60%), Positives = 64/82 (77%)
Frame = -1
Query: 311 DYTLKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTT 370
D L + GL++C DTIVG+ M RG+SGGQ+KRVTT EM+VG FMDEISTGLDSSTT
Sbjct: 466 DDRLCLKGLEVCADTIVGNAMLRGISGGQRKRVTTWEMLVGLANA*FMDEISTGLDSSTT 287
Query: 371 FQIVKCLQQIVHLTEGTILMSL 392
+QI+ L+Q VH+ +GT ++SL
Sbjct: 286 YQILNSLKQCVHILKGTTVISL 221
>TC211803 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter 1, partial
(6%)
Length = 255
Score = 93.2 bits (230), Expect = 2e-19
Identities = 39/83 (46%), Positives = 65/83 (77%)
Frame = +1
Query: 89 MNERQQIIDKIFKVAEEDNEKYLRKFRNRIDKVGIRLPTVEVRFKNLTVEADSFVGSRAL 148
+ ERQ++++++ KVAEEDNE++L K + RID+VG+ +PT+EVR+++L +EA++FVGSRAL
Sbjct: 7 IQERQKLLERLVKVAEEDNERFLLKLKERIDRVGLDIPTIEVRYEHLNIEAEAFVGSRAL 186
Query: 149 PTLPNTALNILESLIGLFGFNTT 171
P+ N+ N++E L +T+
Sbjct: 187 PSFINSVTNVVEGFFNLLHISTS 255
>BM309256 similar to PIR|T02644|T0 ABC-type transport protein homolog
F12C20.5 - Arabidopsis thaliana, partial (9%)
Length = 441
Score = 91.7 bits (226), Expect = 6e-19
Identities = 48/108 (44%), Positives = 66/108 (60%)
Frame = +1
Query: 337 GGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTEGTILMSLLQPA 396
GG KR+TTGE+++GP + LFMDEISTGLD STT+QI++ L+ +GT ++SLLQPA
Sbjct: 22 GGLHKRLTTGELLIGPARVLFMDEISTGLDRSTTYQIIRYLKHSTRALDGTTIVSLLQPA 201
Query: 397 PETFDLFDDIILISEGQVVYQGPREHIVEFFESCGFRCPERKGTADFL 444
PET++LF G G FF++ G + K + FL
Sbjct: 202 PETYELFFVCYSFVRGSDCISGSP*SCC*FFQTNGIQLS*AKKCSRFL 345
Score = 68.6 bits (166), Expect = 6e-12
Identities = 35/87 (40%), Positives = 51/87 (58%), Gaps = 10/87 (11%)
Frame = +2
Query: 371 FQIVKCLQQIVHLTEGTILMSLLQPAPETFDLFDD----------IILISEGQVVYQGPR 420
+Q+V Q ++ L++ + + P+ + F+ +IL+ EGQ+VYQGPR
Sbjct: 95 YQLVLIGQLLIKLSDILSIQPVHLMGPQLYPFFNQLQRPMNCFLYVILLCEGQIVYQGPR 274
Query: 421 EHIVEFFESCGFRCPERKGTADFLQEV 447
E V+FF+ GF CPERK ADFLQEV
Sbjct: 275 EAAVDFFKQMGFSCPERKNVADFLQEV 355
>CA820193 similar to PIR|T04229|T04 ABC-type transport protein F14M19.30 -
Arabidopsis thaliana, partial (23%)
Length = 423
Score = 77.4 bits (189), Expect = 1e-14
Identities = 38/115 (33%), Positives = 69/115 (59%)
Frame = +1
Query: 333 RGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTEGTILMSL 392
RG+SGG+++RV+ G ++ L +DE ++GLDS++ F++++ L+Q TI++S+
Sbjct: 70 RGLSGGERRRVSIGLCLLHDPAVLLLDEPTSGLDSTSAFKVMRILKQTCVSRNRTIILSI 249
Query: 393 LQPAPETFDLFDDIILISEGQVVYQGPREHIVEFFESCGFRCPERKGTADFLQEV 447
QP+ + D I+L+S+GQVV+ G + F S GF P + ++ E+
Sbjct: 250 HQPSFKILACIDRILLLSKGQVVHHGSVATLQAFLHSNGFTVPHQLNALEYAMEI 414
>BG047100 similar to GP|20522008|db pleiotropic drug resistance like protein
{Nicotiana tabacum}, partial (4%)
Length = 229
Score = 69.7 bits (169), Expect = 3e-12
Identities = 38/85 (44%), Positives = 55/85 (64%)
Frame = +2
Query: 39 DEEALKWAAIEKLPTYDRLRTSIMQTFTEGDQPQPGNRQQHKEVDVTKLDMNERQQIIDK 98
DEEALKWAA+EKLPTY+RLR ++ T + G E+DV+ L + ERQ++ ++
Sbjct: 2 DEEALKWAALEKLPTYNRLRKGLL-TASHG---------VANEIDVSDLGIQERQKLFER 151
Query: 99 IFKVAEEDNEKYLRKFRNRIDKVGI 123
+ KVA DNE+ + R ID+VG+
Sbjct: 152 LVKVAV*DNERVSVELRESIDRVGL 226
>BE806563 similar to GP|11994752|db ABC transporter-like protein {Arabidopsis
thaliana}, partial (20%)
Length = 428
Score = 66.6 bits (161), Expect = 2e-11
Identities = 57/174 (32%), Positives = 89/174 (50%), Gaps = 3/174 (1%)
Frame = +1
Query: 233 FVPRKTSAYISQNDVHVGEMTVKETLDFSARCQGVGTRYDLLSELARREKEAGIFPEAEL 292
FV RK ++ Q+DVH +TV ETL ++A + L L+R EK+ AE+
Sbjct: 7 FVKRKVG-FVPQDDVHYPHLTVLETLTYAALLR-------LPKSLSREEKKE----HAEM 150
Query: 293 DLFMKATAVKGTESSLITDYTLKILGLDICKDTIVGDEMN--RGVSGGQKKRVTTG-EMI 349
+ LGL C+++ VG M RG+SGG++KRV+ G EM+
Sbjct: 151 --------------------VIAELGLTRCRNSPVGGCMALFRGISGGERKRVSIGQEML 270
Query: 350 VGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTEGTILMSLLQPAPETFDLF 403
V P+ LF+DE ++GLDS+T IV L + T++ ++ QP+ + +F
Sbjct: 271 VNPS-LLFVDEPTSGLDSTTAQLIVSVLHGLARAGR-TVVATIHQPSSRLYRMF 426
>TC208020 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter 1, partial
(14%)
Length = 765
Score = 63.9 bits (154), Expect = 1e-10
Identities = 36/129 (27%), Positives = 75/129 (57%), Gaps = 5/129 (3%)
Frame = +2
Query: 308 LITDYTLKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDS 367
+ + ++++ L+ ++++VG G+S Q+KR+T +V +FMDE ++GLD+
Sbjct: 158 MFIEEVMELVELNPLRNSLVGLPGVSGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDA 337
Query: 368 STTFQIVKCLQQIVHLTEGTILMSLLQPAPETFDLFDDIILISE-GQVVYQGP----REH 422
+++ ++ V T T++ ++ QP+ + F+ FD++ L+ GQ +Y GP H
Sbjct: 338 RAAAIVMRTVRNTVD-TGRTVVCTIHQPSIDIFEAFDELFLMKRGGQEIYVGPLGRHSTH 514
Query: 423 IVEFFESCG 431
++++FES G
Sbjct: 515 LIKYFESIG 541
>BI701600
Length = 426
Score = 62.8 bits (151), Expect = 3e-10
Identities = 37/86 (43%), Positives = 54/86 (62%)
Frame = +3
Query: 177 TILKNASGIVKPSRMALLLGPPSSGKTTLLLALAGKLDSELRVQGDITYNGHRLNEFVPR 236
TILK +GI +P + +LGP SGK+TLL ALAG+L + G I N +L + V R
Sbjct: 69 TILKGVTGIAQPGEILAVLGPSGSGKSTLLHALAGRLHGP-GLTGTILANSSKLTKPVLR 245
Query: 237 KTSAYISQNDVHVGEMTVKETLDFSA 262
+T +++Q+D+ +TV+ETL F A
Sbjct: 246 RT-GFVTQDDILYPHLTVRETLVFCA 320
>BI892553
Length = 421
Score = 60.1 bits (144), Expect = 2e-09
Identities = 31/83 (37%), Positives = 48/83 (57%), Gaps = 2/83 (2%)
Frame = +3
Query: 364 GLDSSTTFQIVKCLQQIVHLTE--GTILMSLLQPAPETFDLFDDIILISEGQVVYQGPRE 421
GLDS+ ++ ++K + + + T++ S+ QP+ E F LFD++ L+S G+ VY GP
Sbjct: 3 GLDSAASYYVMKRIATLDKKDDVHRTVVASIHQPSSEVFQLFDNLCLLSSGRTVYFGPAS 182
Query: 422 HIVEFFESCGFRCPERKGTADFL 444
EFF S GF CP +D L
Sbjct: 183 AAKEFFASNGFPCPPLMNPSDHL 251
>TC212091 similar to UP|Q9LFG8 (Q9LFG8) ABC transporter-like protein, partial
(12%)
Length = 477
Score = 53.1 bits (126), Expect = 2e-07
Identities = 24/55 (43%), Positives = 39/55 (70%)
Frame = +3
Query: 326 IVGDEMNRGVSGGQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQI 380
++GD +R VSGG+++RV+ G I+ LF+DE ++ LDS++ F +VK LQ+I
Sbjct: 9 VIGDNGHRSVSGGERRRVSIGTDIIHNPIVLFLDEPTSNLDSTSAFMVVKVLQRI 173
>TC218748 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter 2
(Fragment), partial (37%)
Length = 1384
Score = 51.2 bits (121), Expect = 9e-07
Identities = 27/89 (30%), Positives = 55/89 (61%), Gaps = 5/89 (5%)
Frame = +3
Query: 345 TGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTEGTILMSLLQPAPETFDLFD 404
T E++ P+ +FMDE ++GLD+ +++ ++ V T T++ ++ QP+ + F+ FD
Sbjct: 3 TSELVANPS-IIFMDEPTSGLDARAAAIVMRTVRNTVD-TGRTVVCTIHQPSIDIFESFD 176
Query: 405 DIILISE-GQVVYQGP----REHIVEFFE 428
+++L+ + GQ +Y GP H++ +FE
Sbjct: 177 ELLLMKQGGQEIYVGPLGHHSSHLINYFE 263
>TC219347 similar to UP|Q9LF78 (Q9LF78) Mitochondrial half-ABC transporter,
partial (36%)
Length = 1085
Score = 50.1 bits (118), Expect = 2e-06
Identities = 58/251 (23%), Positives = 101/251 (40%), Gaps = 3/251 (1%)
Frame = +1
Query: 174 TKLTILKNASGIVKPSRMALLLGPPSSGKTTLLLALAGKLDSELRVQGDITYNGHRLNEF 233
T+ IL S +V + ++G SGK+T+L L D G I + + E
Sbjct: 85 TERKILDGISFVVPAGKSVAIVGTSGSGKSTILRLLFRFFDPH---SGSIKIDDQNIREV 255
Query: 234 VP---RKTSAYISQNDVHVGEMTVKETLDFSARCQGVGTRYDLLSELARREKEAGIFPEA 290
RK+ + Q+ V + T+ + + L+ E+E ++ A
Sbjct: 256 TLESLRKSIGVVPQDTVLFND-TIFHNIHYG--------------RLSATEEE--VYEAA 384
Query: 291 ELDLFMKATAVKGTESSLITDYTLKILGLDICKDTIVGDEMNRGVSGGQKKRVTTGEMIV 350
+ A+ T Y+ T+VG E +SGG+K+RV +
Sbjct: 385 Q------QAAIHNTIMKFPDKYS-----------TVVG-ERGLKLSGGEKQRVALARAFL 510
Query: 351 GPTKTLFMDEISTGLDSSTTFQIVKCLQQIVHLTEGTILMSLLQPAPETFDLFDDIILIS 410
L DE ++ LDS+T +I+ L+ + + + L A + D+II++
Sbjct: 511 KAPAILLCDEATSALDSTTEAEILSALKSVANNRTSIFIAHRLTTAMQC----DEIIVLE 678
Query: 411 EGQVVYQGPRE 421
G+V+ QGP E
Sbjct: 679 NGKVIEQGPHE 711
>BU550490
Length = 616
Score = 50.1 bits (118), Expect = 2e-06
Identities = 28/45 (62%), Positives = 31/45 (68%)
Frame = +2
Query: 190 RMALLLGPPSSGKTTLLLALAGKLDSELRVQGDITYNGHRLNEFV 234
R+ LLLGPP SGKTTLL ALAGKLD +LRV GH L F+
Sbjct: 71 RLTLLLGPPRSGKTTLLQALAGKLDRDLRV-------GHPLAPFL 184
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.135 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,308,627
Number of Sequences: 63676
Number of extensions: 174763
Number of successful extensions: 1011
Number of sequences better than 10.0: 117
Number of HSP's better than 10.0 without gapping: 982
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 998
length of query: 447
length of database: 12,639,632
effective HSP length: 100
effective length of query: 347
effective length of database: 6,272,032
effective search space: 2176395104
effective search space used: 2176395104
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC123976.3


